Benchmarking Evolutionary Analysis vs. Machine Learning for Protein Folding: A Comprehensive Overview for Biomedical Research
This article provides a systematic comparison of Evolutionary Analysis (EA) and Machine Learning (ML) methodologies for protein structure prediction, a critical task in drug discovery and synthetic biology.
Benchmarking Evolutionary Analysis vs. Machine Learning for Protein Folding: A Comprehensive Overview for Biomedical Research
Abstract
This article provides a systematic comparison of Evolutionary Analysis (EA) and Machine Learning (ML) methodologies for protein structure prediction, a critical task in drug discovery and synthetic biology. It explores the foundational principles of both approaches, detailing key algorithms and their real-world applications in areas like de novo protein design and drug-target interaction prediction. The content addresses significant challenges, including the poor performance of ML tools like AlphaFold on fold-switching proteins and the limitations of molecular docking, while presenting optimization strategies such as the ACE (Alternative Contact Enhancement) method and machine learning-rescored docking. Finally, it offers a rigorous validation framework, benchmarking the performance of leading ML models like AlphaFold, ESMFold, and OmegaFold on metrics of accuracy, speed, and resource consumption to guide researchers in selecting the optimal tool for their specific needs.
The Foundational Paradigms: Unpacking Evolutionary Signals and AI-Driven Prediction in Protein Folding
The Protein Folding Problem and Its Critical Role in Biotechnology and Medicine
The protein folding problem represents one of the most enduring challenges in structural biology. It centers on predicting the precise three-dimensional native structure of a protein from its linear amino acid sequence—a process fundamental to all biological function [1] [2]. This problem has profound implications, as a protein's structure directly determines its function; misfolded proteins are implicated in numerous neurodegenerative diseases, including Alzheimer's, Parkinson's, and ALS [2]. For decades, the scientific community has pursued two complementary computational approaches to tackle this problem: evolutionary algorithm (EA)-based methods, which often leverage co-evolutionary information and physical principles, and machine learning (ML)-based methods, which learn structure-prediction patterns from vast datasets of known protein structures [3] [4]. Understanding the relative strengths, limitations, and optimal applications of these paradigms is critical for researchers and drug development professionals aiming to harness computational power for biological discovery and therapeutic innovation.
Theoretical Framework and the Energy Landscape
The conceptual framework for understanding protein folding is the free energy landscape [5]. In this model, the folding process is visualized as a stochastic search across a multidimensional surface, where the protein spontaneously progresses from an ensemble of unfolded states (U) toward the native conformation (N)—the global free energy minimum [6] [5]. Evolution has selected for amino acid sequences whose energy landscapes are funnel-shaped, efficiently guiding the protein toward its functional structure while avoiding misfolding, aggregation, and long-lived metastable traps [5]. This landscape perspective provides a unified theoretical foundation for both EA and ML strategies, which can be understood as different methods for navigating this conformational space to identify the native state.
Benchmarking Computational Approaches: EA vs. ML
Machine Learning-Assisted Directed Evolution (MLDE)
Directed evolution (DE), a mainstay of protein engineering, mimics natural selection by iteratively applying mutagenesis and functional screening to accumulate beneficial mutations. However, its efficiency is severely hampered by epistasis—non-additive interactions between mutations that create rugged fitness landscapes with multiple local optima [7]. Machine learning-assisted directed evolution (MLDE) strategies address this by using supervised ML models trained on sequence-fitness data to predict high-fitness variants across the entire combinatorial landscape.
A systematic evaluation of MLDE across 16 diverse protein fitness landscapes demonstrated that MLDE consistently outperforms conventional DE [7]. The study found that the advantage of MLDE is most pronounced on landscapes that are challenging for traditional DE, particularly those with fewer active variants and more local optima. Key strategies include:
- Active Learning DE (ALDE): Uses iterative rounds of model prediction and experimental validation to refine exploration [7].
- Focused Training MLDE (ftMLDE): Enhances training set quality using zero-shot predictors, which leverage evolutionary, structural, and stability knowledge to prioritize informative variants without experimental data [7].
Table 1: Performance of MLDE Strategies Across Challenging Landscapes
| MLDE Strategy | Key Mechanism | Advantage over DE | Ideal Use Case |
|---|---|---|---|
| Standard MLDE | Single-round prediction using models trained on random sampling | Moderate | Landscapes with moderate epistasis |
| Active Learning (ALDE) | Iterative model refinement with experimental feedback | High | Resource-intensive screens; highly epistatic landscapes |
| Focused Training (ftMLDE) | Training set enriched by zero-shot predictors | Highest | Landscapes with sparse high-fitness variants |
Deep Learning for Structure Prediction
In the domain of structure prediction, deep learning models have set new standards for accuracy. These systems are trained on massive datasets of known structures from the Protein Data Bank (PDB) and leverage two key information sources: evolutionary history through Multiple Sequence Alignments (MSAs) and physico-chemical constraints [4].
Table 2: Benchmarking of Leading ML Protein Folding Tools
| Model | Key Innovation | Typical PLDDT* (Short Seq) | Inference Time (400 aa) | GPU Memory Use |
|---|---|---|---|---|
| AlphaFold | Transformer network integrating MSAs & physics | 0.89 [8] | ~210 sec [8] | ~10 GB [8] |
| ESMFold | Single-sequence inference using protein language model | 0.93 [8] | ~20 sec [8] | ~18 GB [8] |
| OmegaFold | Balanced design for accuracy & efficiency | 0.76 [8] | ~110 sec [8] | ~10 GB [8] |
| *PLDDT (Predicted Local Distance Difference Test): Confidence score (0-1) where >90 is high, <50 is low. |
A critical limitation of these ML models is their performance on intrinsically disordered proteins (IDPs) and regions (IDRs) [4]. Because they are trained predominantly on structured proteins from the PDB, they are biased toward single, stable conformations. When encountering IDPs, they often output low-confidence scores or unrealistic stable structures, highlighting a fundamental gap in their training data and design [4].
Inverse Folding and Non-Autoregressive Decoding
The inverse folding problem—designing sequences that fold into a target structure—is a critical task for protein engineering. Traditional autoregressive models generate sequences token-by-token, leading to teacher-forcing discrepancies and low efficiency [3]. The DIProT toolkit implements a non-autoregressive generative model that generates and refines the entire sequence in parallel [3]. This approach addresses the teacher-forcing problem and significantly improves generation efficiency, achieving a sequence recovery rate of 54.4% on the TS50 dataset and 50.6% on CATH4.2 [3]. DIProT integrates this model with a user-friendly interface and in-silico evaluation using ESMFold, forming a virtual design loop that allows researchers to incorporate prior knowledge and human feedback [3].
Experimental Protocols and Methodologies
Protocol for Equilibrium Unfolding
This classical experimental method determines the conformational stability of a protein by measuring its unfolding under denaturing conditions [6].
Materials:
- Protein Purification System: For producing pure, concentrated protein sample.
- Urea Stock Solution: High-purity (e.g., 10M) urea in the chosen buffer.
- Spectrofluorometer: Instrument for measuring fluorescence emission (e.g., PTI C-61).
- Circular Dichroism (CD) Spectropolarimeter: For validating secondary structure changes.
- Quartz Cuvettes: High-quality, matched cuvettes for UV spectroscopy.
Procedure:
- Prepare Denaturant Series: Create a series of solutions with incrementally increasing urea concentrations (e.g., 0 M to 8 M), ensuring constant buffer, salt, and protein concentration across all samples.
- Equilibration: Incubate all samples until equilibrium is reached. The required time must be determined empirically for each protein via time-course experiments.
- Reversibility Check: Confirm that the unfolding process is reversible by comparing the unfolding (native -> denatured) and refolding (denatured -> native) curves. The data should overlay.
- Data Collection:
- Fluorescence Emission: Acquire emission spectra (300-400 nm) following excitation at 280 nm (Trp/Tyr) or 295 nm (Trp-specific). The signal at a wavelength showing maximal change between native and unfolded states is plotted against urea concentration.
- Circular Dichroism: Measure CD signals (e.g., at 222 nm for alpha-helix content) across the same denaturant range.
- Data Analysis: Fit the resulting sigmoidal curve to a two-state or multi-state unfolding model to calculate the conformational free energy (ΔG) of unfolding.
Protocol for MLDE and Focused Training
This protocol outlines a computational workflow for implementing machine learning-assisted directed evolution.
Materials:
- Initial Variant Library: Experimental fitness data for a limited set of protein variants (training data).
- Zero-Shot Predictors: Computational tools (e.g., based on evolutionary, structural, or stability knowledge) to score sequences without experimental data.
- MLDE Software: Platforms like those described in [7] for model training and prediction.
Procedure:
- Initial Training Set Construction: Instead of random sampling, use one or more zero-shot predictors to select a focused training set of variants predicted to be high-fitness or informative.
- Model Training: Train a supervised machine learning model (e.g., regression) on the initial training set of sequences and their experimentally measured fitness values.
- Variant Prediction & Selection: Use the trained model to predict the fitness of all possible variants within the defined sequence space.
- Experimental Validation: Synthesize and experimentally test the top in silico predicted variants.
- Active Learning Loop (Optional): Incorporate the new experimental data into the training set and retrain the model for iterative rounds of prediction and validation.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key Reagents and Tools for Protein Folding Research
| Item / Reagent | Function / Application | Example / Key Property |
|---|---|---|
| Chaotropic Denaturants | Induce protein unfolding for equilibrium/kinetic studies. | Urea, Guanidinium HCl (GdmHCl) [6] |
| Reducing Agents | Prevent spurious disulfide bond formation during refolding. | Dithiothreitol (DTT), TCEP, β-mercaptoethanol [6] |
| Proteases | Probe native vs. non-native structure via specific cleavage patterns. | Used in large-scale refolding assays [2] |
| Molecular Chaperones | Assist in proper protein folding in cellular and in-vitro contexts. | Identify proteins unable to refold spontaneously [2] |
| Zero-Shot Predictors | Prioritize variant libraries for MLDE without experimental data. | Tools using evolutionary, structural, or stability data [7] |
| Structure Prediction Servers | In-silico validation of designed protein sequences. | AlphaFold, ESMFold, OmegaFold [8] [3] |
Workflow and Pathway Visualizations
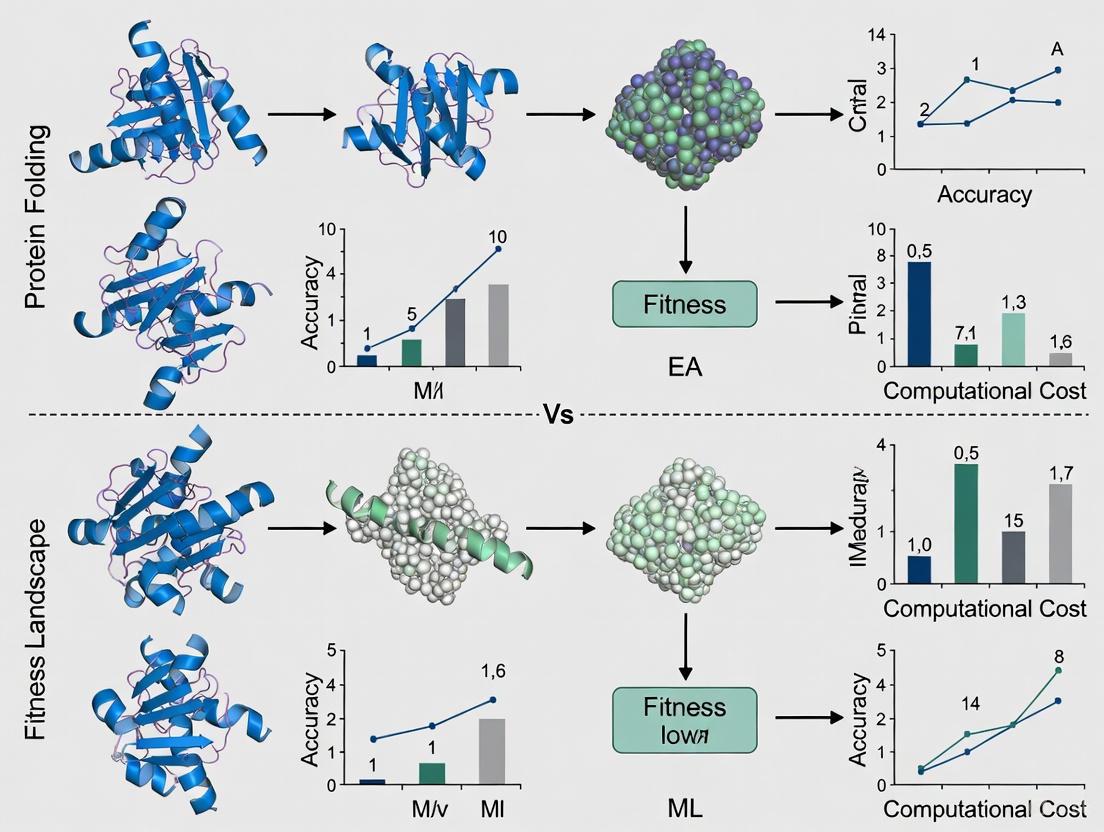
EA vs. ML for Protein Engineering
ML Protein Folding Prediction Pipeline
The benchmarking of evolutionary algorithms and machine learning for solving the protein folding problem reveals a future of complementary integration rather than outright replacement. ML methods, particularly deep learning for structure prediction, have demonstrated unprecedented speed and accuracy for structured proteins, revolutionizing the field [8] [4]. Similarly, MLDE provides a powerful advantage over naive directed evolution on complex, epistatic fitness landscapes [7]. However, EA-based methods and physical principles remain crucial, especially in areas where ML currently fails, such as designing entirely novel folds, modeling intrinsically disordered proteins, and predicting the effects of mutations in de novo designed sequences [4] [2]. The most promising path forward lies in hybrid approaches that leverage the pattern-recognition power of ML with the principled exploration of EA and physical models. This synergistic strategy will be essential for unlocking the next frontier: not just predicting structures, but reliably designing novel proteins for therapeutic and biotechnological applications.
Evolutionary Analysis (EA) represents a powerful, principles-based approach for deciphering the language of protein sequences to infer structure and function. At its core, EA operates on the fundamental biological premise that evolutionary constraints preserve functionally important relationships within and between proteins. When applied to multiple sequence alignments (MSAs), EA can detect co-evolutionary signals—patterns of correlated mutations between residue positions—that reveal which amino acids interact to maintain structural stability and biological function. These signals provide a critical source of information for protein structure prediction, function annotation, and understanding molecular recognition in signaling networks.
The resurgence of EA in structural biology is particularly notable when benchmarked against emerging machine learning (ML) methods. While ML approaches like AlphaFold2 have demonstrated remarkable accuracy, they remain profoundly dependent on the evolutionary information encoded in MSAs as primary inputs [9] [10]. This dependency underscores that EA provides the foundational biological constraints that enable modern ML systems to achieve unprecedented performance, establishing EA not as a competing methodology but as an essential component in the computational structural biology toolkit.
Theoretical Foundations: From Sequences to Structural Constraints
Multiple Sequence Alignments as Evolutionary Records
A Multiple Sequence Alignment is a computational reconstruction of evolutionary history, arranging homologous protein sequences to highlight conserved and variable regions. The construction of informative MSAs begins with searching sequence databases (e.g., UniClust30, UniProt) using tools such as HHblits or Jackhmmer to collect homologous sequences [11] [12]. The quality and depth (number of sequences) of an MSA directly impacts the strength of detectable co-evolutionary signals; alignments with hundreds or thousands of diverse sequences typically yield more reliable predictions [10].
MSAs encode two primary types of evolutionary information:
- Conservation patterns: Residues critical for function or structural stability exhibit lower evolutionary variability.
- Correlation patterns: Pairs of residues that mutate in a coordinated manner often share spatial proximity or functional linkage.
The latter pattern forms the basis for detecting co-evolution and inferring structural constraints.
The Physical Basis of Co-evolution
Coevolution occurs when mutations at one residue position necessitate compensatory mutations at another position to maintain protein fitness. In structural contexts, this frequently arises from physical interactions between residues that form stabilizing contacts. When two residues interact closely—such as in hydrogen bonding, salt bridges, or hydrophobic packing—a mutation that alters side-chain properties at one position may require complementary changes at the interacting position to preserve the interaction geometry and stability [13]. Similarly, in protein-protein interactions, co-evolution maintains complementary surfaces for specific molecular recognition [13].
From an information theory perspective, co-evolving residue pairs contain mutual information about structural constraints. The computational challenge lies in distinguishing direct couplings (which reflect physical constraints) from indirect correlations (which arise from phylogenetic relationships or other confounding factors) [12].
Methodological Approaches: Detecting Co-evolution
Computational Frameworks for Co-evolution Detection
Several computational approaches have been developed to detect co-evolution from MSAs, each with distinct theoretical foundations and implementation strategies:
Table 1: Key Computational Methods for Detecting Co-evolution
| Method | Underlying Principle | Key Tools | Strengths |
|---|---|---|---|
| Direct Coupling Analysis (DCA) | Maximum entropy model estimating direct probabilities of residue pairs | mfDCA, plmDCA, GREMLIN, CCMPred | Direct estimation of coupling parameters; avoids indirect correlations [12] |
| Inverse Covariance Methods | Sparse inverse covariance estimation to identify conditional dependencies | PSICOV | Effectively filters out transitive correlations [12] |
| Meta-Predictors | Machine learning consensus from multiple methods | metaPSICOV, PConsC, PConsC2 | Improved precision by combining orthogonal prediction sets [12] |
| Evolutionary Trace | Phylogenetic tree-based identification of functionally important residues | Evolutionary Trace | Identifies specificity-determining residues; maps functional surfaces [13] |
Experimental Protocols for Method Validation
Protocol 1: Benchmarking Co-evolution Methods for Contact Prediction
- Dataset Curation: Select a diverse set of protein domains with known structures and minimal sequence similarity (e.g., <30% identity) [12].
- MSA Generation: For each target, build an MSA using HHblits (3 iterations, 99% sequence identity threshold, 60% coverage with master sequence) against the UniProt20 database [12].
- Contact Prediction: Apply co-evolution methods (e.g., metaPSICOV, PSICOV, GREMLIN) to predict residue-residue contacts.
- Performance Assessment:
- Define structural contacts as Cβ atoms (Cα for Gly) within 8Å [12].
- Exclude trivial contacts (sequence separation <5).
- Calculate precision as: Precision = True Positives / (True Positives + False Positives)
- Focus evaluation on long-range contacts (sequence separation >23) which are most informative for structure [12].
Protocol 2: De Novo Structure Prediction Using Co-evolution Constraints
- Contact Prediction: Generate top L/5 long-range contact predictions (where L is protein length) using the optimal co-evolution method [12].
- Fragment Assembly: Use contact restraints alongside secondary structure predictions in fragment-based structure prediction software (e.g., FRAGFOLD, Rosetta) [12].
- Decoy Generation: Generate thousands of structural decoys satisfying the contact constraints.
- Model Selection: Identify best models using:
- Satisfaction of predicted contacts
- Knowledge-based energy functions
- Structural quality assessment tools
Benchmarking studies have demonstrated that this approach can generate correct folds for a substantial proportion of targets when reliable MSAs are available [12].
Quantitative Benchmarking of Co-evolution Methods
Performance Comparison Across Protein Classes
Systematic evaluation of co-evolution methods reveals significant variation in performance across different protein structural classes:
Table 2: Performance Comparison of Co-evolution Methods Across SCOP Classes
| Method | All α (%) | All β (%) | α/β (%) | Membrane Proteins (%) | Overall Average Precision (%) |
|---|---|---|---|---|---|
| metaPSICOV Stage 2 | 38.5 | 61.2 | 59.8 | 32.1 | 52.9 |
| PConsC2 | 36.8 | 59.7 | 58.3 | 30.5 | 50.8 |
| GREMLIN | 35.2 | 57.4 | 56.1 | 28.9 | 48.9 |
| PSICOV | 33.7 | 55.8 | 54.6 | 27.3 | 47.1 |
| FreeContact | 30.4 | 52.1 | 51.3 | 24.8 | 43.2 |
Precision values represent the percentage of correct contacts among the top L/5 predictions for each protein class [12].
Key observations from these benchmarks include:
- All-β and α/β proteins generally yield higher precision predictions, likely due to stronger co-evolutionary signals from extensive residue contacts.
- All-α and membrane proteins present greater challenges, with precision values typically 10-20% lower [12].
- Consensus methods (metaPSICOV, PConsC2) consistently outperform individual methods by leveraging complementary strengths.
MSA Depth Requirements for Reliable Prediction
The relationship between MSA depth (number of effective sequences) and prediction accuracy follows a nonlinear pattern:
- Minimum threshold: Approximately 5×L effective sequences (where L is protein length) are needed for basic signal detection [12].
- Saturation point: Diminishing returns observed beyond 50×L sequences for most methods.
- Quality dependence: MSA diversity (sequence variation) proves equally important as raw sequence count.
EA in Modern Protein Structure Prediction Pipelines
Integration with Deep Learning: The AlphaFold2 Paradigm
AlphaFold2 represents the most successful integration of EA principles with deep learning architecture. Its neural network explicitly reasons about evolutionary relationships through several key components:
- Evoformer Architecture: A novel neural network block that jointly processes MSA and pair representations, enabling continuous information exchange between evolutionary and spatial reasoning [9].
- Triangle Attention Mechanisms: Enforce geometric consistency in pairwise predictions through triangle multiplicative updates and triangle attention [9].
- Iterative Refinement: The recycling process allows multiple rounds of structural refinement using the same network [10].
The critical role of MSAs in AlphaFold2's performance is evidenced by:
- Strong correlation between MSA depth and prediction accuracy (pLDDT score) [10].
- Dramatic performance reduction when using single sequences instead of MSAs [14].
- The system's ability to identify co-evolving residue pairs and translate them into spatial constraints [10].
MSA-Free Approaches: The Emerging Role of Protein Language Models
Recent advances in protein language models (pLMs) like ESM represent a shift toward implicit evolutionary learning. These models:
- Pre-training: Learn statistical patterns from billions of protein sequences using self-supervised objectives (masked language modeling) [14] [15].
- Embeddings: Encode evolutionary constraints in dense vector representations without explicit MSAs [15].
- Performance: Approach MSA-based methods for targets with many homologs but excel at speed, processing sequences in seconds rather than minutes [14].
Hybrid approaches like HelixFold-Single combine pLMs with geometric learning components from AlphaFold2, demonstrating competitive accuracy on targets with large homologous families while being significantly faster than MSA-based methods [14].
Research Reagents and Computational Tools
Table 3: Essential Research Reagents and Computational Tools for Evolutionary Analysis
| Tool/Resource | Type | Function | Application Context |
|---|---|---|---|
| HHblits | Software | Rapid MSA generation via iterative hidden Markov model searches | Building deep MSAs from sequence databases [12] |
| UniProt20/UniClust30 | Database | Curated protein sequence databases with clustered sequences | Source of homologous sequences for MSA construction [11] [12] |
| metaPSICOV | Software | Meta-predictor combining multiple co-evolution methods | High-precision contact prediction from MSAs [12] |
| GREMLIN/CCMPred | Software | plmDCA implementation for direct coupling analysis | Residue-residue contact prediction [12] |
| AlphaFold2 | Software | End-to-end deep learning structure prediction | Protein 3D structure prediction using MSA inputs [9] [10] |
| ESMFold | Software | Protein language model for structure prediction | Fast structure prediction without explicit MSA generation [14] |
Evolutionary Analysis remains an indispensable methodology for protein structure and function prediction, even as machine learning approaches dominate recent advancements. The core principles of co-evolution—detecting evolutionarily coupled residues to infer structural constraints—provide biologically grounded signals that enhance the interpretability and reliability of computational predictions. Rather than being supplanted by ML, EA has been productively integrated into state-of-the-art systems where it continues to provide the evolutionary context essential for accurate structure prediction.
Future directions will likely focus on:
- Hybrid approaches combining explicit co-evolution analysis with protein language models.
- Improved contact precision for challenging protein classes (membrane proteins, all-α).
- Integration with experimental data to validate functional predictions from co-evolution analysis.
For researchers benchmarking EA against pure ML approaches, the critical insight is that these methodologies are increasingly synergistic rather than competitive, with EA providing the fundamental biological constraints that guide and validate ML-based predictions.
Workflow Visualization
Evolutionary Analysis Workflow
The field of protein structure prediction has undergone a profound transformation, shifting from a reliance on physics-based simulations to the dominance of deep learning methodologies. This revolution, catalyzed by artificial intelligence (AI), has not only solved a decades-old scientific challenge but has also fundamentally reshaped the toolkit available to researchers and drug development professionals. This technical guide examines this paradigm shift within the context of benchmarking evolutionary algorithm (EA)-inspired methods against machine learning (ML) approaches. We detail the core architectures, provide quantitative performance comparisons, and outline experimental protocols that highlight how ML has overcome the inherent limitations of classical physics-based and EA-driven protein folding models.
The Classical Era: Physics-Based Models and Evolutionary Principles
Before the rise of deep learning, computational protein structure prediction relied heavily on physics-based principles and evolutionary information. These methods were grounded in the paradigm that a protein's native state corresponds to its global free energy minimum [16]. A key breakthrough was the development of fragment assembly, an approach pioneered by methods like Rosetta and QUARK [17]. These methods operated on the principle that local sequence segments prefer local structures found in the database of known proteins. They would identify short (3-9 residue) structural fragments from experimentally solved structures based on sequence and local predicted structure similarity, and then assemble these fragments into full-length models using Monte Carlo or Replica-Exchange Monte Carlo (REMC) simulations guided by knowledge-based or physics-based force fields [17].
Another cornerstone of pre-ML methods was the use of evolutionary coupling analysis derived from Multiple Sequence Alignments (MSAs). The hypothesis was that pairs of residues in contact within a protein's structure would exhibit correlated mutations across evolution. Early methods used simple metrics like mutual information, but accuracy was low due to an inability to distinguish direct from indirect couplings. The introduction of global statistical models, particularly Direct Coupling Analysis (DCA) and Markov Random Fields (MRFs), represented a significant advance by simultaneously considering all pairwise interactions to infer direct contacts, thereby improving prediction accuracy [17].
Despite their ingenuity, these classical approaches faced fundamental limitations. Physics-based force fields were approximations, and accurate computation of the full energy landscape was challenging, often leading to misfolded designs in vitro [16]. Furthermore, the conformational sampling required by methods like fragment assembly was computationally intensive and time-consuming, restricting throughput and the exploration of novel protein folds [16] [17].
Table 1: Comparison of Classical Physics-Based/EA and Modern ML Protein Folding Methods
| Feature | Classical Physics-Based/EA Methods | Modern ML Methods |
|---|---|---|
| Core Principle | Free energy minimization, fragment assembly, evolutionary coupling [16] [17] | Learning sequence-structure mappings from data using deep neural networks [16] [18] |
| Key Algorithms | Rosetta, QUARK, I-TASSER, DCA [17] | AlphaFold2, RoseTTAFold, ESMFold, ProteinMPNN [18] [19] |
| Primary Input | Amino acid sequence, MSAs, predicted local features [17] | Amino acid sequence (and MSAs for some models) [18] [20] |
| Sampling Method | Monte Carlo, REMC, gradient descent [17] | End-to-end forward pass, inference [18] [21] |
| Computational Cost | High (hours to days per target) [16] | Relatively low (seconds to minutes per target) [20] |
| Accuracy (on single domains) | Moderate, struggled with distant homology [17] | High, often approaching experimental accuracy [18] [21] |
| Strength | Physics-based rationale, ability to explore de novo folds | Speed, accuracy, ability to leverage evolutionary scale data |
Diagram 1: Classical protein folding workflow, illustrating the iterative, sampling-based approach.
The Deep Learning Revolution: Architectures and Breakthroughs
The application of deep learning to protein folding represents a fundamental shift from iterative simulation to direct prediction. This transition was marked by the critical assessment of protein structure prediction (CASP) competitions, where ML methods demonstrated unprecedented accuracy.
Key Architectural Innovations
The success of modern ML models stems from several key architectural innovations:
Attention Mechanisms and Transformers: AlphaFold2's core innovation was the Evoformer, a specialized transformer module that processes both the MSA and a pair representation of residues [21]. This allows the model to reason about long-range interactions and co-evolutionary signals simultaneously and globally, overcoming the limitations of earlier local and pairwise statistics like DCA.
End-to-End Differentiable Learning: Unlike classical pipelines with separate stages for feature generation, sampling, and scoring, models like AlphaFold2 are trained end-to-end [18] [17]. This means the entire network is optimized for the final task—producing accurate atomic coordinates—allowing it to learn complex, implicit mappings from sequence to structure that were previously manually engineered.
Equivariance: RoseTTAFold and other models incorporate principles of equivariance, ensuring that their predictions are transformationally invariant (e.g., rotating the input sequence should not change the predicted structure, only its orientation in space). This built-in geometric awareness is crucial for robust structure prediction [18].
The impact of these innovations is best illustrated by the dramatic performance leap in CASP. As shown in Diagram 2, AlphaFold2 achieved a score nearly three times that of the top-tier methods from just six years prior, a milestone considered to have largely solved the single-domain protein folding problem [21].
Diagram 2: Simplified AlphaFold2-style architecture highlighting the Evoformer and end-to-end learning.
From Structure Prediction to Protein Design
The revolution quickly expanded from prediction to design. The inverse problem—finding a sequence that folds into a desired structure—has been tackled by new deep learning models. Inverse folding methods, such as ProteinMPNN and ESM-IF, take a backbone structure as input and generate sequences that are likely to fold into it [18]. This has dramatically improved the success rate and efficiency of de novo protein design.
Furthermore, structure prediction models themselves have been repurposed as generative models. Tools like RFdiffusion use diffusion models, trained on the principles of AF2, to generate novel protein structures either unconditionally or conditioned on specific functional motifs, opening the door to designing proteins not seen in nature [18].
Table 2: Key ML Models in Protein Structure Prediction and Design
| Model | Primary Function | Core Innovation | Typical Use Case |
|---|---|---|---|
| AlphaFold2 [18] [21] | Structure Prediction | Evoformer, end-to-end learning | High-accuracy single-structure prediction from sequence |
| RoseTTAFold [18] | Structure Prediction | 3-track network (sequence, distance, 3D) | Accurate structure prediction, basis for design tools |
| ESMFold [18] [19] | Structure Prediction | Protein language model (single-sequence) | Fast prediction for orphan sequences, high-throughput |
| SimpleFold [20] | Structure Prediction | Flow-matching with standard transformers | Challenges need for complex, domain-specific architectures |
| ProteinMPNN [18] | Inverse Folding/Design | Message-Passing Neural Network | Robust sequence design for given backbones |
| RFdiffusion [18] | De Novo Design | Diffusion model based on RoseTTAFold | Generating novel protein structures and binders |
Experimental Protocols and Benchmarking
Benchmarking the performance of ML against classical methods requires rigorous experimental protocols. The community-wide standard is the CASP (Critical Assessment of protein Structure Prediction) experiment, a biennial blind trial where groups predict the structures of recently solved but unpublished proteins [21].
CASP Benchmarking Protocol
- Target Selection: Organizers release amino acid sequences of proteins whose structures have been experimentally determined but not published.
- Model Submission: Research teams worldwide submit their predicted 3D models within a set timeframe.
- Accuracy Assessment: Predictions are compared to the experimental ground truth using metrics like the Global Distance Test (GDT_TS), which measures the percentage of residues placed within a threshold distance of their correct position [21] [17].
The results from CASP13 (2018) and CASP14 (2020) quantitatively demonstrated ML's supremacy. As shown in Diagram 3, AlphaFold2's median GDT_TS score of ~92 for the hardest targets was comparable to experimental methods, far exceeding the best classical methods [21].
Diagram 3: Qualitative performance leap of ML models in CASP.
Protocol for Iterative Protein Optimization
Beyond static structure prediction, ML guides functional protein optimization. The DeepDE algorithm provides a protocol for directed evolution guided by deep learning [22]:
- Initial Library Construction: Generate a diverse library of ~1,000 protein mutants (e.g., triple mutants) and measure their fitness (e.g., fluorescence for GFP).
- Model Training: Train a supervised deep learning model on the sequence-fitness data from the initial library.
- In Silico Exploration: Use the trained model to virtually screen a vast space of triple mutants and select top candidates for synthesis.
- Experimental Validation: Synthesize and test the predicted high-performance mutants.
- Iteration: Incorporate the new experimental data into the training set and repeat steps 2-4.
This protocol, applied to GFP, achieved a 74.3-fold increase in activity in just four rounds, far surpassing conventional directed evolution [22]. It demonstrates how ML mitigates the "combinatorial explosion" of sequence space by learning a predictive fitness landscape from limited but smartly chosen data.
The Scientist's Toolkit: Research Reagent Solutions
The modern computational protein folding and design workflow relies on a suite of software tools and databases that function as essential "research reagents."
Table 3: Essential Research Reagents for ML-Driven Protein Science
| Item Name | Type | Function / Application | Access |
|---|---|---|---|
| AlphaFold2 [18] [21] | Software Model | High-accuracy protein structure prediction from sequence. | Open source; also via AlphaFold DB |
| RoseTTAFold [18] | Software Model | Accurate structure prediction; base network for design tools like RFdiffusion. | Open source |
| ProteinMPNN [18] | Software Model | Inverse folding for designing sequences that fold into a given backbone. | Open source |
| ESMFold [18] [19] | Software Model | Fast, single-sequence-based structure prediction using a protein language model. | Open source |
| AlphaFold DB [16] | Database | Repository of pre-computed AlphaFold2 predictions for over 200 million sequences. | Publicly accessible |
| PDB | Database | Primary repository for experimentally determined protein structures; used for training and validation. | Publicly accessible |
| FiveFold Framework [19] | Ensemble Method | Generates conformational ensembles by combining five algorithms; useful for disordered proteins and drug discovery. | Methodological framework |
Future Directions and Challenges
Despite its triumphs, the ML revolution continues to confront significant challenges. A primary limitation is the prediction of multiple conformational states. Most models, including AlphaFold2, predict a single, static structure, missing the intrinsic dynamics essential for the function of many proteins, such as enzymes and intrinsically disordered proteins (IDPs) [19]. Emerging ensemble methods like FiveFold, which aggregate predictions from multiple algorithms (AF2, RoseTTAFold, ESMFold, etc.), represent a promising approach to modeling conformational diversity and have shown utility in studying IDPs like alpha-synuclein [19].
Another frontier is the accurate modeling of protein complexes and interactions. While progress is being made, predicting the precise 3D structure of large multi-protein assemblies remains a active area of research. Finally, the "inverse folding" problem, while advanced by tools like ProteinMPNN, is not fully solved. Ensuring that designed sequences are highly designable (i.e., fold reliably into the target structure) and functional requires robust metrics and often iterative experimental validation [18]. The fusion of physics-based principles with deep learning models may hold the key to creating generative models that more accurately characterize the full energy landscape of proteins [18].
The Vast and Constrained Protein Functional Universe
The prediction of a protein's three-dimensional structure from its amino acid sequence stands as a fundamental challenge in structural biology, essential for understanding biological function and accelerating drug discovery. For decades, two distinct computational philosophies have addressed this problem: Evolutionary Algorithms (EA), which leverage physical principles and global optimization to explore conformational space, and Machine Learning (ML), which infers structural patterns from evolutionary information and known protein structures. The recent revolutionary success of deep learning models like AlphaFold2 has dramatically shifted the landscape, establishing a new benchmark for accuracy [9]. However, the core question remains: to what extent can purely physical, search-based methods (EA) compete with or complement data-driven, inference-based methods (ML) in providing accurate, generalizable, and functionally insightful protein models? This review provides a comprehensive benchmarking overview of these competing paradigms, dissecting their methodologies, accuracies, computational demands, and applicability to challenging protein classes like fold-switching proteins and complexes.
Table 1: Core Paradigms in Protein Structure Prediction
| Feature | Evolutionary Algorithm (EA) Approach | Machine Learning (ML) Approach |
|---|---|---|
| Core Philosophy | Physical search-based optimization | Data-driven pattern inference |
| Primary Input | Amino acid sequence & force fields | Amino acid sequence & Multiple Sequence Alignments (MSAs) |
| Representative Method | USPEX [23] | AlphaFold2 [9], ESMFold, OmegaFold [8] |
| Key Strength | Physical realism; potential for novel fold discovery | Unprecedented speed and accuracy for single domains |
| Key Limitation | Computationally intractable for large proteins; force field inaccuracy [23] | Limited by training data; struggles with multiple conformations [24] |
Methodological Deep Dive: Experimental Protocols
The Machine Learning (ML) Pipeline
Modern ML methods, such as AlphaFold2, employ a sophisticated end-to-end neural network architecture. The process begins with input preparation, where the primary amino acid sequence is used to generate a Multiple Sequence Alignment (MSA) and a set of homologous sequences [9]. These are fed into the Evoformer module, a novel neural network block that acts as the system's "engine." The Evoformer processes the inputs through attention-based mechanisms to reason about the spatial and evolutionary relationships between residues, producing a rich representation of the protein's potential structure [9]. This representation is then passed to the structure module, which introduces an explicit 3D structure. Starting from a trivial initial state, this module iteratively refines the atomic coordinates of all heavy atoms through a process called "recycling," resulting in a highly accurate protein structure with precise atomic details [9]. The network is trained end-to-end using a combination of structural losses, including those that emphasize the orientational correctness of residues.
Figure 1: The core workflow of an ML-based protein structure prediction pipeline, as exemplified by AlphaFold2.
The Evolutionary Algorithm (EA) Pipeline
In contrast, the Evolutionary Algorithm approach, as implemented in methods like USPEX, treats structure prediction as a global optimization problem. The algorithm starts with an initial population of random protein conformations. Each structure in this population is then relaxed using molecular mechanics force fields (e.g., Amber, CHARMM, or Oplsaal) via molecular dynamics engines like Tinker or Rosetta to locally minimize its energy [23]. The fitness of each individual in the population is evaluated based on its potential energy or scoring function. A selection process then favors the lowest-energy (fittest) structures to proceed to the next generation. To create new candidate structures, USPEX employs specialized variation operators that generate "offspring" through operations mimicking genetic evolution, such as crossover and mutation. This cycle of selection, variation, and fitness evaluation is repeated for numerous generations, allowing the population to evolve toward conformations with progressively lower energy, ideally converging on the native protein structure [23].
Figure 2: The iterative workflow of an Evolutionary Algorithm (EA) for protein structure prediction.
Benchmarking Performance: Quantitative Comparisons
Accuracy and Reliability Metrics
The performance of protein structure prediction methods is quantitatively assessed using several key metrics. The Global Distance Test (GDT) is a common measure, with a GDT_TS score above 90 generally considered competitive with experimental methods [8]. The predicted Local Distance Difference Test (pLDDT) is a per-residue confidence score where values above 90 indicate high accuracy [9]. For protein complexes, interface-specific metrics like ipTM (interface predicted Template Modeling score) and pDockQ (predicted DockQ score) are used, with higher scores indicating more reliable protein-protein interactions [25].
Table 2: Benchmarking of ML-based Protein Folding Tools [8]
| Protein Length | Method | Running Time (s) | PLDDT | GPU Memory |
|---|---|---|---|---|
| 50 | ESMFold | 1 | 0.84 | 16 GB |
| OmegaFold | 3.66 | 0.86 | 6 GB | |
| AlphaFold (ColabFold) | 45 | 0.89 | 10 GB | |
| 400 | ESMFold | 20 | 0.93 | 18 GB |
| OmegaFold | 110 | 0.76 | 10 GB | |
| AlphaFold (ColabFold) | 210 | 0.82 | 10 GB | |
| 800 | ESMFold | 125 | 0.66 | 20 GB |
| OmegaFold | 1425 | 0.53 | 11 GB | |
| AlphaFold (ColabFold) | 810 | 0.54 | 10 GB |
The benchmarking data reveals a critical trade-off between speed, accuracy, and resource consumption. For shorter sequences (e.g., 50 residues), OmegaFold provides an optimal balance of high accuracy (PLDDT 0.86) and resource efficiency [8]. For longer sequences (e.g., 400 residues), ESMFold demonstrates remarkable speed (20s) and high accuracy (PLDDT 0.93), while AlphaFold remains robust but computationally heavier. In direct performance tests, the EA method USPEX successfully found low-energy conformations for proteins up to 100 residues, with energies comparable to or lower than those generated by the established physical method Rosetta Abinitio [23]. However, the study concluded that current force fields remain a limiting factor for accurate blind prediction via EA.
Performance on Complexes and Alternative Folds
While ML methods excel at predicting single, stable domains, they exhibit significant limitations when proteins adopt multiple conformations. A systematic study found that AlphaFold2 predicts only one conformation for 92% of known dual-folding proteins [24]. This is a critical constraint in the "protein functional universe," as fold-switching proteins are involved in key biological processes like circadian rhythms and transcription regulation [24]. The underlying issue is that standard ML models are trained to output a single, static structure. In contrast, EA methods, by their nature, can sample a diverse landscape of conformations, potentially capturing metastable states. To address this, new methods like Alternative Contact Enhancement (ACE) have been developed, which uncover coevolutionary signatures for both conformations of fold-switching proteins, successfully revealing dual-fold coevolution in 56 out of 56 tested proteins [24].
For protein complexes, AlphaFold3 and ColabFold with templates perform similarly, both outperforming the template-free ColabFold. In assessments of heterodimeric complexes, AlphaFold3 produced the highest fraction of 'high-quality' models (39.8%) and the lowest fraction of 'incorrect' models (19.2%) [25]. The ipTM score and Model Confidence were identified as the most reliable metrics for evaluating these complex predictions [25].
Table 3: Key Resources for Protein Structure Prediction and Validation
| Resource / Reagent | Type | Function / Application |
|---|---|---|
| AlphaFold Database / AlphaFold3 | Software/Web Server | Predicts protein structures and complexes with high accuracy [9] [25] |
| ColabFold | Software | Accessible, cloud-based implementation of AlphaFold2 [25] |
| ESMFold & OmegaFold | Software | Alternative ML tools offering speed/resource advantages [8] |
| USPEX | Software | Evolutionary Algorithm for ab initio protein structure prediction [23] |
| GREMLIN | Software | Infers co-evolved amino acid contacts from MSAs for fold-switching analysis [24] |
| Rosetta (REF2015) | Software Suite | Force field & algorithms for structure prediction & design; used for relaxation & scoring [23] |
| Tinker (Amber/CHARMM) | Software Suite | Molecular dynamics package for structure relaxation & energy calculation [23] |
| GPCRmd, ATLAS | Database | Specialized MD databases for validating dynamics of specific protein families [26] |
| DockQ, pDockQ | Metric | Standardized scores for evaluating quality of protein-protein interfaces [25] |
The benchmarking of Evolutionary Algorithms and Machine Learning for protein folding reveals a nuanced landscape. ML methods, particularly AlphaFold2 and its successors, have achieved unprecedented accuracy for predicting single-domain protein structures, largely solving this aspect of the problem [9] [27]. However, the functional universe of proteins is vast and constrained not by single states but by dynamic conformational landscapes. Here, current ML models show a significant blind spot, often failing to predict functionally critical alternative folds and dynamic conformational changes [24] [26].
EA methods offer a fundamentally different approach based on physical principles and conformational search, proving capable of finding deep energy minima and potentially capturing structural diversity [23]. Their performance, however, is currently limited by computational cost for large proteins and the accuracy of existing force fields. The future of the field lies not in a winner-takes-all outcome but in the integration of both paradigms. ML models can provide powerful starting points and energy surrogates, while EA and physical simulations can be used to refine structures and explore conformational ensembles. Overcoming current limitations will require developing next-generation models that natively predict ensembles, better integrating biophysical constraints into ML, and creating richer training datasets that capture structural diversity, ultimately unlocking a deeper understanding of the vast and constrained protein functional universe.
Protein folding represents one of the most fundamental challenges in computational biology, standing at the intersection of physics, biology, and computer science. The process by which a linear amino acid chain spontaneously folds into a precise three-dimensional structure remains only partially understood, despite decades of research. Two conceptual frameworks—Evolutionary Algorithms (EA) and Machine Learning (ML)—offer distinct approaches to navigating this complex problem space. This technical guide examines the core challenges of combinatorial explosion and evolutionary myopia that constrain both methodologies, providing researchers with experimental protocols, analytical frameworks, and benchmarking data essential for advancing protein folding research.
The protein folding problem is intrinsically linked to astronomical combinatorial complexity. For a typical 100-amino acid protein, the theoretical sequence space encompasses 20^100 possible configurations—a number that exceeds the count of atoms in the observable universe [28]. This combinatorial explosion presents an insurmountable computational barrier for exhaustive search algorithms. Meanwhile, evolutionary myopia describes the limited predictive generalizability of models trained on narrow biological contexts, failing to capture the full diversity of protein structural principles across the tree of life.
Combinatorial Explosion in Protein Folding
The Fundamental Combinatorial Challenge
Combinatorial explosion manifests throughout protein structure prediction and design. The astronomical size of protein sequence spaces makes comprehensive exploration computationally intractable. As Levitt noted in his seminal review, protein folding's "endgame" involves the ordering of amino acid side-chains into a well-defined, closely packed configuration, a process hampered by combinatorial explosion in the number of possible configurations [29]. This challenge is not merely theoretical; it directly impacts the feasibility of computational protein design and structure prediction.
Recent research demonstrates that the genetic architecture of protein stability is remarkably simple despite this combinatorial complexity. Energy models reveal that protein genotypes can be accurately predicted using additive free energy changes with only a small contribution from pairwise energetic couplings [28]. This simplification enables navigation of high-dimensional sequence spaces that would otherwise be computationally prohibitive.
Table 1: Scale of Combinatorial Challenges in Protein Folding
| Aspect | Combinatorial Complexity | Computational Implications |
|---|---|---|
| Sequence Space for 100-aa Protein | 20^100 possible sequences | Exhaustive search impossible; requires heuristic methods |
| Side-chain Packing Configurations | Exponential growth with protein size | Endgame folding requires sophisticated search algorithms [29] |
| Mutational Combinations | 2^34 ≈ 1.7×10^10 for 34 mutation sites | Experimental exploration of high-order mutants extremely challenging [28] |
| Functional Sequence Fraction | <0.2% of 10-aa variants folded (additive model) | Random sampling yields mostly non-functional proteins [28] |
Thermodynamic Frameworks and Dimensional Hardness
A novel thermodynamic theory of intelligence frames combinatorial explosion as the central computational bottleneck in high-dimensional systems. This framework introduces a dimensional hardness parameter (Hd = Γ·τ / C(ρ)·log₂ Deff), where Γ represents entropy flow, τ is the coherence timescale, C(ρ) is the system's coherence, and Deff is the effective dimensionality of the configuration space [30]. Systems maintain structure and adaptivity when Hd <1 but collapse under combinatorial explosion when Hd >1.
This theoretical model has practical implications for protein folding simulations. All-atom molecular dynamics simulations face exponential growth in computational requirements as protein size increases. Recent simulations of protein misfolding reveal that entanglement status changes—where protein sections loop around each other incorrectly—represent a persistent class of misfolding that evades cellular quality control systems [31]. These misfolds are particularly stable and difficult to correct, requiring backtracking and unfolding several steps to correct entanglement status.
Diagram 1: Protein Folding and Misfolding Pathways
Evolutionary Myopia in Biological Systems
Conceptual Framework and Definition
Evolutionary myopia describes the phenomenon where biological systems optimized for immediate fitness advantages develop limitations in long-term adaptability. In protein science, this manifests as limited generalizability of structural principles across phylogenetic boundaries and path dependencies in evolutionary trajectories that constrain future adaptive potential.
This concept finds parallels in human vision research, where myopia development involves complex gene-environment interactions shaped by evolutionary history. Studies of myopia-related genes have detected signatures of adaptation in vision and light perception pathways, with evidence that local adaptation to different light environments during human migration diversified the genetic basis of myopia [32]. This evolutionary specialization potentially contributes to discrepancies in myopia prevalence across modern populations.
Experimental Evidence from Multi-Omics Studies
Integrative transcriptome and proteome analyses of lens-induced myopia in mouse models reveal the molecular basis of this evolutionary mismatch. Researchers identified 175 differentially expressed genes and 646 differentially expressed proteins between treated and control eyes, with insulin-like growth factor 2 mRNA binding protein 1 (Igf2bp1) emerging as a convincing biomarker [33]. The low correlation between transcriptomic and proteomic data highlights the complex regulatory layers between genetic predisposition and phenotypic expression.
Proteomic profiling of form-deprivation myopia in guinea pigs further elucidated 348 differentially expressed proteins in the vitreous body, with calcium signaling pathways playing a critical role in mediating eye changes [34]. These findings demonstrate how evolutionary adaptations to ancient light environments manifest as vulnerabilities under modern conditions.
Table 2: Evolutionary Myopia Signatures in Protein-Related Systems
| System | Evolutionary Adaptation | Modern Vulnerability | Molecular Mechanism |
|---|---|---|---|
| Human Vision | Rhodopsin molecular diversity for different light environments [32] | High myopia prevalence in altered light conditions | Phototransduction pathway genetic variants |
| Protein Fold Stability | Additive energy models with sparse couplings [28] | Misfolding diseases in aging populations | Entanglement errors evading quality control [31] |
| Cellular Quality Control | Efficient degradation of most misfolded proteins | Persistent entanglement misfolds | Buried misfolds invisible to surveillance [31] |
Experimental Methodologies and Benchmarks
High-Dimensional Sequence Space Sampling
Confronting combinatorial explosion requires sophisticated experimental designs that enrich for functional protein sequences. Methodologies for sampling high-dimensional sequence spaces include:
Library Design and Synthesis: Researchers constructed a library containing all combinations of 34 selected mutants (2^34 ≈ 1.7×10^10 genotypes) using a heuristic technique that enriches for conserved fold and function. For each possible starting single amino acid substitution, selections iteratively identified further substitutions that simultaneously maximize the resulting combinatorial mutant's predicted abundance and binding to an interaction partner [28].
AbundancePCA Measurement: Cellular abundance of sampled genotypes was quantified using highly validated pooled selection and abundance protein fragment complementation assays. This approach enabled triplicate abundance measurements for 129,320 variants (0.0007% of sequence space) with high reproducibility (Pearson's r > 0.91) [28].
Energy Model Inference: Additive free energy models were trained on abundance and ligand binding selections quantifying effects of single and double amino acid mutants. Model parameters included Gibbs free energy terms for wild type (ΔGf) and single substitutions (ΔΔGf), with a two-parameter transformation relating folded fraction to AbundancePCA fitness [28].
Diagram 2: High-Throughput Protein Stability Mapping
Classification Benchmarking Frameworks
Large-scale benchmark studies assessing tools for classifying protein-coding and non-coding transcripts reveal systematic challenges in biological sequence analysis. A comprehensive evaluation of 24 tools producing >55 models on 135 datasets identified key bottlenecks [35]:
- Lack of standardized training sets and reliance on homogeneous training data
- Gradual changes in annotated data and absence of gold standards
- Lower performance of end-to-end deep learning models compared to hybrid approaches
- Presence of false positives and negatives in benchmark datasets
These limitations directly impact the assessment of EA versus ML approaches for protein folding. Benchmarking studies must account for dataset bias, with performance metrics contextualized against training data composition and evolutionary distance between training and test cases.
EA vs ML: Comparative Analysis for Protein Folding
Performance Benchmarks and Metrics
The revolutionary success of AlphaFold2 since its 2020 debut demonstrates ML's transformative potential for protein structure prediction [36] [37]. However, evolutionary algorithms maintain distinct advantages for specific protein design challenges. Benchmarking reveals complementary strengths:
Generalization Capability: ML models like AlphaFold achieve remarkable accuracy when predicting structures homologous to training examples but face challenges with entirely novel folds. Evolutionary algorithms employing energy-based scoring functions can explore genuinely novel regions of protein space, albeit at higher computational cost.
Interpretability Trade-offs: EA approaches typically leverage physically interpretable energy models with additive free energy changes and sparse pairwise couplings [28]. In contrast, deep neural networks constitute extremely complicated models with millions of fitted parameters that function as "black boxes" [28].
Data Efficiency: Evolutionary algorithms can navigate high-dimensional sequence spaces with relatively sparse experimental data, as demonstrated by energy models explaining half the fitness variance in combinatorial multi-mutants using only single and double mutant training data [28].
Table 3: EA vs ML Benchmarking for Protein Folding Challenges
| Metric | Evolutionary Algorithms | Machine Learning | Representative Tools |
|---|---|---|---|
| Combinatorial Search | Energy-guided heuristic search | Pattern recognition in known folds | Rosetta, AlphaFold [37] |
| Novel Fold Design | Strong (energy-based exploration) | Limited by training data | – |
| Computational Efficiency | Lower (requires many evaluations) | Higher (after training) | – |
| Experimental Validation Success | 2-8% of 5-aa variants folded [28] | High for structure prediction | AlphaFold (CASP14 winner) [36] |
| Handling Evolutionary Myopia | Physical principles generalize | Limited by training data diversity | – |
Integrated Approaches and Future Directions
The most promising research directions leverage hybrid methodologies that combine physical principles with data-driven pattern recognition. Several integrative strategies show particular promise:
Energy-Based Priors in ML Architectures: Incorporating physicochemical constraints as inductive biases in neural network architectures, combining EA's interpretability with ML's pattern recognition power.
Transfer Learning Across Evolutionary Distance: Using EA-generated synthetic protein families to augment training data for ML models, addressing evolutionary myopia by expanding structural diversity beyond naturally occurring proteins.
Active Learning Frameworks: Iteratively cycling between ML-based predictions and EA-guided experimental validation to rapidly explore high-value regions of sequence space while minimizing experimental burden.
Research Reagent Solutions
Table 4: Essential Research Reagents and Computational Tools
| Reagent/Tool | Function | Application Context |
|---|---|---|
| AbundancePCA | Pooled selection and abundance measurement | High-throughput protein stability quantification [28] |
| 3D-printed lens mounts | Controlled visual form deprivation | Murine myopia induction for evolutionary studies [33] |
| AlphaFold Database | Protein structure predictions | ML-based structure inference benchmark [36] [37] |
| All-atom molecular dynamics | Atomic-scale folding simulation | Protein misfolding mechanism studies [31] |
| RNAChallenge dataset | Standardized classification benchmark | Tool performance evaluation [35] |
| HPLC-EC detection | Neurotransmitter quantification | Dopamine level measurement in myopia studies [33] |
Combinatorial explosion and evolutionary myopia represent fundamental challenges that constrain both evolutionary algorithms and machine learning approaches to protein folding. Combinatorial explosion necessitates sophisticated search strategies and energy-based heuristics to navigate astronomically large sequence spaces. Evolutionary myopia manifests as limited generalizability across evolutionary distances, constraining the predictive power of models trained on narrow biological contexts.
Benchmarking reveals complementary strengths: EA approaches provide physically interpretable models and better novel fold exploration, while ML delivers unprecedented accuracy for structure prediction within its training domain. The most promising research directions integrate these methodologies, combining physical principles with data-driven pattern recognition to overcome both combinatorial explosion and evolutionary myopia.
Future progress will depend on continued development of experimental methods for high-throughput stability mapping, standardized benchmarking datasets that account for evolutionary diversity, and hybrid algorithms that leverage the respective strengths of both evolutionary computation and deep learning. Such integrated approaches offer the greatest potential for unlocking protein folding's remaining mysteries and harnessing this knowledge for therapeutic applications.
From Theory to Practice: Key Algorithms and Transformative Applications in Biopharma
The prevailing paradigm in structural biology has long been that a single amino acid sequence encodes for one stable three-dimensional structure. However, fold-switching proteins challenge this assumption by adopting distinct secondary and tertiary structures, often in response to cellular stimuli [38]. These structural remodelling events play critical biological roles across all kingdoms of life, from regulating the cyanobacterial circadian clock to suppressing human innate immunity during SARS-CoV-2 infection [39] [38]. Despite their biological importance, state-of-the-art deep learning methods like AlphaFold2 systematically fail to predict fold switching, accurately predicting only one conformation for 92% of known dual-folding proteins [39]. This limitation stems from a fundamental challenge: these methods infer structure from evolutionary conservation patterns but appear to miss the coevolutionary signatures specific to alternative folds.
This technical guide explores how Evolutionary Analysis (EA) approaches, specifically Markov Random Fields (MRFs) and the GREMLIN algorithm, address this gap through the novel Alternative Contact Enhancement (ACE) methodology. Unlike machine learning methods that often predict single static structures, ACE successfully revealed coevolution of amino acid pairs corresponding to both conformations in 56 out of 56 tested fold-switching proteins from distinct families [39]. By leveraging evolutionary principles rather than pattern recognition alone, EA provides a powerful complementary approach to ML for predicting protein conformational diversity.
Theoretical Foundation: Evolutionary Analysis for Fold Switching
The Coevolutionary Principle in Protein Structure
The foundation of evolutionary analysis for structure prediction rests on the observation that amino acid pairs that physically interact within a protein structure tend to coevolve over natural selection [40]. When a mutation occurs at one position, compensatory mutations often arise at contacting positions to maintain structural and functional integrity. These evolutionary couplings can be detected through statistical analysis of multiple sequence alignments (MSAs) and used to infer which residues are likely in direct physical contact [39] [40].
Modern implementations use Markov Random Fields (MRFs) to distinguish direct from indirect couplings, addressing the challenge that residues can appear correlated simply because both interact with a third residue [39]. The GREMLIN (Generative Regularized ModeLs of proteINs) algorithm implements an MRF-based approach with several advantages for coevolutionary analysis: it converges to a global minimum as MSA depth increases, generates reasonable predictions from relatively shallow MSAs, and accounts for noncausal correlations through its MRF formalism [39].
Why ML Fails Where EA Succeeds for Dual-Fold Proteins
Machine learning methods like AlphaFold2 rely heavily on the same coevolutionary principle but make different structural assumptions. These systems are trained on static protein structures from the PDB and learn to predict the most thermodynamically stable conformation [41] [42]. For fold-switching proteins, this often results in prediction of only one fold—typically the one with stronger coevolutionary signatures in deep multiple sequence alignments [39].
The key insight behind the ACE approach is that coevolutionary signatures for alternative folds are not absent but are often masked in standard analyses. Single-fold variants within protein superfamilies can dominate the evolutionary signal, drowning out the subtler signatures of fold switching [39]. By strategically analyzing sequence subfamilies with more fold-switching variants, ACE successfully uncovers these hidden coevolutionary patterns.
The ACE Methodology: A Technical Deep Dive
The Alternative Contact Enhancement (ACE) approach employs a sophisticated workflow designed to unmask coevolutionary signals for alternative folds that are typically missed by conventional analyses.
The diagram below illustrates the comprehensive ACE workflow for detecting dual-fold coevolution:
Core Components and Procedures
MSA Generation and Strategic Pruning
The ACE methodology begins by generating a deep multiple sequence alignment using the query sequence known to adopt two distinct folds. Unlike standard approaches that use the deepest possible MSA, ACE strategically prunes this alignment to create successively shallower MSAs with sequences increasingly identical to the query [39]. This systematic pruning creates nested MSAs ranging from diverse superfamilies to specific subfamilies, intentionally unmasking coevolutionary couplings for alternative conformations that are strengthened in specific evolutionary contexts [39].
Dual-Algorithm Coevolutionary Analysis
Each MSA in the nested hierarchy undergoes parallel coevolutionary analysis using two complementary methods:
- GREMLIN: An MRF-based approach that identifies coevolved amino acid pairs through maximum entropy modeling and regularized inference [39]
- MSA Transformer: A language model that uses attention mechanisms to analyze evolutionary patterns both within MSA columns and across individual sequences [39]
This dual-algorithm approach leverages the complementary strengths of both methodologies, with GREMLIN offering robust performance across MSA depths and MSA Transformer sometimes providing superior accuracy for single-fold proteins [39].
Contact Prediction Integration and Filtering
Predictions from all MSAs and both algorithms are combined and superimposed on a single contact map. The contact map uses an asymmetric design to maximize information content, separately displaying contacts unique to each fold [39]. Finally, density-based scanning filters remove noisy predictions while preserving legitimate contacts corresponding to both folds [39].
Contact Categorization Framework
Predicted contacts are systematically categorized into four distinct types:
- Dominant Fold Contacts: Unique contacts corresponding to the experimentally determined structure with greatest overlap to predictions from the deepest MSA
- Alternative Fold Contacts: Unique contacts corresponding to the other experimentally determined structure
- Common Contacts: Predicted contacts overlapping experimentally determined contacts shared by both folds
- Unobserved Contacts: Predicted contacts not overlapping any experimentally determined contacts, which may represent folding intermediates or noise [39]
Table 1: Contact Categorization in ACE Analysis
| Category | Description | Structural Significance |
|---|---|---|
| Dominant Fold Contacts | Unique to the conformation best predicted by deep MSAs | Often but not always the lowest energy state (33% of cases) |
| Alternative Fold Contacts | Unique to the other experimentally determined conformation | Functionally critical alternative state |
| Common Contacts | Shared between both experimentally determined structures | Structural core preserved during fold switching |
| Unobserved Contacts | Not matching any experimental contacts | Potential folding intermediates or prediction errors |
Quantitative Performance Assessment
Enhanced Prediction of Alternative Fold Contacts
The ACE methodology demonstrates substantial improvements over standard coevolutionary analysis approaches that use only deep superfamily MSAs. When applied to 56 fold-switching proteins with sufficiently deep MSAs, ACE achieved mean and median increases of 201% and 187%, respectively, in correctly predicted amino acid contacts uniquely corresponding to alternative conformations [39].
Table 2: Performance Comparison of ACE vs. Standard Approach
| Metric | Standard Approach | ACE Methodology | Improvement |
|---|---|---|---|
| Alternative Fold Contact Prediction | Baseline | 201% mean increase | Substantial enhancement |
| Proteins with Dual-Fold Coevolution | Not detected | 56/56 proteins | 100% success in test set |
| False Positive Rate | Not specified | 0/181 in blind prediction | High specificity |
Validation and Extension to Blind Prediction
The dual-fold coevolution discovered through ACE provides evolutionary evidence that fold-switching has been preserved by natural selection, implying these functionalities provide adaptive advantages [39]. Researchers successfully leveraged ACE-derived contacts to predict two experimentally consistent conformations of a candidate protein with unsolved structure and developed a blind prediction pipeline that correctly identified 13 out of 56 fold-switching proteins (23%) with no false positives (0/181) [39].
Experimental Protocol for ACE Implementation
Step-by-Step Methodology
For researchers seeking to implement the ACE approach, the following detailed protocol provides a practical roadmap:
Input Preparation
- Obtain the amino acid sequence of the protein of interest
- Collect experimentally determined structures for both conformations (if available for validation)
- Format structures with consistent residue numbering
MSA Generation and Processing
Coevolutionary Analysis
- Run GREMLIN analysis on each MSA with default parameters
- Run MSA Transformer on each MSA with attention to both row and column patterns
- Extract top-scoring contacts from each analysis based on coupling scores
Contact Integration and Mapping
- Combine all predicted contacts into unified contact map
- Use asymmetric representation to separate fold-specific contacts
- Map predictions to experimental structures using 8Å heavy atom distance cutoff [39]
Density-Based Filtering
- Implement scanning window approach to identify high-density contact regions
- Filter out low-density predictions likely to be noise
- Retain contacts forming spatially proximate clusters
Validation and Classification
- Categorize contacts as dominant, alternative, common, or unobserved
- Quantify overlap with experimental structures
- Calculate performance metrics for each fold separately
Research Reagent Solutions
Table 3: Essential Research Reagents and Computational Tools for ACE Implementation
| Resource | Type | Function in ACE Protocol | Availability |
|---|---|---|---|
| GREMLIN | Algorithm | MRF-based coevolutionary analysis | Publicly available |
| MSA Transformer | Algorithm | Language model-based contact prediction | Publicly available |
| MMSeq2 | Software | Rapid MSA generation | Publicly available |
| ColabFold | Platform | Integrated MSA generation and structure prediction | Publicly available [40] |
| Protein Data Bank | Database | Experimental structures for validation | Publicly available |
| AlphaFold Database | Database | Structural predictions for comparison | Publicly available [40] |
Comparative Advantages in the EA vs. ML Landscape
When benchmarking Evolutionary Analysis against Machine Learning approaches for protein structure prediction, each methodology demonstrates distinct strengths and limitations:
EA approaches, particularly the ACE methodology, excel where ML methods face fundamental challenges:
- Detection of multiple native states from single sequences [39]
- Identification of coevolutionary patterns specific to subfamilies [39]
- Revealing evolutionary selection for dual-fold functionality [39]
- Blind prediction of fold-switching capability without prior structural knowledge [39]
ML approaches maintain advantages in:
- Speed and scalability for proteome-wide prediction [40]
- Accuracy for single-fold globular proteins [19]
- Accessibility through user-friendly interfaces [42]
- Integration of multiple data types through end-to-end architectures [40]
The most powerful future framework likely combines strengths of both approaches, using EA principles to guide ML models beyond single-structure predictions toward conformational ensembles and dynamic landscapes [42] [19].
Future Directions and Integration Opportunities
The demonstrated success of ACE for identifying dual-fold coevolution suggests several promising research directions:
- Integration with ensemble prediction methods like FiveFold, which combines predictions from five algorithms to model conformational diversity [19]
- Hybrid EA-ML pipelines that use ACE-derived contacts as constraints for deep learning models
- Extension to condition-dependent folding by analyzing MSAs from specific environmental contexts
- Application to drug discovery for targeting alternative conformations in therapeutic development [19]
As the field progresses, the integration of evolutionary analysis with machine learning represents the most promising path toward comprehensively understanding and predicting protein structural diversity, moving beyond single static structures to capture the dynamic reality of proteins in their native biological environments [41].
The prediction of protein three-dimensional structures from amino acid sequences represents a monumental challenge in computational biology. For decades, this "protein folding problem" remained largely unsolved, bottlenecking advancements in fields ranging from drug discovery to fundamental biology. The landscape transformed dramatically with the advent of sophisticated machine learning (ML) methods, particularly deep learning architectures that have achieved unprecedented accuracy. These ML approaches now stand in contrast to earlier methodologies that heavily relied on evolutionary analysis (EA) through multiple sequence alignments (MSAs) and physical energy functions.
This technical guide provides an in-depth analysis of three leading ML powerhouses in protein structure prediction: AlphaFold, ESMFold, and OmegaFold. Each system embodies a distinct architectural philosophy in addressing the folding problem, with varying dependencies on evolutionary information and computational demands. Understanding these core architectures is essential for researchers, scientists, and drug development professionals seeking to leverage these tools effectively and contextualize their performance within the broader paradigm shift from EA-driven to ML-driven folding approaches.
Quantitative Performance Benchmarking
A comprehensive benchmark comparing the three methods reveals critical trade-offs between accuracy, speed, and computational resource requirements, enabling informed selection based on research constraints and objectives.
Table 1: Runtime and Resource Consumption Benchmark (A10 GPU)
| Sequence Length | Method | Running Time (s) | PLDDT | CPU Memory (GB) | GPU Memory (GB) |
|---|---|---|---|---|---|
| 50 | ESMFold | 1 | 0.84 | 13 | 16 |
| OmegaFold | 3.66 | 0.86 | 10 | 6 | |
| AlphaFold* | 45 | 0.89 | 10 | 10 | |
| 400 | ESMFold | 20 | 0.93 | 13 | 18 |
| OmegaFold | 110 | 0.76 | 10 | 10 | |
| AlphaFold* | 210 | 0.82 | 10 | 10 | |
| 800 | ESMFold | 125 | 0.66 | 13 | 20 |
| OmegaFold | 1425 | 0.53 | 10 | 11 | |
| AlphaFold* | 810 | 0.54 | 10 | 10 | |
| 1600 | ESMFold | Failed (OOM) | - | - | 24 |
| OmegaFold | Failed (>6000) | - | - | 17 | |
| AlphaFold* | 2800 | 0.41 | 10 | 10 |
Note: AlphaFold data is based on ColabFold implementation. PLDDT (Predicted Local Distance Difference Test) scores range from 0-1, with higher values indicating greater confidence/accuracy. OOM = Out of Memory. [8]
Table 2: Method Overview and Comparative Strengths
| Method | Developer | Core Innovation | MSA-Dependent | Key Strength | Key Limitation |
|---|---|---|---|---|---|
| AlphaFold | DeepMind | Evoformer & End-to-End Learning | Yes | Exceptional accuracy, especially with MSA | Computationally intensive, complex setup |
| ESMFold | Meta | Single-Sequence Protein Language Model | No | Extreme speed, no MSA needed | Lower accuracy on some targets, high memory use |
| OmegaFold | Various Academics | Protein Language Model & Geometric Transformers | No | Balance of accuracy and MSA-independence | Slower than ESMFold, struggles with long sequences |
The benchmarking data indicates that ESMFold provides the fastest inference for shorter sequences but faces memory constraints with longer proteins. [8] OmegaFold demonstrates superior accuracy on shorter sequences compared to ESMFold while maintaining reasonable resource utilization, making it suitable for resource-constrained environments. [8] AlphaFold achieves the highest accuracy across diverse targets, particularly when reliable MSAs are available, albeit with significantly longer runtimes. [9] [8]
Architectural Breakdown
AlphaFold: End-to-End Geometric Deep Learning
AlphaFold employs a sophisticated, integrated architecture that directly predicts atomic coordinates from sequence data, representing a significant departure from earlier fragment-assembly or physical simulation approaches. [9]
Diagram 1: AlphaFold's Core Architecture with Recycling
The network processes two primary representations throughout its architecture: a pair representation (Nres × Nres) encoding relationships between residues, and an MSA representation (Nseq × Nres) capturing evolutionary information. [9] [43] The revolutionary Evoformer block, the core of AlphaFold's architecture, enables continuous information exchange between these representations through novel operations: [9]
- Triangle Multiplicative Updates: Enforce geometric consistency by using two edges of a triangle to update the third, ensuring physical plausibility.
- Axial Attention Mechanisms: Process rows and columns of the MSA representation efficiently while incorporating pair-level information as biases.
The processed representations feed into the Structure Module, which explicitly represents 3D atomic coordinates as rigid body frames (rotations and translations) for each residue. [9] AlphaFold implements iterative refinement through a "recycling" process where outputs are recursively fed back into the same modules, progressively enhancing accuracy while reducing stereochemical violations. [9] [44]
AlphaFold 3 extends this architecture with the Pairformer (a simplified Evoformer) and a diffusion-based structure decoder that begins with a cloud of atoms and iteratively refines their positions, enabling prediction of complexes involving proteins, DNA, RNA, and small molecules. [44] [43]
ESMFold: Protein Language Model Paradigm
ESMFold represents a fundamentally different approach that leverages unsupervised learning on protein sequences alone, eliminating the computational bottleneck of MSAs.
Diagram 2: ESMFold's Single-Sequence Language Model Approach
The architecture begins with ESM-2, a 15-billion parameter transformer model pre-trained using masked language modeling on millions of protein sequences from UniRef. [45] During this pre-training, the model develops attention patterns that implicitly capture structural interactions between amino acids, effectively internalizing structural constraints from evolutionary patterns without explicit supervision. [45]
These learned representations are passed to a folding block that processes both sequence and pairwise representations, similar to AlphaFold but substantially simplified. [45] Finally, an equivariant transformer converts these representations into precise atomic-level coordinates while maintaining rotational and translational equivariance—a critical property for meaningful structural predictions. [45]
This streamlined architecture enables ESMFold to achieve speeds approximately 60 times faster than AlphaFold, allowing Meta to predict structures for over 600 million metagenomic proteins, though with generally lower accuracy than AlphaFold on challenging targets. [45]
OmegaFold: Geometric Transformers without MSAs
OmegaFold occupies a middle ground, combining protein language model principles with explicit structural reasoning while operating independently of MSAs.
OmegaFold introduces a novel combination of a protein language model and a geometry-inspired transformer to achieve high-resolution structure prediction. [46] [47] Unlike ESMFold, OmegaFold generates pseudo-MSAs internally through its language model, capturing co-evolutionary patterns without external database searches, making it particularly effective for orphan sequences with few homologs. [48] [47]
The architecture employs attention-based geometric transformers that explicitly reason about spatial relationships and protein geometry during the folding process. [47] This approach demonstrates particular strength on shorter protein sequences (up to 400 residues), where it achieves superior accuracy compared to ESMFold and competitive performance with AlphaFold, while maintaining greater computational efficiency than the latter. [8]
Experimental Methodology
Training Protocols
Table 3: Training Data and Objectives
| Method | Training Data | Training Objective | Key Architectural Innovations |
|---|---|---|---|
| AlphaFold | 170,000+ PDB structures; evolutionary databases | End-to-end coordinate prediction with intermediate losses | Evoformer, Triangle multiplicative updates, Iterative recycling |
| ESMFold | Millions of sequences from UniRef; no structural data in pre-training | Masked language modeling followed by structural fine-tuning | Emergent attention maps from language modeling, Equivariant transformers |
| OmegaFold | Curated protein structures; evolutionary sequences | Structure prediction with geometric constraints | Protein language model pre-training, Attention-based geometric transformers |
Validation and Benchmarking
Rigorous validation protocols are essential for meaningful comparison of protein folding methods. The Critical Assessment of Structure Prediction (CASP) experiments serve as the gold-standard blind assessment for protein folding accuracy. [9] [44] In CASP14, AlphaFold achieved a median backbone accuracy of 0.96 Å RMSD95, dramatically outperforming other methods (next best: 2.8 Å), with accuracy competitive with experimental structures in most cases. [9]
Standardized evaluation metrics include:
- GDT (Global Distance Test): Measures structural similarity, with scores above ~90 considered competitive with experimental methods. [44]
- pLDDT (predicted Local Distance Difference Test): Per-residue confidence estimate ranging from 0-100. [9] [8]
- TM-score: Structure similarity measure that is less sensitive to local variations. [9]
For comprehensive benchmarking, tools like PSBench provide large-scale datasets with over one million structural models annotated with multiple quality scores at global, local, and interface levels, enabling rigorous EMA (Estimation of Model Accuracy) comparisons. [49]
Diagram 3: Protein Structure Prediction Benchmarking Protocol
Research Reagent Solutions
Table 4: Essential Resources for Protein Structure Prediction Research
| Resource Category | Specific Tools | Function & Application |
|---|---|---|
| Structure Prediction Servers | AlphaFold Server, COSMIC² (OmegaFold) | Web-based interfaces for structure prediction without local installation |
| Local Implementation Frameworks | ColabFold, OpenFold, OmegaFold (NIH Biowulf) | Optimized implementations for local deployment, often with reduced hardware requirements |
| Benchmarking & Validation Suites | PSBench, CASP Assessment Tools | Large-scale datasets and evaluation pipelines for rigorous method comparison |
| Specialized Computing Resources | NVIDIA A100/A10 GPUs, High-CPU servers | Hardware acceleration for training and inference of large models |
| Biological Databases | Protein Data Bank (PDB), UniRef, MGnify | Source databases for training data, templates, and multiple sequence alignments |
| Structure Analysis Tools | PyMOL, ChimeraX, VMD | Visualization and analysis of predicted protein structures |
The architectural evolution from AlphaFold to ESMFold and OmegaFold represents a fascinating trajectory in computational biology. AlphaFold's sophisticated, EA-integrated approach demonstrates the peak of accuracy achievable through carefully engineered deep learning architectures that explicitly incorporate evolutionary and physical constraints. ESMFold's protein language model paradigm showcases the emergent structural understanding possible through scaling unsupervised learning on sequences alone, prioritizing speed and scalability. OmegaFold strikes a balance, maintaining independence from MSAs while incorporating explicit geometric reasoning.
For the research and drug development professional, selection criteria should be guided by specific use cases: AlphaFold for maximum accuracy when computational resources and MSA information are available; ESMFold for high-throughput screening of large sequence databases; and OmegaFold for orphan sequences or when operating under computational constraints. As these methods continue to evolve, the integration of their complementary strengths will likely define the next generation of protein structure prediction tools, further closing the gap between computational prediction and experimental determination in structural biology.
The field of de novo protein design seeks to create novel proteins with specified structural and functional properties from scratch, rather than modifying existing natural proteins. This represents a paradigm shift, moving beyond the constraints of natural evolutionary history to access a vastly larger protein functional universe [16]. Recent breakthroughs in artificial intelligence (AI), particularly generative models, have dramatically accelerated our ability to design proteins computationally. These advancements are primarily driven by two complementary classes of technologies: structure-based diffusion models like RFdiffusion, which generate protein backbone structures, and Protein Language Models (PLMs), which understand and generate protein sequences based on evolutionary principles [18] [50].
This technical guide provides an in-depth examination of these core technologies, their methodologies, and their integration. Framed within the context of benchmarking evolutionary algorithm (EA) versus machine learning (ML) approaches for protein folding and design, it details how modern AI tools are enabling the systematic exploration of protein sequence and structure space. The guide is structured to equip researchers and drug development professionals with a comprehensive understanding of the current state-of-the-art, its experimental validation, and the practical tools required for implementation.
Core Technologies and Mechanisms
RFdiffusion: Structure-Based Generative Modeling
RFdiffusion is a generative model for protein backbones based on a denoising diffusion probabilistic model (DDPM) framework. It was developed by fine-tuning the RoseTTAFold structure prediction network on protein structure denoising tasks [51].
Architectural Foundation and Representation
The model utilizes the RoseTTAFold architecture, which operates on a residue-based frame representation comprising:
- A Cα coordinate
- An N–Cα–C rigid orientation for each residue [51]
This representation is rotationally equivariant, meaning predictions are independent of the global orientation of the input structure, a crucial property for working with 3D molecular data.
Diffusion and Denoising Process
The core generative process involves a forward noising process and a learned reverse denoising process:
- Forward Noising: Training inputs are generated by perturbing native protein structures from the PDB with Gaussian noise for up to 200 steps. For translations, Cα coordinates are perturbed with 3D Gaussian noise. For residue orientations, the method uses Brownian motion on the manifold of rotation matrices [51].
- Reverse Denoising: To generate a novel protein backbone, the process starts from random noise. RFdiffusion iteratively predicts a denoised structure from the noisy input. Each residue frame is updated by stepping toward this prediction with controlled noise addition, progressively refining the structure over multiple steps [51].
Table 1: RFdiffusion Training and Architectural Specifications
| Component | Specification | Functional Significance |
|---|---|---|
| Base Network | Fine-tuned RoseTTAFold | Leverages pre-learned structural knowledge from protein folding |
| Training Loss | Mean-squared error (m.s.e.) between frame predictions and true structure | Promotes continuity of global coordinate frame between timesteps |
| Training Strategy | Self-conditioning | Conditions on previous predictions between timesteps; improves performance |
| Output | Protein backbone structure (Cα atoms and orientations) | Provides scaffold for subsequent sequence design |
Conditioning for Functional Design
A key advantage of RFdiffusion is its ability to incorporate conditioning information during the generation process, enabling solutions to targeted design challenges. Conditioning types include [51]:
- Partial sequence information
- Fold information (e.g., secondary structure and block-adjacency)
- Fixed functional-motif coordinates (e.g., for enzyme active sites or binding interfaces)
Diagram 1: RFdiffusion Generation Workflow
Protein Language Models (PLMs): Sequence-Based Generative Modeling
Protein Language Models represent a complementary approach that treats protein sequences as textual data, applying natural language processing techniques to learn the underlying "grammar" and "syntax" of proteins.
Architectural Approaches and Training
PLMs are typically based on transformer or other deep learning architectures and are trained on massive datasets of protein sequences, such as UniRef or the MGnify Protein Database, which contains nearly 2.4 billion non-redundant sequences [16]. Training methodologies include:
- Unsupervised Learning: Models learn general features from large, diverse datasets of unlabeled protein sequences [52].
- Weak-Positive Only Learning: Utilizes limited sets of evolutionarily related proteins that lack experimentally assayed fitness labels [52].
- Supervised Learning: Trained on variant sequences of a specific target protein with associated functional labels [52].
Functional Applications
PLMs can be deployed for various protein design tasks:
- Conditional Sequence Generation: Creating sequences based on structural constraints, functional descriptions, or control tags [53].
- Inverse Folding: Generating sequences that are likely to fold into a given backbone structure, with models like ProteinMPNN achieving remarkable success in designing sequences for monomeric proteins, binders, and oligomers [18].
- Function-Guided Design: Generating novel proteins based on textual functional descriptions or functional keywords [53].
Quantitative Performance and Benchmarking
Performance Metrics for Protein Design
Evaluating de novo protein design methods requires multiple metrics to assess different aspects of design quality. The PDFBench benchmark introduces a comprehensive set of 22 metrics covering [53] [54]:
- Sequence Plausibility: Measures how protein-like the generated sequences are.
- Structural Fidelity: Assesses whether the designed protein folds into the intended structure.
- Language-Protein Alignment: Evaluates how well the generated protein matches the input functional description or keywords.
- Novelty and Diversity: Quantifies how distinct the designs are from naturally occurring proteins.
RFdiffusion Performance Data
RFdiffusion has demonstrated state-of-the-art performance across multiple design challenges:
Table 2: RFdiffusion Experimental Performance Metrics
| Design Challenge | In Silico Success Rate | Experimental Validation | Key Achievement |
|---|---|---|---|
| Unconditional Monomer Design | High diversity and accuracy | 6/300-residue and 3/200-residue designs characterized; correct topology and high thermostability [51] | Generates elaborate structures with little similarity to training data |
| Protein Binder Design | Confirmed by experimental structures | Cryo-EM structure of designed binder with influenza haemagglutinin nearly identical to design model [51] | High accuracy in interface design |
| Symmetric Oligomer Design | High success rate in silico | Hundreds of designed symmetric assemblies characterized [51] | Enables complex supramolecular structures |
| Metal-Binding Proteins | Not explicitly quantified | Hundreds of metal-binding proteins experimentally characterized [51] | Accurate scaffolding of functional sites |
Comparative Performance in Function-Guided Design
The PDFBench benchmark provides comparative data for various models on function-guided design tasks. Performance varies significantly across models and evaluation metrics, highlighting the importance of multi-faceted benchmarking [53].
Integrated Experimental Protocols
RFdiffusion and ProteinMPNN Workflow for De Novo Design
The most successful current protocol combines RFdiffusion for structure generation with ProteinMPNN for sequence design, following a two-stage paradigm [51] [53]:
Stage 1: Backbone Generation with RFdiffusion
- Define Design Objective: Specify whether the goal is unconditional generation, motif scaffolding, binder design, or symmetric assembly.
- Configure Conditioning: If applicable, provide conditioning information such as fixed motif coordinates, symmetry operations, or target interface residues.
- Generate Backbones: Run the RFdiffusion sampling process to produce candidate backbone structures. For challenging design problems, generate thousands of candidates.
- Filter Backbones: Select a subset of backbones based on structural criteria (e.g., compactness, secondary structure composition, similarity to design objective).
Stage 2: Sequence Design with ProteinMPNN
- Input Backbone Structures: Provide the selected backbones to ProteinMPNN.
- Generate Sequences: Sample multiple sequences (typically 8 per design) for each backbone structure [51].
- Filter Sequences: Select sequences based on confidence metrics, evolutionary novelty, and other relevant criteria.
Stage 3: In Silico Validation
- Structure Prediction: Use AlphaFold2 or ESMFold to predict the structure of each designed sequence.
- Success Criteria: Define in silico success as:
- High confidence (mean pAE < 5)
- Global backbone RMSD < 2 Å to the designed structure
- Local backbone RMSD < 1 Å on any scaffolded functional site [51]
- Select Candidates: Choose top candidates for experimental characterization.
Experimental Validation Protocol
After computational design, experimental validation is essential:
- Gene Synthesis: Chemically synthesize DNA sequences encoding the designed proteins.
- Protein Expression: Express proteins in appropriate expression systems (e.g., E. coli, insect cells).
- Purification: Purify proteins using standard chromatography techniques.
- Biophysical Characterization:
Diagram 2: Protein Design and Validation Pipeline
Table 3: Key Computational and Experimental Resources for AI-Driven Protein Design
| Resource Type | Specific Tools/Databases | Primary Function | Access |
|---|---|---|---|
| Structure Prediction | AlphaFold2, RoseTTAFold, ESMFold | Predict 3D structure from amino acid sequence | Open source / Web servers |
| Structure Databases | Protein Data Bank (PDB), AlphaFold DB, ESM Metagenomic Atlas | Provide experimental and predicted structures for training and analysis | Public databases |
| Sequence Design | ProteinMPNN, ESM-IF, EvoDiff | Generate sequences for given backbone structures ("inverse folding") | Open source |
| Backbone Generation | RFdiffusion, AF2-design, Proteus | Generate novel protein backbone structures de novo | Open source |
| Benchmarking | PDFBench, CASP | Standardized evaluation of protein design and prediction methods | Public benchmarks |
| Experimental Validation | X-ray crystallography, Cryo-EM, Circular Dichroism | Confirm structural accuracy and stability of designs | Core facilities |
Generative AI methods, particularly RFdiffusion and Protein Language Models, have dramatically advanced the field of de novo protein design. By combining structure-based diffusion with sequence-based language modeling, researchers can now design novel proteins with specified folds and functions at an unprecedented success rate—approaching 20% experimental success rates for some applications [50]. These tools enable the exploration of regions in protein sequence and structure space that natural evolution has not sampled, potentially unlocking new solutions for therapeutic, catalytic, and synthetic biology challenges.
The integration of these computational methods with robust experimental validation, as exemplified by the RFdiffusion and ProteinMPNN pipeline, represents the current state-of-the-art. As benchmarking frameworks like PDFBench continue to standardize evaluation, and as models incorporate more biochemical knowledge, we can anticipate further acceleration in our ability to design functional proteins de novo, ultimately expanding access to the vast untapped potential of the protein functional universe.
AI-Enabled Molecular Docking for Drug Discovery
Molecular docking, a cornerstone of computational drug design, is undergoing a transformative shift from traditional physics-based simulations to artificial intelligence-driven methodologies. This paradigm shift addresses critical bottlenecks in conventional drug discovery, where prolonged timelines, substantial costs, and inherent uncertainties impede development workflows [56]. AI-enabled docking leverages deep learning models to directly predict protein-ligand binding conformations and associated binding free energies, bypassing computationally intensive conformational searches through advanced parallel computing capabilities [56]. This technical guide provides an in-depth analysis of current AI docking methodologies, their performance benchmarks, experimental protocols, and their contextual relationship to broader machine learning advances in protein structure prediction, fulfilling a critical need for researchers and drug development professionals navigating this rapidly evolving landscape.
Methodological Landscape: AI Docking Architectures
The current ecosystem of AI-enabled molecular docking encompasses three primary architectural paradigms, each with distinct mechanistic approaches and performance characteristics that researchers must understand for proper implementation.
Generative Diffusion Models
Generative diffusion models represent the most recent innovation in docking methodology, operating through a progressive denoising process that refines random initial ligand poses into precise binding conformations [56]. These models, including SurfDock and DiffBindFR, demonstrate exceptional pose prediction accuracy, with SurfDock achieving remarkable RMSD ≤ 2Å success rates of 91.76% on benchmark datasets like the Astex diverse set [56]. The underlying architecture operates through a forward process that gradually adds noise to known crystal structures, training the model to learn the reverse transformation that recovers native poses from noise. During inference, the model samples from a noise distribution and iteratively refines the pose through a learned denoising function, effectively navigating the complex conformational space to identify energetically favorable binding geometries.
Regression-Based Models
Regression-based architectures, including KarmaDock and QuickBind, employ deep neural networks to directly map input features of protein binding pockets and ligand structures to either binding affinity values or atomic coordinates of the bound pose [56]. These models typically utilize graph neural networks (GNNs) or transformer-based architectures to process structural information, learning complex patterns from vast datasets of known protein-ligand complexes [57]. While offering computational efficiency, regression models frequently struggle with physical plausibility, often producing chemically invalid structures with incorrect bond lengths, angles, or steric clashes despite favorable RMSD metrics [56]. This limitation stems from their direct coordinate prediction approach without explicit enforcement of molecular mechanics constraints.
Hybrid AI-Traditional Methods
Hybrid methodologies, exemplified by Interformer, integrate AI-driven scoring functions with traditional conformational search algorithms [56]. These approaches leverage the sampling capabilities of physics-based docking engines like AutoDock Vina while enhancing pose ranking through learned scoring functions trained on structural data. The hybrid paradigm offers a balanced approach, maintaining the physical validity advantages of traditional methods while incorporating the pattern recognition capabilities of deep learning. This architecture typically demonstrates superior performance in virtual screening scenarios where both binding pose accuracy and affinity prediction are crucial for hit identification [56].
Performance Benchmarking: Quantitative Analysis
Comprehensive evaluation across multiple dimensions reveals the distinct performance characteristics of each docking methodology, providing critical insights for method selection in specific research contexts.
Table 1: Docking Performance Across Method Classes (Success Rates %)
| Method Category | Representative Methods | Pose Accuracy (RMSD ≤ 2Å) | Physical Validity (PB-Valid) | Combined Success (RMSD ≤ 2Å & PB-Valid) |
|---|---|---|---|---|
| Traditional | Glide SP, AutoDock Vina | 65.29-77.65 | 94.12-97.65 | 63.53-75.88 |
| Generative Diffusion | SurfDock, DiffBindFR | 75.66-91.76 | 40.21-63.53 | 33.33-61.18 |
| Regression-Based | KarmaDock, QuickBind | 15.38-42.35 | 10.96-47.79 | 3.27-23.28 |
| Hybrid AI | Interformer | 55.88-77.04 | 85.29-94.12 | 51.76-72.35 |
Table 2: Performance Across Dataset Difficulties (Success Rates %)
| Method Category | Astex Diverse Set (Known Complexes) | PoseBusters Set (Unseen Complexes) | DockGen Set (Novel Pockets) |
|---|---|---|---|
| Traditional | 75.88 | 70.59 | 63.53 |
| Generative Diffusion | 61.18 | 39.25 | 33.33 |
| Regression-Based | 23.28 | 15.38 | 3.27 |
| Hybrid AI | 72.35 | 64.12 | 51.76 |
The performance stratification clearly demonstrates that traditional and hybrid methods maintain superior physical validity and combined success rates across all dataset difficulties, while generative models excel specifically in pose accuracy for known complexes but struggle with novel targets [56]. This performance pattern highlights a critical generalization challenge in current AI docking methods, particularly when encountering proteins with low sequence similarity to training data or novel binding pocket architectures.
Experimental Protocols: Benchmarking Methodology
Rigorous evaluation protocols are essential for meaningful performance assessment and method comparison. The following standardized methodology ensures reproducible benchmarking across different research environments.
Dataset Curation and Preparation
- Astex Diverse Set: Curate a collection of known protein-ligand complexes with high-resolution crystal structures (≤2.0Å), ensuring diverse protein families and ligand chemotypes [56].
- PoseBusters Benchmark: Prepare complexes strictly excluded from training datasets to evaluate generalization to unseen targets, with particular attention to complex binding motifs and challenging steric environments [56].
- DockGen Dataset: Compile structures featuring novel binding pockets with low structural similarity to proteins in common training sets, focusing on orphan targets with therapeutic relevance [56].
- Pre-processing: Standardize all protein structures by removing water molecules, adding hydrogen atoms, and assigning appropriate protonation states using tools like OpenBabel or RDKit. Prepare ligand structures through geometry optimization and tautomer enumeration.
Evaluation Metrics and Implementation
- Pose Accuracy: Calculate root-mean-square deviation (RMSD) of heavy atoms between predicted and experimental ligand poses after optimal structural alignment of protein binding sites. Apply a 2Å threshold for success classification [56].
- Physical Validity: Utilize PoseBusters validation toolkit to assess chemical and geometric consistency, including bond length validity (within 4σ of standard values), bond angle validity (within 4σ of standard values), stereochemistry preservation, absence of protein-ligand atomic clashes, and proper ring conformation [56].
- Interaction Recovery: Quantify recovery of key molecular interactions (hydrogen bonds, halogen bonds, π-π stacking, salt bridges) through computational geometry approaches comparing predicted poses to experimental reference structures.
- Virtual Screening Performance: Evaluate using enrichment factors (EF1, EF10) and area under the ROC curve (AUC) in ligand benchmarking experiments with known active and decoy compounds [56].
Execution Parameters
- Computational Environment: Conduct experiments on systems with high-performance GPUs (e.g., NVIDIA A100 or H100) to ensure consistent inference times across methods.
- Sampling Parameters: For methods with configurable sampling, use consistent exhaustiveness parameters (e.g., 32 for Vina-based methods) and generate 20 poses per ligand to ensure adequate conformational coverage.
- Scoring and Ranking: Employ each method's native scoring function for pose ranking without post-processing to evaluate real-world performance.
Diagram 1: Docking benchmarking workflow (76 characters)
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of AI-enabled molecular docking requires familiarity with both computational tools and experimental validation methodologies.
Table 3: Essential Research Reagents and Computational Tools
| Tool/Reagent | Type | Primary Function | Application Context |
|---|---|---|---|
| SurfDock | Software | Generative diffusion model for pose prediction | High-accuracy binding pose generation for well-characterized targets |
| Glide SP | Software | Traditional physics-based docking with hybrid AI scoring | Virtual screening campaigns requiring high physical validity |
| PoseBusters | Validation Toolkit | Automated physical plausibility assessment | Quality control for docking predictions pre-experimental validation |
| CETSA | Experimental Assay | Cellular target engagement validation in intact cells | Confirmation of binding predictions in physiologically relevant environments |
| AlphaFold3 | Structure Prediction | Protein-ligand complex structure prediction | Generating structural templates for targets lacking experimental structures |
| AutoDock Vina | Software | Traditional conformational search algorithm | Baseline comparisons and hybrid method implementations |
| Interformer | Software | Hybrid AI-traditional docking method | Balanced approach for both pose accuracy and physical validity |
Contextual Framework: Docking in Protein Structure Prediction
AI-enabled molecular docking represents a critical downstream application within the broader ecosystem of machine learning advances in structural biology, particularly following revolutionary developments in protein folding prediction. The relationship between these domains is both sequential and synergistic, with accurate protein structure prediction serving as a foundational prerequisite for reliable molecular docking [49].
The benchmarking paradigms established for protein folding methods, including those evaluating ESMFold, OmegaFold, and AlphaFold, provide valuable methodological frameworks for assessing AI docking approaches [8]. These include standardized metrics like PLDDT (predicted local distance difference test) for folding accuracy that parallel docking assessment through RMSD and physical validity checks [8]. Furthermore, the computational resource considerations documented in protein folding benchmarking—including running time, CPU memory, and GPU memory utilization—directly inform infrastructure requirements for deploying AI docking solutions in research and production environments [8].
The evolutionary trajectory from traditional molecular dynamics simulations to AI-powered folding prediction mirrors the ongoing transition in docking methodologies, with both fields grappling with balancing accuracy, computational efficiency, and physical plausibility [58]. This contextual relationship underscores the importance of considering AI-enabled molecular docking not as an isolated technological advancement, but as an integral component of the computational structural biology toolkit, increasingly essential for accelerating drug discovery pipelines and improving therapeutic development success rates [59].
Diagram 2: AI docking in structure prediction (67 characters)
The comprehensive benchmarking of AI-enabled molecular docking methods reveals distinct performance patterns that should guide methodological selection for specific research applications. For virtual screening campaigns prioritizing hit identification, hybrid AI-traditional methods provide the optimal balance of computational efficiency and physical validity. For binding mode analysis of lead compounds with established activity, generative diffusion models offer superior pose accuracy, though require experimental validation to address physical plausibility limitations. Regression-based approaches currently serve primarily as rapid screening tools for large compound libraries despite their physical validity challenges. As AI methodologies continue to evolve, integration with experimental validation through techniques like CETSA for cellular target engagement confirmation remains essential for bridging the gap between computational prediction and biomedical reality, ultimately accelerating therapeutic development through robust, AI-enabled structure-based drug design.
Engineering Novel Enzymes and Antibodies for Synthetic Biology
The field of synthetic biology is undergoing a profound transformation, moving away from traditional, labor-intensive protein engineering methods toward computationally driven design. For decades, engineering novel enzymes and antibodies relied heavily on directed evolution—an iterative process of random mutagenesis and screening—and rational design, which required extensive structural knowledge [60] [61]. These methods, while successful, were often described as Sisyphean tasks due to the practically immeasurable size of protein sequence space, where a typical-length protein can fold into 10^300 possible configurations [60]. The advent of artificial intelligence (AI) and machine learning (ML) has fundamentally shifted this paradigm. AI systems, notably DeepMind's AlphaFold which earned the 2024 Nobel Prize in Chemistry, have revolutionized structure prediction [60] [41]. Furthermore, the rise of generative AI models and inverse folding approaches has flipped the traditional script, enabling researchers to design novel protein sequences for desired structures and functions from scratch, thereby accelerating the development of biocatalysts and therapeutics for synthetic biology applications [60] [42].
AI and Machine Learning Foundations for Protein Design
Key Computational Models and Their Applications
The integration of AI into protein engineering has been catalyzed by several foundational computational models that address different aspects of the design problem. These tools have evolved from predicting static structures to designing functional biomolecules and modeling their dynamic interactions.
Table 1: Key AI Models in Protein Engineering and Their Primary Applications
| Model Name | Type | Primary Application in Synthetic Biology | Key Advancement |
|---|---|---|---|
| AlphaFold 2 & 3 [60] [42] | Structure Prediction Neural Network | Predicts 3D protein structures and multi-molecular complexes (proteins, DNA, RNA, ligands). | Achieved atomic accuracy in structure prediction; AF3 extends to biomolecular complexes. |
| RFdiffusion [60] [42] | Generative Design (Diffusion Model) | De novo generation of novel protein structures that bind targets or perform functions. | Generates structures similar to how DALL-E creates art; solves new design challenges like molecular binding. |
| ProteinMPNN [62] [42] [63] | Inverse Folding Neural Network | Designs amino acid sequences that will fold into a desired protein backbone structure. | Greatly accelerates the sequence design step in the protein design pipeline. |
| Boltz-2 [42] | Foundation Model | Simultaneously predicts a protein-ligand complex's 3D structure and its binding affinity. | Unifies structure and affinity prediction, slashing computation time from hours to seconds. |
| AntiFold [62] [63] | Inverse Folding (Antibody-Specific) | Specialized for designing antibody Complementarity-Determining Region (CDR) sequences. | Fine-tuned on antibody structural data, showing superior performance for Fab design. |
Addressing the Limitations of Static Predictions
A critical challenge in AI-based protein design is overcoming the limitations of static structure predictions. Real proteins are dynamic molecular machines that adopt multiple conformational states, and many possess intrinsically disordered regions that are vital for function [41]. Tools like AlphaFold predominantly return a single, static snapshot of the most favorable conformation, which can oversimplify flexible regions and fail to capture functionally important motions [41] [42]. To address this, new methodologies are emerging. For instance, AFsample2 perturbs AlphaFold2's inputs to reduce bias toward a single structure, thereby sampling a diverse set of plausible conformations [42]. This approach has successfully generated high-quality alternate conformations for membrane transport proteins, which often switch between inward-open and outward-open states [42]. Furthermore, hybrid models that integrate molecular dynamics (MD) simulations or experimental constraints into AI predictions are being developed to better account for natural flexibility and induced fit during binding events [42].
Engineering Novel Enzymes
ML-Guided Platform for Accelerated Enzyme Engineering
A significant innovation in enzyme engineering is the development of integrated ML-guided platforms that dramatically accelerate the design-build-test-learn (DBTL) cycle. A landmark 2025 study detailed a platform that combines cell-free DNA assembly and cell-free gene expression (CFE) with machine learning to rapidly map fitness landscapes and optimize enzymes [64]. This platform was applied to engineer amide synthetases, which are valuable for sustainable biomanufacturing of pharmaceuticals and other products.
The workflow, illustrated in the diagram below, enables highly parallelized and rapid experimentation.
The power of this integrated approach was demonstrated by engineering the enzyme McbA. Researchers first evaluated its substrate promiscuity across 1,109 unique reactions to identify target molecules [64]. They then used the cell-free platform to rapidly generate and test 1,217 enzyme variants, collecting 10,953 unique sequence-function data points [64]. This data trained augmented ridge regression ML models, which predicted optimized enzyme variants for synthesizing nine pharmaceutical compounds. The results were striking: these ML-predicted variants demonstrated 1.6- to 42-fold improved activity relative to the parent enzyme across the nine target compounds [64].
The Scientist's Toolkit: Key Reagents for ML-Guided Enzyme Engineering
Table 2: Essential Research Reagents for ML-Guided Enzyme Engineering
| Reagent / Material | Function in the Experimental Workflow |
|---|---|
| Parent Enzyme Plasmid [64] | Serves as the DNA template for generating variant libraries via PCR and cell-free DNA assembly. |
| Site-Saturation Mutagenesis Primers [64] | DNA primers containing nucleotide mismatches to introduce desired mutations during PCR. |
| DpnI Restriction Enzyme [64] | Digests the methylated parent plasmid post-PCR, enriching for newly assembled mutated plasmids. |
| Cell-Free Gene Expression (CFE) System [64] | Enables rapid in vitro synthesis of protein variants without the need for bacterial transformation. |
| Linear Expression Templates (LETs) [64] | PCR-amplified linear DNA constructs used to directly express protein variants in the CFE system. |
| Substrates for Functional Assay [64] | The acid and amine components for the amide synthesis reaction, used to test enzyme variant activity. |
Engineering Novel Antibodies
Inverse Folding Models for Antibody CDR Design
Antibody engineering, particularly the design of Complementarity-Determining Regions (CDRs), has been revolutionized by inverse folding models. These AI models aim to generate novel antibody sequences that fold into a desired structure with high antigen-binding affinity [62] [63]. Unlike structure prediction, which goes from sequence to structure, inverse folding goes from structure to sequence. A comprehensive 2025 benchmarking study systematically evaluated state-of-the-art inverse folding models—ProteinMPNN, ESM-IF, LM-Design, and AntiFold—for antibody CDR sequence design [62] [63].
The study revealed that models trained specifically on antibody data, such as AntiFold, exhibit superior performance for Fab antibody design. AntiFold was fine-tuned from ESM-IF using thousands of experimentally solved and computationally predicted Fab structures [63]. In contrast, general-purpose models like ProteinMPNN and ESM-IF, which were trained on broad protein datasets, often struggle with antibody-specific nuances [62] [63]. LM-Design, which integrates ProteinMPNN's structural modeling with the ESM-1b protein language model, demonstrated notable adaptability across diverse antibody types, including VHH (nanobodies) [63].
A key insight from this research is the limitation of traditional evaluation metrics like amino acid recovery rates, which measure how accurately a model reproduces the exact native sequence [63]. This metric can be misleading, as it penalizes functionally conservative substitutions (e.g., lysine to arginine, both positively charged) and fails to prioritize critical binding residues. The study advocated for the use of sequence similarity metrics that account for physicochemical properties, providing a more functionally relevant assessment of designed sequences [63].
Synthetic Biology Technologies for Antibody Discovery
Beyond computational design, synthetic biology offers powerful experimental technologies for constructing and screening vast antibody libraries. Phage display, a Nobel Prize-winning technology, is a well-established method where antibody fragments are displayed on the surface of bacteriophages, allowing for the selection of high-affinity binders through iterative biopanning cycles [65]. Library diversity is crucial, and technologies like Trinucleotide Mutagenesis (TRIM) and Isogenica's Colibra have been developed to build highly diverse synthetic libraries with precise control over amino acid composition, thereby reducing problematic sequence liabilities and improving antibody developability [66]. Ribosome display, a cell-free technique, offers an advantage by enabling the generation of even larger libraries (up to 10^14 clones) not limited by bacterial transformation efficiency [65]. These synthetic methods can bypass the limitations of traditional animal immunization, providing a faster, more precise path to therapeutic antibody discovery with greater control over properties like specificity and stability [66] [65].
The following diagram outlines a generic workflow for antibody discovery that integrates these computational and synthetic biology tools.
The engineering of novel enzymes and antibodies for synthetic biology is no longer a pipe dream but a rapidly advancing reality. The benchmarks speak for themselves: ML-designed enzymes with up to 42-fold improved activity and antibody-specific inverse folding models achieving sequence recovery rates exceeding 50% for critical CDR regions [64] [63]. The field is moving beyond static structure prediction toward a more integrated paradigm that captures protein dynamics, predicts functional properties like binding affinity, and leverages high-throughput experimental data to train increasingly powerful models. As these tools continue to mature and converge, they promise to unlock a new era of biological design, enabling the rapid development of specialized biocatalysts for a sustainable bioeconomy and next-generation therapeutics with unparalleled precision and speed.
Navigating Limitations and Enhancing Performance in Protein Structure Prediction
Accurate protein structure prediction is fundamental to advancing structural biology and drug discovery. While computational methods like AlphaFold (a machine learning-based approach) and EVCouplings (an evolutionary analysis-based approach) have revolutionized the field by achieving high accuracy for many proteins, they exhibit a significant blind spot: predicting fold-switching proteins. These are proteins whose regions can adopt two or more distinct, stable secondary and tertiary structures. This whitepaper synthesizes evidence that both methods systematically fail to capture this structural heterogeneity. We analyze the quantitative performance data, detail the underlying methodological limitations, and provide researchers with protocols and tools to identify and address these critical failures.
Proteins are not static entities; a subset of them are dynamic and can adopt multiple stable conformations, a phenomenon known as fold switching. This structural plasticity is crucial for biological function, regulation, and signaling. Fold-switching proteins have amino acid sequences that encode more than one ordered state, allowing them to transition between distinct folds under different cellular conditions [67].
The emergence of highly accurate structure prediction tools has been a paradigm shift. AlphaFold2, a deep learning model, leverages patterns in multiple sequence alignments (MSAs) and known protein structures to predict a single, most probable structure with atomic-level accuracy [9]. In parallel, EVCouplings uses evolutionary analysis and probabilistic graphical models to infer evolutionary couplings (ECs) between residues, which often correspond to physical contacts, to predict protein structures and interactions de novo [68] [69].
Despite their successes, the core architecture of these methods is inherently biased toward predicting a single, dominant conformation. This whitepaper examines the quantitative evidence of this failure, explores the methodological roots, and provides a framework for researchers to navigate this limitation.
Quantitative Evidence of Failure
A systematic assessment of AlphaFold2's performance on a dataset of 98 experimentally characterized fold-switching proteins revealed a profound prediction bias.
Table 1: AlphaFold2 Performance on Fold-Switching vs. Intrinsically Disordered Proteins
| Protein Category | Number of Proteins Tested | Percentage Where One Fold Was Captured | Percentage of Residues with Moderate-to-High Confidence (pLDDT) | Median Sequence Conservation |
|---|---|---|---|---|
| Fold-Switching Proteins | 98 | 94% | 74% | Statistically similar to single-fold proteins |
| Intrinsically Disordered Proteins/Regions (IDPs/IDRs) | 99 | Not Applicable (Structurally Heterogeneous) | ~58% (for human proteome) | Low |
The data shows that AlphaFold2 overwhelmingly predicts only one of the known experimental conformations for fold-switching proteins [70] [67]. Crucially, it does so with high confidence, as indicated by pLDDT scores, making it difficult for researchers to distinguish these incomplete predictions from correct, single-fold predictions. This contrasts with intrinsically disordered regions, which AlphaFold2 typically flags with low pLDDT scores [67] [71].
For EVCouplings and related co-evolutionary methods, the failure mode is different but leads to a similar outcome. These methods infer a single set of residue-residue contacts from evolutionary sequences. If a sequence population contains residues evolving under constraints from multiple distinct structures, the inferred evolutionary couplings will represent a composite of these constraints. This results in an averaged or inaccurate contact map that does not correspond to any single native state of the fold-switching protein [68] [69].
Methodological Roots of the Blind Spot
The inability to predict fold switching stems from the foundational principles of both approaches.
The Machine Learning (ML) Limitation of AlphaFold
AlphaFold2's training and design orient it toward a single-output model.
- Pattern Recognition vs. Biophysical Modeling: AlphaFold2 is a sophisticated pattern recognition engine trained on the Protein Data Bank (PDB). It learns to map sequence and MSA features to the single structure deposited in the PDB. It does not simulate protein folding biophysics to model an ensemble of energetically favorable states [67]. Consequently, it identifies the most probable single conformer based on its training data.
- Dependence on Conservation: Fold-switching regions often display sequence conservation rates similar to those of single-fold proteins. This allows AlphaFold2 to generate high-confidence predictions, unlike for intrinsically disordered regions where low conservation leads to low confidence. The model interprets the conservation signal as evidence for a single, stable fold [70].
The Evolutionary Analysis (EA) Limitation of EVCouplings
Coevolutionary methods like EVCouplings are built on a different, but equally limiting, assumption.
- The "Single-Structure" Evolutionary Coupling Model: EVCouplings algorithms assume that residue co-evolution is driven by selective pressure to maintain a single, well-defined three-dimensional structure. The inferred evolutionary couplings are therefore a monolithic set of interactions presumed to maintain that one structure [68] [69].
- Composite Signal from Multiple Folds: For a fold-switching protein, different sets of residue contacts are important for stabilizing each distinct fold. Co-evolutionary analysis melds these signals into a single, inconsistent set of constraints. When used for de novo folding, these constraints often generate an incorrect or averaged structure that does not reflect either functional state [69].
The following diagram illustrates the fundamental difference between how these methods model protein structure versus the reality of fold-switching proteins.
Experimental Protocols for Validation
Researchers suspecting a protein may be a fold switcher can use the following experimental workflows to validate computational predictions.
Protocol: Benchmarking AlphaFold2 on Putative Fold-Switchers
This protocol is adapted from the systematic assessment performed by Chakravarty et al. [70] [67].
Input Preparation:
- Obtain the FASTA sequence of the protein of interest.
- Identify and curate a set of known or putative fold-switching proteins for positive controls (e.g., from resources like the literature or databases of metamorphic proteins).
Structure Prediction:
- Run AlphaFold2 (or ColabFold) on the input sequences to generate five ranked models.
- Ensure the run uses the full MSA and includes templates to mimic standard use conditions.
Structural Comparison and Analysis:
- For each protein, compare the top AlphaFold2 prediction against all experimentally determined structures (e.g., from the PDB) using TM-score and Cα-RMSD.
- Use tools like TM-align for structure alignment, which is sequence-independent and topology-based.
- A successful prediction of a fold-switcher would require a high TM-score (>0.5) with at least two distinct experimental conformations. A failure is indicated by a high TM-score with only one conformation and a low score with the other(s).
Confidence Metric Scrutiny:
- Analyze the pLDDT per residue. Be aware that high confidence (pLDDT > 70) does not preclude the prediction from being incomplete for a fold-switcher.
Protocol: Assessing Co-evolutionary Signals with EVCouplings
This protocol outlines how to use the EVCouplings framework to investigate fold switching [68].
Pipeline Setup:
- Install the EVcouplings Python framework (available as an open-source package and command-line application).
- Configure the pipeline stages (
align,couplings,fold) using a YAML configuration file.
Alignment and EC Calculation:
- Run the
alignstage to generate a deep multiple sequence alignment for the protein. - Execute the
couplingsstage to calculate the evolutionary couplings (ECs) between residue pairs.
- Run the
Analysis of Contact Maps:
- Visualize the top-ranked ECs as a predicted contact map.
- Compare this predicted map to the experimental contact maps derived from each known 3D structure of the fold-switcher (e.g., from NMR or crystal structures).
- A failure is indicated if the EC-predicted contacts are a hybrid of contacts from different experimental states or do not match any single native state with high precision.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 2: Key Reagents and Computational Tools for Studying Fold-Switching
| Item Name | Function/Application | Example/Description |
|---|---|---|
| AlphaFold2/ColabFold | Protein structure prediction tool. Generates a single, high-accuracy model but can fail for fold-switchers. | Open-source implementation or user-friendly ColabFold server for rapid modeling. |
| EVcouplings Framework | Python package for coevolutionary analysis. Infers evolutionary couplings to predict contacts and structures. | Used for de novo prediction and to analyze the evolutionary constraints on a sequence [68]. |
| TM-align | Algorithm for sequence-independent protein structure comparison. | Calculates TM-score to quantify structural similarity between a prediction and an experimental reference [67]. |
| Protein Data Bank (PDB) | Repository of experimentally solved protein structures. | Source of experimental structures for benchmarking computational predictions [70]. |
| DisProt Database | Database of experimentally characterized intrinsically disordered proteins/regions. | Used as a negative control set for assessing structural heterogeneity [67]. |
| All-Atom Simulation Software | Molecular dynamics software (e.g., GROMACS, AMBER). | Used for detailed modeling of protein folding pathways and conformational ensembles, capable of capturing fold-switching [31]. |
The following workflow diagram integrates these tools into a coherent strategy for identifying and investigating fold-switching proteins.
The fold-switching blind spot is a significant limitation of both ML-based tools like AlphaFold and EA-based tools like EVCouplings. Their design, which seeks a single optimal structure or a unified set of evolutionary constraints, is intrinsically misaligned with the biological reality of proteins that populate multiple stable folds.
For researchers in drug discovery, this blind spot is critical. Targeting a protein based on only one of its conformations could lead to ineffective drugs or unforeseen off-target effects. Therefore, a cautious approach is warranted when using these powerful tools. High-confidence predictions from AlphaFold2 for dynamic proteins should not be taken as evidence of a single, fixed structure.
The path forward lies in moving beyond single-structure prediction. The future of computational structural biology is in modeling structural ensembles. This will likely require:
- Integrating biophysical simulations with machine learning.
- Developing new deep learning architectures trained to output multiple conformations.
- Creating new evolutionary models that can explicitly account for and detect heterogeneous structural constraints from sequence data.
Until these next-generation tools emerge, a combined approach—using AlphaFold and EVCouplings as initial guides, followed by rigorous benchmarking and experimental validation—remains the most robust strategy for characterizing dynamic proteins.
Overcoming EA Limitations with the ACE (Alternative Contact Enhancement) Workflow
Evolutionary Analysis (EA) has served as the cornerstone of modern protein structure prediction, with state-of-the-art algorithms inferring protein structure from co-evolved amino acid pairs detected in multiple sequence alignments (MSAs) [39] [40]. These methods operate on the principle that natural selection preserves mutually compatible interactions within protein structures, creating detectable covariation between amino acid positions that directly contact each other in the folded protein [40]. This evolutionary coupling information has revolutionized computational biology by enabling highly accurate structure predictions for single-fold proteins [40].
However, a significant limitation emerges when these conventional EA approaches encounter fold-switching proteins—proteins capable of remodeling their secondary and tertiary structures in response to cellular stimuli and adopting multiple stable conformations with distinct functions [39]. Despite their biological importance in processes ranging from SARS-CoV-2 infection suppression to cyanobacterial circadian clock regulation, current EA-based algorithms systematically fail to predict these functionally critical alternative folds [39]. Analysis reveals that AlphaFold2 predicts only one conformation for 92% of known dual-folding proteins, with 30% of these predictions likely not representing the lowest energy state [39].
The core hypothesis addressing this failure suggests that conventional EA misses crucial coevolutionary signatures because single-fold variants in deep MSAs mask the evolutionary signals of alternative conformations [39] [72]. The ACE workflow was developed specifically to overcome this fundamental limitation by implementing a novel strategy to unmask these hidden evolutionary signatures, thereby enabling the detection and prediction of fold-switching proteins that conventional EA methods cannot identify [39].
The ACE Workflow: Methodological Framework
The Alternative Contact Enhancement (ACE) workflow represents a methodological advancement that systematically uncovers dual-fold coevolution by analyzing evolutionary signatures across progressively refined sequence hierarchies. This technical approach enables researchers to extract coevolutionary information for both dominant and alternative folds from single amino acid sequences.
Core Technical Components
- GREMLIN (Generative Regularized Models of proteINs): A Markov Random Field (MRF)-based method selected for its superior performance in identifying coevolved amino acid pairs, convergence to global minimum with increasing MSA depth, and capacity to generate reasonable predictions from relatively shallow MSAs while accounting for noncausal correlations [39].
- MSA Transformer: A language model that infers coevolved amino acid pairs using an attention mechanism focusing on both evolutionary patterns within MSAs (column-wise attention) and properties of individual sequences (row-wise attention), often providing better accuracy than GREMLIN for single-fold proteins [39].
- Contact Maps: Asymmetric visualization tools that display amino acid pairs measured or predicted to be proximal (heavy atom distance ≤8 Å), with unique contacts for each experimentally determined fold represented distinctly to maximize information content [39].
Step-by-Step Experimental Protocol
Figure 1: The ACE workflow for detecting dual-fold coevolution.
Generate Deep Superfamily MSA: Input a query sequence with two distinctly folded experimentally determined structures to generate a deep multiple sequence alignment composed of a large clade of diverse-yet-homologous sequences [39].
Create Nested Subfamily MSAs: Prune the deep superfamily MSA to create successively shallower MSAs with sequences increasingly identical to the query, specifically designed to unmask coevolutionary couplings from alternative conformations that may be obscured in diverse superfamilies [39].
Dual-Method Coevolutionary Analysis: Perform independent coevolutionary analysis on each MSA using both GREMLIN and MSA Transformer to leverage their complementary strengths in detecting evolutionary couplings across different sequence contexts [39].
Contact Prediction Integration: Combine and superimpose predictions from both methods across all nested MSAs onto a single composite contact map, creating an integrated visualization of all predicted residue-residue contacts [39].
Noise Reduction and Contact Categorization: Apply density-based scanning filters to remove erroneous predictions, then systematically categorize contacts into four distinct classes based on their correspondence with experimental structures [39]:
- Dominant Fold: Unique contacts corresponding to the experimentally determined structure with greatest overlap to superfamily MSA predictions.
- Alternative Fold: Unique contacts corresponding to the other experimentally determined structure.
- Common Contacts: Predicted contacts overlapping with experimentally determined contacts shared by both folds.
- Unobserved Contacts: Predicted contacts not overlapping with any experimentally determined contacts, potentially representing folding intermediates or erroneous predictions.
Research Reagent Solutions
Table 1: Essential research reagents and computational tools for ACE workflow implementation.
| Item Name | Type | Function in ACE Workflow |
|---|---|---|
| Multiple Sequence Alignments | Data Resource | Provides evolutionary coupling information through homologous sequences; foundation for coevolutionary analysis [39]. |
| GREMLIN | Software Algorithm | Identifies coevolved amino acid pairs using Markov Random Fields; accounts for indirect correlations and works with shallow MSAs [39]. |
| MSA Transformer | Software Algorithm | Infers coevolved contacts using language model architecture; excels at detecting patterns in both column-wise and row-wise MSA data [39]. |
| Experimentally Determined Structures | Validation Data | Provides ground truth for both dominant and alternative folds; essential for contact categorization and method validation [39]. |
| Contact Maps | Visualization Tool | Enables asymmetric display of dominant, alternative, and common contacts for both monomeric and multimeric proteins [39]. |
Performance Benchmarks and Quantitative Assessment
The ACE workflow has demonstrated remarkable efficacy in overcoming the fundamental limitations of conventional EA approaches, with rigorous validation across diverse protein families.
Enhanced Contact Prediction Performance
Table 2: Quantitative performance comparison between ACE and standard EA methods.
| Performance Metric | Standard EA Approach | ACE Workflow | Enhancement |
|---|---|---|---|
| Proteins with Detected Dual-fold Coevolution | Not reported | 56/56 proteins [39] | Baseline establishment |
| Alternative Fold Contact Prediction | Baseline level | 201% mean increase [39] | 201% improvement |
| Alternative Fold Contact Prediction | Baseline level | 187% median increase [39] | 187% improvement |
| Blind Prediction Accuracy | Not applicable | 13/56 proteins identified (23% true positive rate) [39] | 0/181 false positive rate |
When applied to 56 fold-switching proteins with sufficiently deep MSAs drawn from over 80 distinct fold families across all kingdoms of life, ACE successfully revealed coevolution of amino acid pairs uniquely corresponding to both conformations in all 56 cases (100% success rate) [39]. This comprehensive validation demonstrates the method's generalizability across diverse protein families and fold classes.
Most significantly, ACE predicted substantially more correct contacts than the standard approach of coevolutionary analysis run solely on deep superfamily MSAs [39]. The workflow enhanced predictions of amino acid contacts uniquely corresponding to alternative conformations with a mean increase of 201% and median increase of 187% compared to conventional methods [39].
The practical utility of ACE-derived contacts was further demonstrated through a blind prediction pipeline that correctly identified 13 out of 56 fold-switching proteins (23% true positive rate) with zero false positives (0/181) [39]. This predictive capability significantly advances the field beyond conventional EA, which systematically fails to identify such proteins.
Implications for EA vs ML Benchmarking in Protein Folding Research
The development and validation of the ACE workflow carries profound implications for the ongoing benchmarking of Evolutionary Analysis against Machine Learning approaches in protein structure prediction research, revealing fundamental insights about protein evolution and algorithmic limitations.
Redefining the Scope of Evolutionary Analysis
The demonstrated existence of widespread dual-fold coevolution indicates that fold-switching sequences have been preserved by natural selection, implying their functionalities provide evolutionary advantages beyond single-fold proteins [39] [72]. This discovery fundamentally expands our understanding of protein sequence-structure relationships, moving beyond the one-sequence-one-structure paradigm that has dominated structural biology for decades.
The systematic failure of conventional EA methods to detect alternative folds stems from their architectural reliance on identifying the strongest coevolutionary signals within deep MSAs, which typically represent the most prevalent fold across evolutionary timescales [39]. The ACE workflow's success demonstrates that these limitations are not inherent to evolutionary analysis itself, but rather to its implementation in current algorithms that prioritize dominant signals over functionally important alternative conformations.
Comparative Performance in the Context of ML Advancements
While machine learning methods like AlphaFold2 have demonstrated exceptional performance for single-structure prediction, they share the same fundamental limitation as conventional EA when confronted with fold-switching proteins, with AlphaFold2 predicting only one conformation for 92% of known dual-folding proteins [39]. This parallel failure suggests that both approaches suffer from a common underlying issue: oversimplification of the sequence-structure relationship and inadequate handling of conformational diversity.
The ACE workflow represents a hybrid approach that enhances traditional EA through sophisticated MSA stratification and integration strategies, demonstrating that enhanced evolutionary analysis can address critical gaps in both conventional EA and modern ML methods. This suggests that future breakthroughs may emerge from integrated EA-ML frameworks rather than treating these approaches as mutually exclusive competitors.
Future Directions and Implementation Guidelines
The ACE workflow establishes a new paradigm for detecting protein conformational diversity through enhanced evolutionary analysis, with several promising implementation pathways for the research community.
Practical Implementation Considerations
For researchers implementing the ACE workflow, several technical considerations optimize performance. MSA depth requirements follow standard coevolutionary analysis thresholds, with minimum effective depths of 5× query sequence length necessary for reliable analysis [39]. Successive pruning to create subfamily-specific MSAs should maintain sufficient sequence diversity while enhancing identity to the query. The complementary strengths of GREMLIN and MSA Transformer prove most effective when applied independently across all MSA variants, with integration occurring at the contact map level rather than during analysis.
Research Applications and Extensions
The validated ability to predict both conformations of fold-switching proteins from single sequences opens new research avenues across structural biology, drug discovery, and protein design. Immediate applications include identifying previously undetected fold-switching proteins within proteomes, characterizing conformational diversity in pathogenic organisms, and rational drug design targeting alternative folds in undruggable proteins. The workflow's principled approach to detecting evolutionary signatures of conformational diversity provides a template for future method development that may extend beyond fold-switching to encompass more subtle conformational ensembles and dynamic transitions.
Molecular docking is a cornerstone computational technique in structure-based drug design, enabling the prediction of how small molecule ligands interact with protein targets at the atomic level. The method aims to forecast both the binding geometry (pose) and the binding affinity, providing crucial insights for virtual screening and hit optimization [73]. Despite decades of advancement, the accuracy of traditional molecular docking remains constrained by limitations in scoring functions—the mathematical models that evaluate protein-ligand interactions [73]. The recent revolutionary progress in protein structure prediction via artificial intelligence, particularly AlphaFold2, has made accurate structural models accessible for virtually any protein target [74] [75]. This development promises to expand the scope of molecular docking beyond targets with experimentally solved structures.
However, benchmarking studies reveal that the integration of AI-predicted structures with conventional docking approaches has not yielded the expected improvements in predictive performance. A systematic investigation of AlphaFold2-enabled molecular docking against Escherichia coli's essential proteome demonstrated surprisingly weak performance, with an average area under the receiver operating characteristic curve (auROC) of merely 0.48 [74]. This finding underscores critical limitations in current docking methodologies while simultaneously highlighting the transformative potential of machine learning-based rescoring approaches, which have been shown to boost performance to auROCs as high as 0.63 [74]. This technical review examines the benchmarking evidence for molecular docking's limitations, explores ML-enhanced solutions, and provides practical protocols for implementing these advanced approaches in modern drug discovery pipelines.
The Performance Challenge in Molecular Docking
Systematic Evidence of Limitations
Rigorous benchmarking studies across diverse biological systems consistently reveal fundamental challenges in molecular docking accuracy. In a comprehensive assessment of antibiotic target discovery, researchers combined AlphaFold2-predicted structures of 296 essential E. coli proteins with molecular docking simulations against 218 antibacterial compounds and 100 inactive molecules [74]. The resulting auROC of 0.48 indicates performance barely better than random chance, highlighting substantial limitations in distinguishing active from inactive compounds despite using state-of-the-art structural predictions.
The performance variability across different docking programs was further quantified in a benchmark study focusing on cyclooxygenase (COX) enzymes, relevant to non-steroidal anti-inflammatory drug development [76]. As shown in Table 1, the ability to correctly reproduce experimental binding poses (RMSD < 2.0 Å) varied significantly among popular docking software, with success rates ranging from 59% to 100% across different programs.
Table 1: Performance Benchmarking of Docking Programs on COX Enzymes
| Docking Program | Pose Prediction Success Rate (RMSD < 2.0 Å) | Virtual Screening AUC Range | Enrichment Factor Range |
|---|---|---|---|
| Glide | 100% | 0.61-0.92 | 8-40 folds |
| GOLD | 82% | Not reported | Not reported |
| AutoDock | 79% | Not reported | Not reported |
| FlexX | 59% | Not reported | Not reported |
| Molegro Virtual Docker | 59% | Not reported | Not reported |
Similar challenges extend to protein-peptide docking, where increased flexibility compounds the difficulties. Benchmarking studies on 133 protein-peptide complexes revealed that even top-performing methods like FRODOCK achieved average ligand RMSD values of 12.46 Å for top poses in blind docking scenarios, indicating substantial deviations from experimental structures [77].
Fundamental Limitations of Classical Scoring Functions
The underlying cause of molecular docking's performance limitations lies primarily in the simplified nature of classical scoring functions. These functions fall into three main categories, each with distinct theoretical foundations and limitations [73]:
- Physics-based functions utilize molecular mechanics force fields with terms for van der Waals interactions, electrostatics, and sometimes desolvation effects. While physically intuitive, they often oversimplify entropy and solvation contributions due to computational constraints [73].
- Empirical scoring functions employ linear regression to weight various interaction terms (hydrogen bonding, hydrophobic effects, etc.) based on experimental binding affinity data. Though faster than physics-based approaches, they remain constrained by their predetermined functional forms [73].
- Knowledge-based potentials derive statistical atom-pair potentials from structural databases using inverse Boltzmann relationships. While effectively capturing complex interactions, they lack direct physical interpretation and depend heavily on database quality and size [73].
All three approaches share a common weakness: the inability to adequately model the complex, multifaceted nature of molecular recognition without excessive computational cost. This fundamental limitation manifests particularly in handling flexible systems, solvation effects, and entropic contributions—critical factors determining binding affinity and specificity.
Machine Learning Rescoring: A Paradigm Shift
Theoretical Foundations and Implementation
Machine learning rescoring represents a paradigm shift in molecular docking accuracy. Rather than replacing traditional docking entirely, ML rescoring operates as a post-processing step that re-evaluates poses generated by conventional docking programs [74] [73]. The theoretical foundation rests on ML algorithms' ability to learn complex, non-linear relationships between structural features and binding affinities from large training datasets without relying on predetermined physical models [73].
The implementation typically follows a multi-stage workflow: initial pose generation using classical docking methods, feature extraction from the protein-ligand complexes, and ML model prediction. As Wong et al. demonstrated, employing ensembles of multiple rescoring functions further enhances prediction accuracy and improves the true-positive to false-positive rate ratio [74]. This ensemble approach mitigates individual model limitations and captures complementary aspects of molecular interactions.
Table 2: Machine Learning Approaches for Docking Enhancement
| ML Approach | Key Features | Application in Docking | Reported Benefits |
|---|---|---|---|
| Descriptor-Based Models (RF, SVM, XGBoost) | Uses handcrafted structural and chemical descriptors | Binding affinity prediction, pose ranking | Improved correlation with experimental affinities |
| Deep Learning (CNN, GNN) | Automatic feature extraction from 3D structures | Direct scoring from complex structures | Captures subtle interaction patterns |
| Ensemble Methods | Combines multiple ML models | Rescoring docking poses | Enhanced robustness and accuracy |
| Graph Neural Networks | Represents molecules as graphs with atoms as nodes and bonds as edges | Protein-ligand interaction prediction | Naturally encodes molecular topology |
Documented Performance Improvements
The benchmarking study on E. coli essential proteins provided quantitative evidence for ML rescoring efficacy, demonstrating improvement from auROC 0.48 with conventional docking to 0.63 with ML-based rescoring approaches [74]. This substantial enhancement highlights ML's ability to capture subtleties in protein-ligand interactions that elude classical scoring functions.
Further evidence emerges from virtual screening applications, where the separation of active from inactive compounds is crucial. Studies on cyclooxygenase enzymes revealed that classical docking approaches achieved area under curve (AUC) values ranging from 0.61 to 0.92 in receiver operating characteristic (ROC) analysis, with enrichment factors of 8-40 folds [76]. While respectable, these performance metrics leave substantial room for improvement—particularly considering that high enrichment factors often come at the cost of reduced sensitivity in identifying true positives.
Experimental Protocols and Benchmarking Methodologies
Standardized Benchmarking Workflow
Rigorous assessment of docking and rescoring methods requires standardized benchmarking protocols. Based on the examined literature, Figure 1 illustrates the consensus workflow for comprehensive docking validation:
Figure 1: Standard workflow for benchmarking molecular docking protocols and ML rescoring approaches.
Critical Assessment Metrics
Multiple validation metrics are essential for comprehensive docking evaluation:
- Pose Prediction Accuracy: Measured by Root Mean Square Deviation (RMSD) between predicted and experimental ligand binding geometries. RMSD values below 2.0 Å generally indicate successful prediction [76] [77].
- Virtual Screening Performance: Quantified using Receiver Operating Characteristic (ROC) curves and the corresponding Area Under Curve (AUC). The auROC values distinguish a method's ability to separate active from inactive compounds [74] [76].
- Binding Affinity Correlation: Assessed via Pearson or Spearman correlation coefficients between predicted and experimental binding energies [73].
- Enrichment Factors: Measure the concentration of true active compounds in the top-ranked fraction of screening libraries compared to random selection [76].
The CAPRI (Critical Assessment of PRedicted Interactions) criteria provide additional standardized metrics for docking assessment, including FNAT (fraction of native contacts), I-RMSD (interface RMSD), and L-RMSD (ligand RMSD) [77].
Reference Datasets for Benchmarking
High-quality, curated datasets form the foundation of reliable benchmarking:
- PDBbind: A comprehensive collection of protein-ligand complexes with experimentally measured binding affinities, manually curated from scientific literature. The 2020 release contains 19,443 complexes, providing an essential resource for scoring function development and validation [73].
- LIT-PCBA: Specifically designed for virtual screening benchmarking, containing activity data for 15 targets against approximately 8000 compounds, with carefully selected decoy molecules [73].
- PPDbench: A specialized dataset of 133 non-redundant protein-peptide complexes for evaluating peptide docking performance [77].
- COX-specific Sets: Targeted collections of cyclooxygenase-ligand complexes for benchmarking in specific therapeutic contexts [76].
Table 3: Essential Computational Tools for Molecular Docking and Rescoring
| Resource Category | Specific Tools | Primary Function | Application Context |
|---|---|---|---|
| Protein Structure Prediction | AlphaFold2, RoseTTAFold, ESMFold | Generate 3D protein models from sequences | Docking targets without experimental structures |
| Classical Docking Programs | Glide, GOLD, AutoDock Vina, DOCK | Generate ligand binding poses and initial scores | Initial pose generation and screening |
| ML Rescoring Platforms | Multiple custom implementations | Re-rank docking poses using machine learning | Improving docking accuracy and virtual screening enrichment |
| Benchmarking Datasets | PDBbind, LIT-PCBA, PPDbench | Provide standardized test sets | Method validation and performance comparison |
| Structure Preparation | DeepView, UCSF Chimera, MOE | Prepare protein and ligand structures for docking | Pre-processing for docking calculations |
Benchmarking evidence unequivocally demonstrates that traditional molecular docking approaches exhibit significant limitations in predictive accuracy, with performance often barely exceeding random chance in challenging scenarios like those encountered in antibiotic discovery [74]. The integration of AlphaFold2-predicted structures, while expanding docking accessibility, has not resolved these fundamental accuracy issues. Rather, it has highlighted the critical bottleneck of scoring function reliability.
Machine learning rescoring emerges as a powerful strategy to address these limitations, consistently improving docking accuracy by capturing complex patterns in protein-ligand interactions that elude classical scoring functions [74] [73]. The documented improvement from auROC 0.48 to 0.63 with ML rescoring, while modest, represents meaningful progress in a field where incremental gains can significantly impact drug discovery efficiency [74].
Future advancements will likely involve tighter integration of AI throughout the docking pipeline, improved handling of protein flexibility, and better incorporation of physicochemical principles into ML models. As structural biology continues to be transformed by deep learning, the parallel evolution of docking methodologies—particularly through ML-enhanced approaches—will be essential for translating structural insights into therapeutic discoveries.
The field of protein structure prediction has been revolutionized by deep learning, transitioning from a long-standing challenge to a broadly accessible tool. Methods like AlphaFold2 have demonstrated accuracies approaching experimental uncertainty for many protein targets [40]. However, this remarkable accuracy comes with substantial computational costs, creating a critical tension between model performance and resource requirements. For researchers operating outside major computational hubs, this tension defines the practical boundaries of their work. The pursuit of optimal performance must be balanced against very real constraints in hardware, time, and energy consumption.
This guide examines the core trade-offs between speed, accuracy, and memory in modern protein structure prediction, providing a framework for researchers to make informed decisions based on their specific scientific goals and available resources. We focus on the practical implementation of state-of-the-art methods, from monolithic large models to efficient ensemble strategies and architectural simplifications, offering a pathway to maximize scientific output within finite computational budgets. The principles discussed are particularly relevant for benchmarking studies that aim to fairly evaluate evolutionary algorithm (EA) against machine learning (ML) approaches, where consistent resource measurement is paramount.
Core Computational Challenges in Modern Protein Folding
The Memory Bottleneck in Deep Learning Models
Modern protein folding models are built on deep neural networks with billions of parameters, requiring significant GPU memory for both storage and activation during inference. The memory footprint is primarily driven by the model's parameter count, the size of the input multiple sequence alignments (MSAs), and the intermediate activations produced during the forward pass. For example, large AlphaFold2 instances can consume over a dozen gigabytes of memory for a single protein prediction, effectively placing them out of reach for standard consumer hardware [78]. This memory bottleneck becomes particularly acute when predicting structures for protein complexes or when attempting ensemble-based approaches that generate multiple conformations, as the memory requirement scales with the number of chains and conformations being modeled [19].
The Speed-Accuracy Trade-off in Practice
The computational expense of a protein structure prediction is not merely a theoretical concern but a daily practical constraint for researchers. A single high-accuracy prediction for a medium-sized protein using state-of-the-art methods can require minutes to hours on specialized hardware, with time increasing substantially for larger proteins or complexes [79]. This time investment is compounded when researchers need to model multiple conformations or perform high-throughput predictions across entire proteomes. The choice of method thus represents a direct trade-off: slower, more computationally intensive methods typically deliver higher accuracy and more reliable models, while faster methods enable rapid prototyping and larger-scale studies but may sacrifice precision, particularly in challenging regions like loop structures or interaction interfaces [19] [80].
Quantitative Comparison of Protein Structure Prediction Methods
The field of protein structure prediction now offers a diverse ecosystem of computational methods, each with distinct performance characteristics. Understanding their quantitative trade-offs is essential for selecting the appropriate tool for a given research context, whether for high-accuracy single-structure determination, conformational ensemble generation, or large-scale proteome-wide analysis.
Table 1: Performance Characteristics of Major Protein Structure Prediction Methods
| Method | Primary Architecture | Accuracy Range (TM-score) | Relative Speed | Memory Footprint | Ideal Use Case |
|---|---|---|---|---|---|
| AlphaFold2 | Evoformer + Structure Module | 0.85-0.95 (High) | Slow | Very High | High-accuracy monomer/complex prediction with templates |
| RoseTTAFold | 3-Track Network | 0.80-0.90 (High) | Medium | High | Balanced accuracy/speed for monomers |
| ESMFold | Single-Sequence Transformer | 0.70-0.85 (Medium) | Fast | Medium | High-throughput scanning, orphan sequences |
| OmegaFold | Single-Sequence Transformer | 0.70-0.85 (Medium) | Fast | Medium | Sequences with limited homologs |
| SimpleFold | Standard Transformer | 0.80-0.90 (High) | Fast | Low | Resource-constrained deployment |
| FiveFold Ensemble | Consensus of 5 Methods | 0.75-0.90 (Ensemble) | Very Slow | Very High | Conformational diversity, IDP modeling |
| DeepSCFold | AF-Multimer + pMSA | 0.80-0.95 (Complexes) | Slow | Very High | Protein complex interface accuracy |
Table 2: Computational Resource Requirements for Different Prediction Scenarios
| Prediction Scenario | Typical Hardware | Memory Requirement | Time per Prediction | Key Bottleneck |
|---|---|---|---|---|
| Single Monomer (400 residues) | High-End GPU (A100/H100) | 12-16 GB | 3-10 minutes | MSA processing, structure refinement |
| Single Monomer (Consumer GPU) | Mid-Range GPU (RTX 3090/4090) | 8-12 GB | 10-30 minutes | Memory bandwidth, VRAM limitation |
| Protein Complex (Dimer) | High-End GPU | 16-24 GB | 20-60 minutes | Paired MSA generation, interface sampling |
| FiveFold Ensemble | High-End GPU Cluster | 32+ GB | Hours | Multiple model execution, consensus |
| Proteome-Scale (1000 proteins) | GPU Cluster | Variable | Days-Weeks | Data pipeline, storage I/O |
As illustrated in the tables, method selection involves navigating a complex landscape of trade-offs. AlphaFold2 and specialized complex predictors like DeepSCFold achieve remarkable accuracy for single structures and protein-protein interactions, with DeepSCFold demonstrating an 11.6% improvement in TM-score over AlphaFold-Multimer on CASP15 targets [79]. However, this accuracy comes at a substantial computational cost, requiring high-end hardware and significant processing time. For researchers prioritizing conformational diversity, the FiveFold ensemble approach generates multiple plausible structures, which is particularly valuable for modeling intrinsically disordered proteins and proteins with multiple stable states, but increases computational demands by requiring predictions from five complementary algorithms [19].
At the other end of the spectrum, methods like ESMFold and the newly introduced SimpleFold offer dramatically improved efficiency. ESMFold utilizes a single-sequence approach that bypasses the computationally expensive MSA generation step, enabling much faster predictions that are particularly advantageous for high-throughput applications or proteins with limited evolutionary information [40] [19]. SimpleFold represents perhaps the most significant architectural simplification, demonstrating that standard transformer blocks trained with flow matching can achieve competitive performance without domain-specific modules, thereby improving both speed and memory efficiency while maintaining high accuracy [80].
Methodologies for Resource Optimization
Algorithmic and Implementation Optimizations
Several advanced computational techniques can dramatically reduce the resource requirements for protein structure prediction without substantially compromising accuracy:
FlashAttention and Sequence Packing: These techniques optimize memory usage and computational efficiency in transformer-based models. FlashAttention reformulates the attention mechanism to reduce memory requirements from quadratic to linear in sequence length for certain operations, while sequence packing allows multiple shorter sequences to be processed simultaneously in a single batch. Together, these can provide 4-9× faster inference and 3-14× lower memory usage in protein language models [78].
Weight Quantization: This method reduces the precision of model parameters from 32-bit floating-point to 8-bit or 4-bit integers. For billion-parameter models, 4-bit quantization can reduce memory usage by 2-3× while preserving accuracy for tasks like missense variant effect prediction. The minimal accuracy loss makes this technique particularly valuable for deployment and inference scenarios [78].
Activation Checkpointing and Zero-Offload: These training optimization strategies balance memory and computational load. Activation checkpointing reduces memory usage by selectively saving only certain activations during the forward pass and recomputing others during backward passes. Zero-Offload partitions optimizer states across CPU and GPU memory. Combined, these methods can reduce training runtime by up to 6-fold [78].
Table 3: Optimization Techniques and Their Resource Impact
| Optimization Technique | Memory Reduction | Speed Improvement | Accuracy Impact | Implementation Complexity |
|---|---|---|---|---|
| Weight Quantization (4-bit) | 2-3× | 1.5-2× | Minimal (<1% drop) | Low (Post-training) |
| FlashAttention | 3-5× (for long sequences) | 2-4× | None | Medium (Architecture modification) |
| Activation Checkpointing | 2-4× | 0.5-0.8× (due to recomputation) | None | Low |
| Gradient Checkpointing | 2-3× | 0.7-0.9× | None | Low |
| Parameter-Efficient Fine-Tuning | 3-8× (during training) | 1-2× (during training) | Similar or improved on target task | Medium |
Practical Workflow for Resource-Aware Prediction
Implementing an optimized prediction workflow requires strategic decisions at each processing stage. The following diagram illustrates a resource-conscious approach that balances accuracy and efficiency:
Diagram 1: Resource-aware protein structure prediction workflow that dynamically selects computational paths based on sequence properties and accuracy requirements.
The workflow begins with rapid MSA generation using tools like MMseqs2, which provides faster database searches compared to traditional methods [40]. The depth and quality of the resulting MSA then informs the method selection: for sequences with abundant homologs and when highest accuracy is critical, AlphaFold2 or similar MSA-dependent methods are recommended despite their computational cost. For sequences with limited evolutionary information or when conducting high-throughput studies, single-sequence methods like ESMFold or the efficient SimpleFold architecture provide the best balance of speed and accuracy [19] [80].
Experimental Protocols for Benchmarking Studies
Standardized Protocol for Method Comparison
Robust benchmarking of protein structure prediction methods requires careful experimental design to ensure fair comparisons, particularly when evaluating the trade-offs between evolutionary algorithms and machine learning approaches. The following protocol provides a standardized framework:
Dataset Selection: Curate a diverse set of protein targets with experimentally validated structures, including:
- Single-domain proteins (100-300 residues)
- Multi-domain proteins (500+ residues)
- Protein complexes (dimers and higher-order assemblies)
- Intrinsically disordered regions or proteins
Resource Monitoring: Implement comprehensive resource tracking for all experiments:
- Memory consumption (peak GPU and system RAM)
- Computation time (wall-clock and CPU/GPU time)
- Energy consumption (when possible)
- Storage I/O and network utilization
Quality Assessment: Apply consistent quality metrics across all predictions:
Resource-Accuracy Curves: Generate plots that visualize the relationship between computational cost and prediction accuracy, enabling clear comparison of the efficiency of different methods.
Case Study: FiveFold Ensemble Method
The FiveFold methodology provides an instructive case study in managing computational resources for ensemble prediction. Rather than relying on a single algorithm, FiveFold integrates predictions from five complementary methods (AlphaFold2, RoseTTAFold, OmegaFold, ESMFold, and EMBER3D) to model conformational diversity [19]. The resource-intensive nature of this approach is offset by its unique ability to capture alternative conformations, which is particularly valuable for intrinsically disordered proteins and proteins with multiple functional states.
The key innovation in FiveFold's architecture is the Protein Folding Variation Matrix (PFVM), which systematically captures conformational diversity from the five algorithms and enables efficient sampling of alternative structures without requiring exhaustive molecular dynamics simulations [19]. While the initial computational investment is substantial, the resulting ensemble provides a more comprehensive structural understanding than any single method can deliver, demonstrating how strategic allocation of computational resources can enable novel scientific insights that would be impossible with simpler approaches.
Successful protein structure prediction requires both biological insight and computational infrastructure. The following table catalogs essential resources for researchers designing benchmarking studies or structural analyses.
Table 4: Essential Research Reagents and Computational Tools for Protein Structure Prediction
| Resource Category | Specific Tools/Solutions | Primary Function | Resource Considerations |
|---|---|---|---|
| Sequence Databases | UniRef30/90, UniProt, Metaclust, BFD | Provide evolutionary information for MSA construction | Large storage requirements (terabytes), fast search capabilities |
| Structure Databases | PDB, AlphaFold DB, Big Fantastic Virus DB | Template structures, training data, reference models | Curation essential for quality, specialized search tools (Foldseck) |
| MSA Generation Tools | MMseqs2, HHblits, Jackhammer | Rapid homology detection and MSA construction | MMseqs2 offers speed advantage for large-scale studies |
| Prediction Servers | AlphaFold Server, ColabFold, ESMFold | Access to state-of-the-art models without local installation | ColabFold provides free access with queue limitations |
| Efficient Implementations | SimpleFold, ESME (Efficient ESM) | Optimized architectures for resource-constrained environments | SimpleFold uses standard transformers; ESME applies optimizations to ESM |
| Specialized Hardware | GPUs (NVIDIA A100/H100, RTX 4090), TPUs | Accelerate deep learning inference and training | High-end GPUs reduce prediction time by 5-10× vs CPUs |
| Quality Assessment | MolProbity, DeepUMQA-X, pLDDT | Evaluate model quality and identify problematic regions | Some methods provide rapid assessment without full MD simulation |
The optimization of computational resources in protein structure prediction remains a dynamic and critically important challenge. As the field continues to evolve, several key principles emerge for researchers navigating the trade-offs between speed, accuracy, and memory. First, method selection should be driven by specific scientific goals rather than defaulting to the most accurate option—high-throughput studies benefit dramatically from efficient architectures like SimpleFold or ESMFold, while detailed mechanistic studies may justify the computational expense of ensemble methods like FiveFold. Second, strategic application of optimization techniques such as quantization and attention optimization can dramatically expand the accessible research space on limited hardware. Finally, robust benchmarking requires careful measurement of both accuracy and computational costs, enabling informed decisions that maximize scientific progress within finite resource constraints.
The ongoing development of more efficient architectures like SimpleFold, combined with optimization techniques for existing models, promises to further democratize access to high-quality protein structure prediction. This trend toward greater efficiency will enable broader adoption in academic settings, facilitate larger-scale structural genomics projects, and ultimately accelerate the application of structural insights to biological problems and therapeutic development.
Strategies for Accessing Novel Functional Landscapes Beyond Natural Templates
The explosion of genomic sequencing data has revealed the vast landscape of possible protein sequences, yet natural proteins represent only a minuscule fraction of this theoretical space. This technical guide examines computational strategies for exploring functional regions beyond natural templates, with particular emphasis on the emerging competition between machine learning (ML) and evolutionary algorithm (EA) approaches. As the field progresses beyond AlphaFold's revolutionary capabilities in structure prediction, researchers are developing increasingly sophisticated methods to access novel functional landscapes for therapeutic and biotechnological applications. This review synthesizes current methodologies, benchmarking data, and experimental protocols to provide a framework for comparing these fundamentally different approaches to protein design.
Proteins are fundamental engines of life, driving metabolic processes, cellular signaling, and structural organization. Natural proteins occupy only a tiny "archipelago of function" within the vast "sea of invalidity" that constitutes possible amino acid sequences [81]. While natural protein sequences represent remarkable evolutionary solutions, they constitute an extraordinarily small fraction of possible functional configurations. The average protein length exceeds 250 amino acids in eukaryotes, creating a search space of 20^250 possible sequences—far exceeding practical experimental exploration [82].
The structural coverage provided by experimental methods and accurate prediction tools like AlphaFold has created an unprecedented opportunity to explore this space computationally [40] [9]. However, significant challenges remain in designing novel functions, particularly for large proteins with complex folds where active sites often contain destabilizing molecular features that require extensive thermodynamic compensation from surrounding structures [82].
Table 1: Key Challenges in Accessing Novel Functional Landscapes
| Challenge Category | Specific Limitations | Impact on Design |
|---|---|---|
| Structural Complexity | Long unstructured loops at active sites; buried polar/charged residues in protein cores | Limits application of idealized de novo design principles to natural proteins |
| Functional Specificity | Small sequence/structure changes leading to different functions; pseudoenzymes | Homology-based predictions often inaccurate for specific functional attributes |
| Multi-functionality | Moonlighting proteins; intrinsic disorder; context-dependent functions | Single-function design paradigms insufficient for complex biological contexts |
| Stability-Function Tradeoffs | Destabilizing functional features in active sites | Requires large structural frameworks for thermodynamic compensation |
Computational Architectures for Novel Function Exploration
Machine Learning-Mediated Design
ML-based approaches, particularly deep learning architectures, have revolutionized protein structure prediction and are increasingly applied to design. These methods leverage patterns learned from existing protein databases to generate novel sequences and structures.
AlphaFold Architecture and Evolution: The AlphaFold system represents a landmark in protein structure prediction, with AlphaFold 2 achieving atomic accuracy competitive with experimental methods in many cases [9]. Its architecture incorporates novel neural network components including the Evoformer block that processes multiple sequence alignments (MSAs) and pairwise features, and a structure module that generates explicit 3D coordinates. The system demonstrated median backbone accuracy of 0.96 Å in CASP14, dramatically outperforming competing methods [9].
AlphaFold 3 Advancements: The recently introduced AlphaFold 3 incorporates a substantially updated diffusion-based architecture capable of predicting joint structures of complexes including proteins, nucleic acids, small molecules, ions, and modified residues [83]. Key innovations include:
- Replacement of the Evoformer with a simpler Pairformer module
- Direct prediction of raw atom coordinates using a diffusion module
- Elimination of stereochemical losses through multiscale diffusion
- Cross-distillation to reduce hallucination in unstructured regions
This architecture demonstrates substantially improved accuracy for protein-ligand interactions compared to state-of-the-art docking tools, and much higher accuracy for protein-nucleic acid interactions compared to nucleic-acid-specific predictors [83].
Protein Language Models: Methods like ESMFold leverage protein language models trained on millions of sequences to predict structures without explicit multiple sequence alignments [40]. For sequences with fewer homologs, these models can outperform MSA-dependent methods, suggesting they have learned fundamental principles of protein folding from sequence statistics alone [40].
Evolutionary Algorithm Approaches
Evolutionary Algorithms Simulating Molecular Evolution (EASME) represent a fundamentally different approach that mimics natural evolutionary processes to explore sequence space [81].
Core EASME Methodology: EASME employs evolutionary algorithms with DNA string representations, biologically accurate molecular evolution, and bioinformatics-informed fitness functions. Unlike ML approaches that primarily interpolate within known sequence space, EASME aims to expand beyond natural templates by simulating evolutionary processes [81].
Operational Modes: EASME can operate in two primary modes:
- "Unknown to Known": Evolves random sequences toward a known consensus sequence, effectively reconstructing extinct evolutionary intermediates
- "Known to Unknown": Forward-evolves known sequences toward desired phenotypic characteristics, functioning as a "fast forward" button on evolution [81]
Advantages for Novel Function Discovery: Proponents argue that EASME holds unique advantages for understanding the "why" behind protein function, not just the "what." The method can produce human-comprehensible design rules and potentially explore functional regions discontinuous from natural sequence space [81].
Hybrid and Integrated Approaches
Many successful design strategies combine evolutionary constraints with atomistic calculations. These approaches use evolutionary information to mitigate risks of misfolding and aggregation, focusing atomistic design on sequence subspaces highly enriched for functional solutions [82].
Evolution-Guided Atomistic Design: This methodology involves assembling backbone fragments from natural proteins followed by sequence design biased toward mutations observed in natural homologs. This preserves critical buried hydrogen bond networks often eliminated by purely physical design calculations [82]. The approach has successfully generated functional antibodies, enzymes, and protein-protein interactions with dozens of mutations from any natural protein while maintaining structural accuracy and function [82].
Benchmarking EA vs. ML Approaches
Direct comparison between EA and ML approaches reveals distinct strengths and limitations for different aspects of novel function discovery.
Table 2: Comparative Performance of EA vs. ML Protein Design Strategies
| Performance Metric | ML-Based Approaches | EA-Based Approaches | Specialized Hybrid Methods |
|---|---|---|---|
| Structure Prediction Accuracy | High (AF3: atomic accuracy for monomers/complexes) [83] | Limited (dependent on accurate fitness proxies) | Moderate to High (leverages evolutionary constraints) [82] |
| Novel Sequence Generation | Extrapolation within sequence space | Exploration beyond natural templates [81] | Guided exploration near functional regions |
| Computational Efficiency | High resource requirements for training | Variable (depends on fitness evaluation complexity) | Moderate to High |
| Interpretability | Low ("black box" models) | High (human-comprehensible rules) [81] | Moderate |
| Experimental Success Rate | Improving (especially for monomers) | Proof-of-concept established [81] | Demonstrated for antibodies, enzymes [82] |
| Handling Multi-molecule Complexes | Strong (AF3 handles proteins, nucleic acids, ligands) [83] | Limited to specified interaction networks | Limited to protein-protein interactions |
Performance Across Functional Classes
Both approaches show varying success depending on the functional class being targeted:
Enzyme Design: ML approaches benefit from large catalytic site databases but struggle with subtle mechanistic differences in superfamilies like the enolase superfamily, where similar structures catalyze different reactions [84]. EA approaches can potentially explore alternative mechanistic solutions but require accurate fitness functions representing catalytic efficiency.
Binding Interface Design: ML methods like AlphaFold Multimer and AlphaFold 3 show high accuracy for protein-protein interfaces [85] [83]. EA approaches have demonstrated success in designing specific interactions, such as toxin-antidote pairs in Wolbachia [81].
Therapeutic Protein Design: ML approaches dominate antibody and miniprotein design, with recent successes in designing oral therapeutics like Th17 antagonist miniproteins [86]. EA approaches offer potential for exploring non-immunogenic sequences distant from natural human proteins.
Experimental Protocols and Methodologies
ML-Based Design Workflow
Data Curation and Preprocessing:
- Collect diverse protein structures from PDB (≈200,000 structures)
- Generate multiple sequence alignments using MMSeq2 or similar tools
- Annotate functional sites, ligand interactions, and structural features
Model Training Protocol:
- Implement geometric and physical constraints as loss functions
- Use iterative refinement through recycling (AF2) or diffusion (AF3)
- Train with self-distillation on model predictions to expand structural diversity
Validation and Selection:
- Use pLDDT and PAE metrics from AlphaFold to assess prediction confidence
- Employ structural clustering to identify diverse solutions
- Filter designs by structural plausibility metrics (steric clashes, bond geometry)
EA-Based Design Workflow
Fitness Function Development:
- Define biophysical objectives (stability, binding affinity, catalytic efficiency)
- Incorporate evolutionary constraints from natural homologs
- Implement multi-objective optimization for conflicting design goals
Evolutionary Operations:
- Apply mutation operators with rates reflecting molecular evolution patterns
- Implement recombination between promising variants
- Use elitism selection to preserve high-performing solutions
Validation and Iteration:
- Use molecular dynamics to assess stability of designed proteins
- Employ docking simulations for interaction designs
- Implement experimental feedback to refine fitness functions
Table 3: Essential Research Resources for Novel Protein Design
| Resource Category | Specific Tools/Platforms | Primary Function | Access Method |
|---|---|---|---|
| Structure Prediction | AlphaFold 2/3, RoseTTAFold, ESMFold | Protein structure prediction from sequence | ColabFold server, local installation |
| Evolutionary Analysis | HMMER, MMseqs2, Clustal Omega | Multiple sequence alignment, homology detection | Web servers, command line |
| Protein Design Suites | Rosetta, ProteinMPNN, RFdiffusion | De novo protein design, sequence optimization | Academic licenses, web servers |
| Quality Assessment | MolProbity, QMEAN, VoroMQA | Structure validation, model quality estimation | Web servers, standalone packages |
| Specialized Databases | AFDB (>214M models), PDB, DisProt, MoonProt | Structural templates, functional annotations | Publicly accessible websites |
| Molecular Visualization | PyMOL, ChimeraX, UCSF Chimera | Structure analysis, figure generation | Academic licenses, open source |
Future Directions and Outstanding Challenges
The field of novel protein design continues to evolve rapidly, with several promising directions emerging:
Integration of Physical Principles: Both ML and EA approaches increasingly incorporate physical and biological knowledge about protein structure. ML models like AlphaFold embed physical constraints directly into their architecture [9], while EA approaches use molecular dynamics simulations as fitness proxies [81].
Multimodal Biomolecular Design: AlphaFold 3's capability to handle proteins, nucleic acids, ligands, and modifications points toward truly integrated biomolecular design [83]. This creates opportunities for designing complete molecular machines rather than isolated components.
Experimental Design Automation: High-throughput experimental validation is creating feedback loops that improve computational methods. The decreasing cost of DNA synthesis and gene assembly enables larger-scale testing of designed proteins.
Explainable AI for Protein Design: As noted in benchmarking studies, a key advantage of EA approaches is their human-comprehensible decision processes [81]. Future ML developments may incorporate explainable AI components to make design rules more transparent.
Despite significant progress, substantial challenges remain. Predicting functions that lack clear structural correlates, designing allosteric regulation, and creating proteins with multiple specific functions continue to challenge both EA and ML approaches. The integration of these complementary methodologies represents the most promising path toward truly novel functional landscapes beyond natural templates.
Rigorous Benchmarking: Validating and Comparing EA and ML Model Performance
The revolutionary accuracy of deep learning systems like AlphaFold2 in predicting protein structures from amino acid sequences has fundamentally transformed structural biology [40] [9]. However, the rapid emergence of multiple machine learning (ML)-based prediction tools necessitates rigorous, standardized benchmarking to guide researchers, scientists, and drug development professionals in selecting and applying these methods appropriately. Establishing common ground for comparison is especially critical for a broader thesis contrasting evolutionary-based (EA) and machine learning (ML) approaches to protein folding. EA methods traditionally leverage evolutionary information from Multiple Sequence Alignments (MSAs) of homologous proteins to infer structural constraints. In contrast, newer ML models, while still utilizing MSAs, employ sophisticated neural networks to learn the complex mapping from sequence to structure, with some language model-based approaches like ESMFold even bypassing the need for explicit MSAs [40]. Benchmarking these paradigms requires a consistent framework evaluating not just prediction accuracy, but also computational efficiency and resource consumption—key factors for practical deployment in both academic and industrial settings. This guide details the core metrics essential for this task: pLDDT for assessing local prediction confidence, running time for practical feasibility, and memory usage for hardware requirements.
Core Benchmarking Metrics Explained
pLDDT (Predicted Local Distance Difference Test)
The pLDDT is a per-residue local confidence score estimated by AlphaFold and other models, scaled from 0 to 100 [87]. It is based on the local distance difference test (lDDT), a superposition-free score that evaluates the correctness of a predicted structure by checking the conservation of inter-atomic distances within a local neighborhood [87] [88].
Interpreting pLDDT Scores
The pLDDT score provides a reliable estimate of local model quality. The following table details the standard interpretation of its value ranges:
Table 1: Interpretation of pLDDT Confidence Scores
| pLDDT Range | Confidence Level | Typical Structural Interpretation |
|---|---|---|
| > 90 | Very high | Both backbone and side chains are typically predicted with high accuracy. |
| 70 - 90 | Confident | Usually a correct backbone prediction, but with potential misplacement of some side chains. |
| 50 - 70 | Low | The prediction should be treated with caution; the region may be unstructured or poorly predicted. |
| < 50 | Very low | The region is likely highly flexible or intrinsically disordered. These regions are unlikely to adopt a well-defined structure [87]. |
It is crucial to understand that pLDDT is a measure of local confidence. A high pLDDT for all domains of a protein does not necessarily indicate confidence in their relative positions or orientations, as the score does not measure confidence at such large scales [87].
pLDDT as a Proxy for Flexibility
Although designed as a confidence metric, pLDDT has shown a significant correlation with protein flexibility. Large-scale studies comparing pLDDT to flexibility metrics derived from Molecular Dynamics (MD) simulations and NMR ensembles have found that regions with low pLDDT often correspond to flexible regions in the protein [88]. However, this correlation is not perfect. AlphaFold's pLDDT can struggle to capture flexibility in the presence of interacting partners and may in some cases predict conditionally folded states of intrinsically disordered regions (IDRs) that only become structured when bound to a partner [87] [88]. For example, AlphaFold2 predicts the eukaryotic translation initiation factor 4E-binding protein 2 (4E-BP2) with a high-confidence helical structure, which in nature it only adopts in its bound state [87].
Running Time and Memory Usage
Beyond accuracy, the practical utility of a protein folding tool is determined by its computational demands: running time and memory usage. These metrics determine the hardware requirements, cost, and scalability of predictions, especially for large proteins or high-throughput applications.
Running time is the total time a model takes to predict a structure from a protein sequence, typically measured in seconds. Memory usage is often broken down into CPU (system) memory and GPU memory, both measured in Gigabytes (GB). These metrics are highly dependent on the length of the input protein sequence, with longer sequences requiring exponentially more computation and memory [8] [89].
Quantitative Benchmarking of Major Protein Folding Tools
A comparative benchmark of three leading ML methods—AlphaFold, OmegaFold, and ESMFold—highlights the trade-offs between accuracy, speed, and resource consumption. The following data, collected on a g5.2xlarge A10 GPU, provides a direct comparison across key metrics [8].
Table 2: Benchmarking Results for Protein Folding Tools (A10 GPU)
| Sequence Length | Tool | Running Time (s) | pLDDT | CPU Memory (GB) | GPU Memory (GB) |
|---|---|---|---|---|---|
| 50 | ESMFold | 1 | 0.84 | 13 | 16 |
| OmegaFold | 3.66 | 0.86 | 10 | 6 | |
| AlphaFold | 45 | 0.89 | 10 | 10 | |
| 400 | ESMFold | 20 | 0.93 | 13 | 18 |
| OmegaFold | 110 | 0.76 | 10 | 10 | |
| AlphaFold | 210 | 0.82 | 10 | 10 | |
| 800 | ESMFold | 125 | 0.66 | 13 | 20 |
| OmegaFold | 1425 | 0.53 | 10 | 11 | |
| AlphaFold | 810 | 0.54 | 10 | 10 | |
| 1600 | ESMFold | Failed (OOM) | Failed | Failed | 24 |
| OmegaFold | Failed (>6000) | Failed | Failed | 17 | |
| AlphaFold | 2800 | 0.41 | 10 | 10 |
OOM = Out of Memory. Best values for each metric and length are highlighted.
Performance Analysis and Tool Selection
The data reveals distinct performance profiles suitable for different applications:
- ESMFold demonstrates superior speed, particularly on shorter sequences (50 and 100 residues), but is less accurate than OmegaFold or AlphaFold and uses the most memory, which can lead to out-of-memory errors on long sequences [8].
- OmegaFold shows a strong balance of speed, accuracy, and resource efficiency for shorter sequences (up to 400 residues). It achieves high pLDDT scores while using less CPU and GPU memory than ESMFold, making it a cost-effective and reliable choice for shorter proteins [8].
- AlphaFold (via ColabFold) is generally the slowest but often the most accurate, especially on shorter sequences. A key advantage is its consistent and efficient GPU memory usage across varying sequence lengths, making it the most robust tool for predicting very long sequences (e.g., 1600 residues) where other models fail [8].
Experimental Protocols for Benchmarking
To ensure reproducible and fair comparisons, follow these detailed methodologies.
Protocol for Measuring Running Time and Memory
Objective: To quantitatively measure the computational resource requirements of protein structure prediction tools.
Hardware/Software: Standardized computing node (e.g., cloud instance with A10 GPU), latest versions of target software (AlphaFold/ColabFold, OmegaFold, ESMFold), system monitoring tools (e.g., time, nvidia-smi).
Procedure:
- Sequence Selection: Curate a set of protein sequences of varying, defined lengths (e.g., 50, 100, 200, 400, 800, 1600 residues).
- Environment Isolation: Run each prediction tool in a clean, isolated environment (e.g., Docker container) to prevent library conflicts and ensure accurate resource measurement.
- Execution and Timing: For each sequence and tool, execute the prediction command and record the wall-clock time from initiation to completion.
- Memory Profiling: Use profiling tools to record the peak CPU and GPU memory allocated during the inference process. The PyTorch profiler is effective for this [89].
- Data Collection: Repeat the measurement multiple times to account for system variability and report the average values.
Protocol for Evaluating pLDDT and Model Accuracy
Objective: To assess the perceived confidence (pLDDT) and ground-truth accuracy of predicted models. Data Sources: Protein sequences for prediction, experimentally determined structures (e.g., from the PDB) for validation. Procedure:
- Model Prediction: Generate 3D structure models for the target sequences using the tools being benchmarked.
- pLDDT Extraction: Parse the output files (e.g., PDB or mmCIF) to extract the per-residue and average pLDDT scores.
- Accuracy Validation (Optional but Recommended):
- Superposition-based metrics: Calculate the Root-Mean-Square Deviation (RMSD) or the Template Modeling Score (TM-score) between the predicted model and the experimental structure after optimal superposition [90] [9]. TM-score is less sensitive to global outliers than RMSD.
- Local accuracy verification: Use the local Distance Difference Test (lDDT) to evaluate the local geometric accuracy of the model against the experimental structure, which validates the meaning of the pLDDT score [87] [9].
Advanced Optimization and Emerging Challenges
Optimization of Computational Workloads
The high computational cost of models like AlphaFold has spurred development of optimization methods. Dynamic Axial Parallelism (DAP) is a strategy that distributes large input tensors (like the MSA and pair representations) across multiple GPUs, significantly improving inference latency for both forward and backward passes [89]. Benchmarking shows that a 2-GPU configuration can offer over 100% strong scaling efficiency for the forward pass [89].
Another key innovation is AutoChunk, an automated algorithm designed to reduce peak memory consumption during inference. It works by intelligently partitioning large computational operations into smaller "chunks," trading a slight increase in computation time for a substantial reduction in memory. Experiments show AutoChunk can reduce memory usage by over 80% for long sequences, enabling the prediction of proteins that would otherwise cause out-of-memory errors [89].
Challenges in Docking and Complex Prediction
While pLDDT is reliable for single-chain confidence, benchmarking becomes more complex for protein-protein interactions. AlphaFold-multimer (AFm) has limitations, particularly when proteins undergo significant conformational changes upon binding [91]. In these cases, AFm's success rate can drop significantly, especially for challenging targets like antibody-antigen complexes [91].
Hybrid approaches that combine deep learning with physics-based methods are emerging as a powerful solution. For example, the AlphaRED pipeline uses AFm to generate initial structural templates and then refines them using a physics-based replica exchange docking algorithm (ReplicaDock 2.0). This approach can rescue failed AFm predictions, generating acceptable-quality models for 63% of benchmark targets where AFm alone struggled [91]. This underscores that no single metric or tool is sufficient for all scenarios, and benchmarks must be tailored to the biological question.
Table 3: Key Computational Tools for Protein Structure Benchmarking
| Tool / Resource | Type | Primary Function | Reference/Source |
|---|---|---|---|
| AlphaFold / ColabFold | Software | High-accuracy protein structure prediction using MSAs and templates. | [DeepMind], [github.com/sokrypton/ColabFold] [40] [9] |
| ESMFold | Software | Fast protein structure prediction using a protein language model (no explicit MSA needed). | [github.com/facebookresearch/esm] [8] [40] |
| OmegaFold | Software | Accurate protein structure prediction, performs well on short sequences. | [github.com/HeliXonProtein/OmegaFold] [8] |
| AlphaFold Database (AFDB) | Database | Repository of pre-computed AlphaFold predictions for multiple proteomes. | [alphafold.ebi.ac.uk] [40] |
| Protein Data Bank (PDB) | Database | Repository of experimentally determined 3D structures of proteins and nucleic acids. | [rcsb.org] [40] |
| ReplicaDock 2.0 | Software | Physics-based replica exchange docking for sampling protein-protein complexes. | [github.com/Graylab/ReplicaDock2] [91] |
| Foldseck | Software | Rapid search of structural similarities in large databases. | [github.com/soedinglab/foldseck] [40] |
Establishing robust benchmarks based on pLDDT, running time, and memory usage is not an academic exercise but a practical necessity for navigating the modern landscape of protein structure prediction. As this guide illustrates, the choice between state-of-the-art tools involves inherent trade-offs. AlphaFold often leads in accuracy, ESMFold in raw speed, and OmegaFold offers a compelling balance for shorter sequences. The field is moving beyond single-metric assessments towards integrated pipelines, such as AlphaRED, which combine the pattern recognition of ML with the physical rigor of traditional biophysical methods. For researchers embarking on a thesis comparing EA and ML paradigms, these benchmarks provide the essential, objective foundation required to critically evaluate the strengths, limitations, and optimal application domains of each approach, ultimately accelerating progress in structural biology and drug discovery.
The field of protein structure prediction has been revolutionized by deep learning, transitioning from a long-standing challenge to a routinely applied technology. Within this landscape, AlphaFold2, ESMFold, and OmegaFold represent three prominent yet methodologically distinct approaches. This analysis provides a technical comparison of these tools, framing them within a broader benchmarking context of evolutionary analysis (EA) against machine learning (ML)-first strategies. Understanding their core architectures, performance characteristics, and computational demands is crucial for researchers and drug development professionals to select the optimal tool for a given application, balancing precision, speed, and resource constraints.
Methodological Foundations & Core Architectures
The fundamental divergence between these models lies in their use of evolutionary information and their underlying neural network architectures.
AlphaFold2 (EA-Centric): AlphaFold2 relies heavily on evolutionary information derived from Multiple Sequence Alignments (MSAs) of homologous proteins. Its core innovation is the Evoformer block—a complex neural network module that processes the MSA and residue-pair representations through a series of attention mechanisms and triangle multiplicative updates. This allows the network to reason about spatial and evolutionary relationships simultaneously [9]. The processed information is then passed to a structure module that iteratively refines the 3D atomic coordinates [9].
ESMFold (ML-First): ESMFold adopts a starkly different, alignment-free approach. It is built upon a protein language model (PLM) that is pre-trained on millions of protein sequences. This PLM generates informative sequence embeddings that implicitly capture evolutionary constraints without the need for explicit MSA construction. These embeddings are fed directly into a folding module to predict the 3D structure, resulting in a dramatic increase in inference speed [92] [93].
OmegaFold (Hybrid ML): OmegaFold represents a hybrid pathway. It also eliminates the need for MSAs but uses a different architecture. It combines a protein language model with a geometry-guided transformer model called the Geoformer. This model learns both single-residue and pairwise-residue embeddings, which are then used to build the structure based on geometric principles [94] [93].
The diagram below illustrates the distinct input and information flow for each of these three core architectures.
Performance Benchmarking and Quantitative Analysis
Systematic benchmarking on a large set of 1,336 protein chains from the PDB (deposited between 2022 and 2024, ensuring no training data overlap) provides a clear hierarchy of accuracy among the three tools.
Table 1: Overall Accuracy Metrics on Recent PDB Structures
| Method | Median TM-score | Median RMSD (Å) | Key Strength |
|---|---|---|---|
| AlphaFold2 | 0.96 [95] [96] | 1.30 Å [95] [96] | Highest overall accuracy |
| ESMFold | 0.95 [95] [96] | 1.74 Å [95] [96] | Excellent speed-accuracy trade-off |
| OmegaFold | 0.93 [95] [96] | 1.98 Å [95] [96] | Strong on orphan proteins/antibodies [94] |
While AlphaFold2 achieves the highest median accuracy, the performance gap is often negligible for many proteins, suggesting that faster models can be sufficient for large-scale screening [95] [96].
Computational Efficiency and Resource Requirements
A critical trade-off exists between accuracy and computational cost. The MSA generation step required by AlphaFold2 is computationally expensive, making it significantly slower than its alignment-free counterparts.
Table 2: Computational Performance Comparison (A10 GPU)
| Method | Inference Speed (Seconds, 400 resid.) | CPU Memory | GPU Memory | Architecture |
|---|---|---|---|---|
| AlphaFold2 | ~210 sec [8] | ~10 GB [8] | ~10 GB [8] | MSA-Dependent |
| ESMFold | ~20 sec [8] | ~13 GB [8] | ~18 GB [8] | Single-Sequence PLM |
| OmegaFold | ~110 sec [8] | ~10 GB [8] | ~10 GB [8] | Single-Sequence PLM + Geoformer |
ESMFold is typically the fastest, being 10-30 times faster than AlphaFold2 in many practical scenarios [95] [93]. OmegaFold strikes a balance, often faster than AlphaFold2 but slower than ESMFold. For longer sequences (>800 residues), memory can become a limiting factor for all models [8].
Experimental Protocols and Practical Application
A Standardized Benchmarking Workflow
To objectively compare these tools, researchers can implement the following workflow, which mirrors the methodology used in large-scale evaluations [95] [92] [93].
- Test Set Curation: Compile a set of protein sequences with experimentally determined structures (e.g., from the PDB) that were released after the training cut-off dates of the models to ensure a blind test. A size of over 1,000 chains is recommended for statistical power [95].
- Structure Prediction: Run each target sequence through AlphaFold2 (or ColabFold), ESMFold, and OmegaFold using standardized hardware and software environments.
- Structure Comparison: Superimpose the predicted model (P) onto the experimental structure (E) using tools like US-align or TM-align.
- Metric Calculation:
- TM-score: Measures global fold similarity. A score >0.8 indicates the same correct fold, with 1.0 being a perfect match [92].
- RMSD: Measures the average distance between equivalent atoms after superposition, in Angstroms (Å). Lower values indicate higher local accuracy [92].
- pLDDT: The model's internal confidence score per residue. Scores >90 are high confidence, while scores <50 are low confidence [92] [9].
- Analysis: Aggregate scores across the entire test set to calculate median TM-scores and RMSD values, as shown in Table 1.
The Scientist's Toolkit: Key Research Reagents
The following table details essential computational "reagents" and their functions for conducting such an analysis.
Table 3: Essential Research Reagents for Protein Structure Prediction Benchmarking
| Research Reagent | Function & Purpose | Example/Note |
|---|---|---|
| Protein Data Bank (PDB) | Source of "ground truth" experimental structures for benchmark set creation and validation [92]. | Use recently deposited structures to avoid data leakage [95]. |
| ColabFold | A popular, accessible implementation of AlphaFold2 that speeds up MSA generation with MMseqs2 [92]. | Lowers the barrier to running AlphaFold2. |
| Robetta/AlphaFold Server | Web servers for protein structure prediction; useful for individual predictions without local installation [92]. | |
| CAMEO & CASP Datasets | Standardized, continuous blind tests for objectively evaluating prediction accuracy [92] [93]. | Used for independent validation of model performance. |
| TM-score & RMSD Scripts | Computational metrics to quantitatively compare a predicted structure against an experimental reference [92]. | Essential for objective performance benchmarking. |
| ProtBert | A protein language model used to generate sequence embeddings that can help predict when AlphaFold2's added accuracy is necessary [95]. | Useful for developing meta-prediction classifiers. |
Discussion and Strategic Implementation
When to Use Which Model: A Decision Framework
The choice between these models is not one of finding a single "best" tool, but of selecting the right tool for the specific research goal and context. The following diagram outlines a decision framework to guide researchers.
Limitations and Advanced Considerations
While these tools are powerful, understanding their limitations is vital for rigorous research.
- Conformational Dynamics: A significant weakness of all three models is their bias toward predicting single, thermodynamically stable conformations. They generally struggle to model the intrinsic conformational ensembles and dynamics of proteins, such as alternative allosteric states or fold-switching proteins [94].
- Physical Realism: Recent studies questioning co-folding models like AlphaFold3 have highlighted that their impressive performance may sometimes stem from pattern recognition rather than a deep understanding of physical principles like steric clashes and electrostatic interactions [97].
- Dependence on Training Data: The models are susceptible to "structural memorization" of proteins in their training set, which can limit their ability to generalize to truly novel folds or conformations [94] [97].
To address the limitation of conformational dynamics, ensemble methods like the FiveFold approach have been developed. This methodology runs multiple prediction algorithms (including AlphaFold2, ESMFold, and OmegaFold) on the same target and integrates the results to generate a spectrum of plausible conformations, providing a more realistic view of a protein's dynamic landscape, which is especially useful for studying intrinsically disordered proteins and for drug discovery [19].
The comparative analysis of AlphaFold2, ESMFold, and OmegaFold reveals a field shaped by the complementary strengths of evolutionary analysis and machine learning-first strategies. AlphaFold2 remains the gold standard for maximum prediction accuracy when computational resources and time are not constraints. In contrast, ESMFold and OmegaFold offer transformative speed and efficiency for high-throughput applications, with accuracy that is sufficient for many practical purposes.
For the researcher, the decision is strategic. The choice depends on the specific balance required between precision, scale, and resource allocation. As the field evolves, the integration of these tools into ensemble methods and the ongoing development of models that better capture protein dynamics and physical principles will further expand the frontiers of structural bioinformatics and drug discovery.
The emergence of deep learning has revolutionized protein structure prediction, with AlphaFold2 setting a benchmark for accuracy. However, for specific applications involving short protein sequences, alternative models offer distinct advantages. This whitepaper examines the performance of three leading deep learning methods—OmegaFold, ESMFold, and AlphaFold—in predicting structures for short sequences. Through quantitative benchmarking of runtime, accuracy, and computational resource utilization, we demonstrate that OmegaFold achieves a superior balance of prediction accuracy and operational efficiency for sequences under 400 residues. Its unique MSA-free architecture, leveraging a protein language model and geometry-inspired transformer, enables high-resolution de novo prediction while consuming significantly less memory, making it an ideal candidate for production environments and the study of orphan proteins and antibodies.
The protein folding problem—predicting a protein's three-dimensional structure from its amino acid sequence—has been a central challenge in computational structural biology for decades [98]. The advent of deep learning has precipitated a paradigm shift, with methods like AlphaFold2 (AF2) achieving accuracy competitive with experimental structures [40] [99]. These tools now provide invaluable models for research, from rational drug development to mutation analysis [99].
Despite these advances, the computational cost and specific architectural requirements of these models can be prohibitive for certain applications. AlphaFold2 and RoseTTAFold, for instance, rely heavily on Multiple Sequence Alignments (MSAs) to extract co-evolutionary information, which is computationally expensive to generate and may be unavailable for proteins with few homologs [100] [18]. In response, a new generation of protein language model (pLM)-based methods, including ESMFold and OmegaFold, has emerged. These models predict structure from a single sequence, offering substantial speed improvements and applicability to "orphan" proteins lacking evolutionary context [101] [102] [100].
This whitepaper frames these developments within a broader thesis on benchmarking evolutionary analysis (EA) versus machine learning (ML) for protein folding. While MSA-dependent methods like AF2 leverage explicit evolutionary analysis, pLM-based methods implicitly learn evolutionary constraints from vast sequence databases during pre-training [101]. We conduct a focused performance analysis on short protein sequences, a critical use-case in drug discovery (e.g., peptides, antibodies), where computational efficiency and accuracy are paramount. Benchmarking data reveals that OmegaFold consistently delivers an optimal trade-off, providing high-accuracy predictions with remarkable resource efficiency [8].
Benchmarking Performance on Short Sequences
To quantitatively evaluate the practical performance of these tools, we benchmarked three leading models—ESMFold, OmegaFold, and AlphaFold (via ColabFold)—on an A10 GPU, measuring running time, predictive accuracy (using pLDDT score), and memory consumption across varying sequence lengths [8].
Quantitative Performance Metrics
The following table summarizes the key performance metrics for sequences up to 400 residues in length, a critical range for many targeted applications.
Table 1: Benchmarking Results for Short Sequences (Lengths 50-400) [8]
| Sequence Length | Model | Running Time (seconds) | PLDDT Score | CPU Memory (GB) | GPU Memory (GB) |
|---|---|---|---|---|---|
| 50 | ESMFold | 1 | 0.84 | 13 | 16 |
| OmegaFold | 3.66 | 0.86 | 10 | 6 | |
| ColabFold | 45 | 0.89 | 10 | 10 | |
| 100 | ESMFold | 1 | 0.30 | 13 | 16 |
| OmegaFold | 7.42 | 0.39 | 10 | 7 | |
| ColabFold | 55 | 0.38 | 10 | 10 | |
| 200 | ESMFold | 4 | 0.77 | 13 | 16 |
| OmegaFold | 34.07 | 0.65 | 10 | 8.5 | |
| ColabFold | 91 | 0.55 | 10 | 10 | |
| 400 | ESMFold | 20 | 0.93 | 13 | 18 |
| OmegaFold | 110 | 0.76 | 10 | 10 | |
| ColabFold | 210 | 0.82 | 10 | 10 |
Analysis of Benchmarking Results
The data reveals a clear performance hierarchy for short sequences:
- Running Time: ESMFold is the fastest method for sequences up to 200 residues, but its speed advantage diminishes at length 400. OmegaFold is significantly faster than ColabFold (up to 12x for 50-residue sequences) while being only slightly slower than ESMFold for shorter lengths [8].
- Predictive Accuracy (pLDDT): For the shortest sequences (50 and 100 residues), OmegaFold achieves the highest or competitive pLDDT scores, indicating superior reliability. While ESMFold can achieve high scores on some longer targets, its performance is highly variable, as seen in the low score at length 100 [8].
- Resource Efficiency: OmegaFold demonstrates superior memory management, consistently using less CPU and GPU memory than ESMFold and similar or less GPU memory than ColabFold. This efficiency is critical for deployment in resource-constrained environments or for scaling to high-throughput prediction tasks [8].
The benchmark concludes that "OmegaFold's balance of speed, accuracy, and resource efficiency makes it an excellent choice for public-serving platforms, particularly for protein sequences with lengths up to 400" [8].
Architectural Foundations
The performance disparities are a direct consequence of the underlying architectural philosophies of each model.
MSA-Dependent vs. MSA-Free Paradigms
- AlphaFold2 (ColabFold): An MSA-dependent model, AF2 relies on generating deep MSAs to infer co-evolutionary signals, which are processed through its Evoformer and structural modules to produce a structure [40] [100]. This process is computationally intensive but provides high accuracy for sequences with many homologs.
- ESMFold and OmegaFold (pLM-based): These are MSA-free models. They use protein language models (pLMs) pre-trained on millions of sequences to generate per-residue and residue-pair representations that implicitly capture evolutionary and structural constraints, bypassing the need for explicit MSA generation [100] [102].
OmegaFold's Technical Innovation
OmegaFold incorporates a novel combination of a protein language model (OmegaPLM) and a geometry-inspired transformer model trained on protein structures [102]. This architecture allows it to perform high-resolution de novo protein structure prediction directly from a primary sequence. The integration of the geometry-inspired transformer enables the model to reason more effectively about spatial relationships, contributing to its high accuracy even without MSAs. This makes it particularly effective for proteins with limited or no homologous sequences, such as orphan proteins and fast-evolving antibodies [102].
Experimental Protocols for Benchmarking
To ensure reproducibility and rigorous comparison, the following experimental protocol outlines the key steps for benchmarking protein structure prediction tools.
Diagram Title: Protein Structure Prediction Benchmarking Workflow
Detailed Methodologies
Dataset Selection: Curate a set of protein sequences with known experimental structures (e.g., from the PDB) to serve as ground truth. The set should include sequences of varying lengths, with a focus on the short sequence range (<400 residues). To test model generalization, include targets from the CASP Free-Modeling (FM) domains, where no homologous structures are available [99] [103].
Computational Environment Configuration: Standardize the hardware and software environment to ensure a fair comparison. The benchmark should be run on a machine with a dedicated GPU (e.g., NVIDIA A10). Use containerization (Docker/Singularity) to ensure consistent software versions and dependencies for each model. For ColabFold, use the standard implementation with MMseqs2 for MSA generation [8] [103].
Model Execution and Data Collection:
- Running Time: Measure the wall-clock time from job submission to completion of the structure prediction. This includes MSA generation time for ColabFold.
- Accuracy Metrics: Calculate the pLDDT score provided natively by each model. Optionally, for sequences with known experimental structures, compute the TM-score or RMSD to evaluate global fold accuracy.
- Resource Utilization: Monitor CPU and GPU memory usage throughout the prediction process, recording the peak consumption.
The Scientist's Toolkit
Table 2: Essential Research Reagents and Computational Tools
| Item | Function in Research | Application Context |
|---|---|---|
| GPU (A10) | Provides accelerated parallel computing for deep learning model inference. | Essential for running all featured structure prediction models in a time-efficient manner [8]. |
| DeepMSA2 | A hierarchical pipeline for constructing high-quality Multiple Sequence Alignments (MSAs) from genomic and metagenomic databases. | Used to generate optimized input MSAs for MSA-dependent models like AlphaFold2, improving tertiary and quaternary structure prediction [103]. |
| ColabFold | A streamlined, accessible implementation of AlphaFold2 that uses MMseqs2 for fast MSA generation and runs via Google Colab notebooks. | Lowers the barrier to entry for running AlphaFold2, suitable for users without local high-performance computing resources [40]. |
| FiveFold Framework | An ensemble method that combines predictions from five different algorithms (AF2, RoseTTAFold, OmegaFold, ESMFold, EMBER3D) to model conformational diversity. | Used for advanced studies of intrinsically disordered proteins and conformational ensembles, beyond single-structure prediction [19]. |
| Foldseck | A tool for rapid structural similarity searches and alignments in large protein structure databases. | Enables efficient comparison of predicted models against existing structures in the PDB or AlphaFold Database for functional annotation [40]. |
Discussion and Implications
The benchmarking data firmly establishes OmegaFold's utility for short-sequence protein structure prediction. Its operational advantages—notably lower memory footprint and competitive speed—coupled with robust accuracy make it exceptionally well-suited for specific research and development contexts.
Practical Applications and Use Cases
- High-Throughput Screening: In drug discovery pipelines where thousands of short peptides or protein fragments require rapid structural characterization, OmegaFold's speed and resource efficiency enable scalable deployment [8].
- Orphan Proteins and Antibody Design: For proteins with few evolutionary relatives, such as orphan proteins or fast-evolving viral antigens and antibodies, MSA-dependent methods struggle. OmegaFold's MSA-free architecture fills this critical gap, providing reliable structural insights where other tools fail [102] [100].
- Integrated Ensemble Approaches: The FiveFold methodology, which leverages multiple prediction algorithms including OmegaFold, demonstrates the value of using specialized tools as part of a consensus-building strategy. This approach better captures conformational diversity, which is crucial for understanding protein function and for structure-based drug design, particularly for disordered proteins [19].
Limitations and Future Directions
While OmegaFold excels with short sequences, its accuracy on longer, complex multidomain proteins may not surpass that of AlphaFold2, which benefits from deep MSAs [8] [100]. Furthermore, like other current deep learning models, OmegaFold is a structure predictor, not a simulator of the physical folding process. It struggles to capture the full energy landscape and conformational dynamics of proteins [18] [98].
Future developments will likely focus on integrating the strengths of both paradigms. This includes developing models that use pLM embeddings as a primary source of information while incorporating physical principles to improve the modeling of dynamics and interactions. Advances in MSA construction, as seen with DeepMSA2, will also continue to push the accuracy of MSA-dependent methods, ensuring both evolutionary analysis and machine learning remain vital to the future of structural bioinformatics [103].
Within the expanding toolkit of protein structure prediction, the choice of model is increasingly dictated by the specific application. This whitepaper, situated within a broader benchmarking effort, demonstrates that while AlphaFold2 remains the gold standard for general-purpose prediction, OmegaFold establishes a distinct niche. For short sequences under 400 residues, it delivers a superior balance of computational efficiency and predictive accuracy. Its MSA-free nature not only provides speed and resource advantages but also unlocks the structural modeling of orphan proteins and designed antibodies, thereby expanding the frontiers of accessible structural biology. As the field progresses, the integration of specialized, efficient tools like OmegaFold into consensus frameworks and drug discovery pipelines will be instrumental in tackling increasingly complex biological challenges.
The dominant paradigm in structural biology has long been the sequence-structure-function relationship, wherein a single amino acid sequence encodes one stable three-dimensional structure that determines its biological function. However, an emerging class of fold-switching proteins (also known as metamorphic proteins) challenges this central dogma by adopting two distinct sets of stable secondary and tertiary structures, transitioning between them in response to cellular stimuli [104]. These structural transitions modulate critical biological functions, including suppression of human innate immunity during SARS-CoV-2 infection, control of bacterial virulence gene expression, and maintenance of cyanobacterial circadian rhythms [104].
Despite revolutionary advances in machine learning (ML) for protein structure prediction, state-of-the-art algorithms including AlphaFold2, trRosetta, and EVCouplings systematically fail to predict these alternative conformations, typically predicting only a single fold for most known dual-folding proteins [104]. This limitation stems from a fundamental methodological divide: these ML approaches infer structure from co-evolutionary information in multiple sequence alignments (MSAs), potentially missing evolutionary signatures preserved for maintaining dual-fold capability [104] [40].
This technical guide examines the experimental confirmation of dual-fold coevolution, positioning Evolutionary Analysis (EA) methods as complementary to ML approaches in the broader context of protein folding benchmarking. We provide validation methodologies, quantitative performance assessments, and practical experimental protocols for researchers investigating protein metamorphosis.
Evolutionary Basis of Dual-Fold Coevolution
Evolutionary Selection of Fold-Switching Sequences
The failure of ML methods to predict alternative folds initially suggested two competing hypotheses: (1) fold-switching proteins are rare evolutionary byproducts not selected for dual conformations, or (2) both conformations are evolutionarily selected, but standard prediction strategies miss their signatures. Recent evidence strongly supports the second hypothesis, indicating that fold-switching sequences have been preserved by natural selection, implying their functionalities provide evolutionary advantage [104].
The discovery of widespread dual-fold coevolution demonstrates that both conformations of fold-switching proteins are evolutionarily selected. This finding has profound implications for protein structure prediction, suggesting that current ML methods may be fundamentally limited in their ability to capture metamorphic capability due to their reliance on evolutionary couplings that assume a single dominant fold [104].
Limitations of Current Structure Prediction Algorithms
Benchmarking studies reveal specific limitations in ML approaches for predicting non-standard protein behaviors:
- AlphaFold2 performance variability: While achieving remarkable accuracy for many single-fold proteins, AlphaFold2 predicts only one conformation for 92% of known dual-folding proteins, with 30% of these predictions likely not representing the lowest energy state [104].
- Peptide structure challenges: AlphaFold2 shows shortcomings in predicting Φ/Ψ angles, disulfide bond patterns, and correlation between low RMSD structures and low pLDDT rankings for peptides, particularly for mixed secondary structure soluble peptides [105].
- Method-specific strengths: OmegaFold demonstrates superiority for shorter sequences and cases with limited homologous sequences, while ESMFold excels in speed but may sacrifice some accuracy [8].
Table 1: Performance Benchmarking of ML Protein Structure Prediction Tools
| Method | Strengths | Limitations with Fold-Switching Proteins | Best Use Cases |
|---|---|---|---|
| AlphaFold2 | High accuracy for single-fold proteins, extensive database | Predicts only one fold for 92% of dual-fold proteins; 30% incorrect lowest energy state | Proteins with deep MSAs, single-fold prediction [104] [40] |
| OmegaFold | Accurate for short sequences, works without MSA | Limited benchmarking on fold-switching proteins | Short sequences (<400 AA), limited homology [8] [105] |
| ESMFold | Very fast prediction, no MSA required | Lower accuracy than AF2 for proteins with MSAs | High-throughput screening, proteins without homologs [8] [40] |
| RoseTTAFold | Approaches AF2 accuracy, different architecture | Similar limitations for alternative conformations | Alternative to AF2 with different implementation [105] |
The ACE Methodology: Experimental Framework for Validating Dual-Fold Coevolution
ACE Workflow and Implementation
The Alternative Contact Enhancement (ACE) approach was developed specifically to detect coevolutionary signatures for both conformations of fold-switching proteins. This methodology successfully revealed coevolution of amino acid pairs uniquely corresponding to both conformations in 56 out of 56 fold-switching proteins from distinct families tested [104].
Core Experimental Protocol
The ACE methodology employs a systematic approach to uncover dual-fold coevolution:
Input Preparation: A query sequence with two distinct experimentally determined structures serves as the starting point [104].
Multiple Sequence Alignment Generation:
- Generate a deep MSA using HMMER3.3.2 (jackhmmer) against UniRef90 with iterative search parameters:
-N 10 --incdomE 10E-20 --incE 10E-20[106]. - Prune the deep MSA to create successively shallower subfamily MSAs with sequences increasingly identical to the query using HHSUITE's QID filter [104] [106].
- Generate a deep MSA using HMMER3.3.2 (jackhmmer) against UniRef90 with iterative search parameters:
Coevolutionary Analysis:
Contact Prediction Integration:
- Combine predictions from both methods across all nested MSAs.
- Superimpose these predictions on a single contact map for comprehensive analysis [104].
Signal Enhancement and Filtering:
- Apply density-based scanning to remove noise from predicted contacts.
- Categorize contacts as: "Dominant fold," "Alternative fold," "Common" (shared by both folds), or "Unobserved" [104].
Quantitative Validation of Dual-Fold Coevolution
Experimental Performance Metrics
The ACE method demonstrates substantial improvement over standard coevolution analysis:
- Alternative conformation contacts: 201% mean increase (187% median) in correctly predicted contacts for alternative conformations [104].
- Overall accuracy: 111% mean increase (107% median) in total correctly predicted contacts across all 56 tested proteins [104].
- Noise suppression: Experimental unobserved contacts amplified significantly less (42% mean, 47% median increase) than alternative or correctly predicted contacts [104].
Table 2: Quantitative Performance of ACE vs. Standard Methods on 56 Fold-Switching Proteins
| Performance Metric | Standard Approach | ACE Enhancement | Improvement |
|---|---|---|---|
| Alternative Fold Contacts | Baseline | 201% mean increase | 187% median increase |
| Total Correct Contacts | Baseline | 111% mean increase | 107% median increase |
| Unobserved Contacts | Baseline | 42% mean increase | 47% median increase |
| Success Rate | Not applicable | 56/56 proteins | 100% |
Experimental Validation and Blind Prediction
The ACE methodology enabled two critical advances in protein structure prediction:
Prediction of unknown conformations: Using ACE-derived contacts, researchers successfully predicted two experimentally consistent conformations of a candidate NusG protein with <30% sequence identity to both of its PDB homologs [104].
Blind prediction pipeline: Development of a blind prediction pipeline for fold-switching proteins that correctly identified 13/56 known fold-switchers (23%) with a false-positive rate of 0/181 [104].
Research Reagent Solutions for Dual-Fold Validation
Table 3: Essential Research Reagents and Tools for Dual-Fold Coevolution Studies
| Reagent/Tool | Function/Application | Implementation Notes |
|---|---|---|
| ACE Software | Detects coevolutionary signatures for alternative folds | Python implementation; requires GREMLIN and MSA Transformer [106] |
| GREMLIN | Infers coevolved residue pairs using Markov Random Fields | Superior performance for contact prediction; converges with deep MSAs [104] |
| MSA Transformer | Protein language model for coevolution detection | Uses attention mechanisms; often better than GREMLIN for single-fold proteins [104] |
| HMMER3.3.2 | Generates deep multiple sequence alignments | jackhmmer implementation with tuned E-values for optimal coverage [106] |
| HHSUITE | Filters MSAs by sequence identity | Creates nested subfamily MSAs to enhance alternative fold signals [104] |
| AlphaFold2 | Reference structure prediction | Benchmark against ACE predictions; identifies single-fold bias [104] [40] |
Integrated Benchmarking Framework: EA vs. ML
Comparative Strengths and Applications
The integrated benchmarking of EA versus ML approaches reveals complementary strengths:
- EA strengths: Detection of alternative conformations, identification of coevolutionary signatures for fold switching, evolutionary interpretation of metamorphic capability [104].
- ML strengths: High-accuracy single-fold prediction, rapid proteome-scale application, excellent performance on proteins with deep MSAs [40].
- Integrated approach: Combining EA's sensitivity to alternative folds with ML's atomic-level accuracy provides the most comprehensive structural understanding [104] [40].
The experimental confirmation of dual-fold coevolution through the ACE methodology represents a significant advance in protein structure prediction, addressing a fundamental limitation of current ML approaches. The validation across 56 diverse fold-switching proteins demonstrates that metamorphic capability is an evolutionarily selected trait with widespread biological implications.
For researchers and drug development professionals, these findings highlight the importance of:
- Considering multiple conformations in functional characterization of proteins, particularly for therapeutic targets.
- Applying specialized EA methods like ACE when investigating proteins with suspected conformational switching.
- Integrating EA and ML approaches for comprehensive structural understanding rather than relying exclusively on single-method predictions.
As AI-driven protein design expands to explore novel regions of the protein functional universe [16], accounting for potential dual-fold capability will be crucial for designing proteins with controlled conformational dynamics. The benchmarking framework presented here provides a pathway for developing next-generation prediction tools that capture the full structural complexity of metamorphic proteins.
The field of protein science stands at a transformative crossroads, shaped by two powerful paradigms: evolutionary analysis (EA) and machine learning (ML). For decades, evolutionary principles, particularly the analysis of co-evolving residues across multiple sequence alignments (MSAs), provided the foundational framework for computational structure prediction [40]. The recent emergence of deep learning systems like AlphaFold2 (AF2) represents a revolutionary shift, achieving unprecedented accuracy by integrating these evolutionary principles with sophisticated neural network architectures [40]. This whitepaper examines the critical intersection of these approaches within protein structure prediction, analyzing the points of their convergence and, more critically, the points of their divergence. Framed within the context of benchmarking for protein folding overview research, we provide a systematic analysis of performance metrics, delineate the limitations of both methodologies, and offer standardized protocols for their evaluation, serving as a guide for researchers and drug development professionals navigating this complex landscape.
Convergent Foundations: How EA Principles Power ML Success
The remarkable success of modern ML protein structure prediction tools is not a wholesale departure from traditional EA but rather a deep integration and enhancement of its core principles.
The Central Role of Evolutionary Coupling
The cornerstone of both EA and ML approaches is the concept of evolutionary coupling [40]. The fundamental insight is that the mutual compatibility of interactions within a protein structure imposes selection pressure on its amino acid sequence. Consequently, mutations at one position often necessitate compensatory mutations at a spatially proximal, interacting position to maintain structural integrity and function. EA methodologies traditionally extracted these residue-residue contact probabilities from MSAs using statistical methods like direct coupling analysis (DCA) to distinguish direct from indirect couplings [40].
ML as an Evolutionary Information Integrator
Deep learning architectures, particularly AlphaFold2's Evoformer, subsume and extend this evolutionary analysis. They do not merely calculate pairwise couplings; they process the entire MSA through a specialized transformer to model complex, higher-order dependencies and patterns that are difficult to capture with simpler statistical models [40]. This allows the network to build a potent, co-evolution-informed potential of what the native structure should be. In this sense, ML serves as a powerful engine for EA, leveraging the same fundamental biological signal but with enhanced pattern recognition capabilities. Protein language models like ESMFold represent a further evolution, implicitly learning these evolutionary patterns from single sequences by being pre-trained on massive sequence databases, effectively internalizing the rules of evolution [40].
Divergent Realities: The Benchmarking Gap Between Prediction and Experiment
Despite their convergence on a theoretical foundation, a significant gap emerges when the predictions of EA/ML systems are benchmarked against experimental reality. This divergence highlights the limitations of both current ML models and the evolutionary data that inform them.
Quantitative Performance Gaps
Systematic benchmarking against experimental structures reveals specific, quantifiable shortcomings in ML predictions. The following table summarizes key performance gaps identified in a comprehensive analysis of nuclear receptor structures.
Table 1: Quantitative Performance Gaps between AlphaFold2 and Experimental Structures
| Metric | Performance Gap | Biological Implication |
|---|---|---|
| Ligand-Binding Pocket Volume | Systematic underestimation by 8.4% on average [107] | Impacts accuracy for structure-based drug design and ligand docking studies. |
| Domain Variability | LBDs show higher variability (CV=29.3%) vs. DBDs (CV=17.7%) [107] | Highlights challenge in predicting conformational flexibility in functional domains. |
| Functional Asymmetry | Fails to capture conformational diversity in homodimers [107] | Misses biologically relevant states where experimental structures show asymmetry. |
| Severe Deviation Cases | Positional divergence >30 Å and RMSD of 7.7 Å in a two-domain protein [108] | Demonstrates potential for catastrophic failure, often linked to unusual conformations or limited data. |
Limitations of the Evolutionary and ML Paradigms
The gaps illustrated in Table 1 stem from several fundamental limitations:
- Training Data Bias: ML models are trained on experimental structures from the PDB, which predominantly represent stable, low-energy conformations. This leads to a bias toward predicting single, ground-state structures and an inability to capture the full spectrum of functionally important conformational states, including folding intermediates and metamorphic states [109] [107].
- The Static Structure Problem: AF2 and similar tools predict a single, static structure. However, proteins are dynamic entities. They cannot natively represent the folding pathway, functional dynamics, or the phenomenon of fold-switching seen in metamorphic proteins, which share highly similar sequences but adopt distinct topologies [108] [109].
- Extrapolation Failure: As demonstrated by the case of the two-domain protein with a 7.7 Å RMSD error, ML models struggle with proteins that have unusual conformations, insufficient homologous sequences for a deep MSA, or high topological complexity [108]. This indicates a limitation in generalizing beyond the training distribution.
Benchmarking Frameworks: From Static Accuracy to Dynamic Fidelity
The established practice of benchmarking using static energy and force errors (e.g., MAE, RMSE) is insufficient for evaluating models for practical simulation tasks [110]. A more holistic framework is required.
The MLIPAudit Benchmarking Suite
To address this, initiatives like MLIPAudit provide a standardized framework for evaluating Machine Learned Interatomic Potentials (MLIPs). It shifts the focus from static errors to performance on downstream application tasks [110]. The suite includes benchmarks for:
- Small Organic Compounds: Testing dihedral scans, conformer selection, and interaction energies.
- Biomolecules: Evaluating backbone sampling, folding dynamics, and stability of proteins and flexible peptides.
- Molecular Liquids and Solvated Systems: Assessing the prediction of bulk properties and solvation effects.
This approach recognizes that models with similar static force errors can perform vastly differently in dynamic simulations, and it provides a community resource for transparent model comparison [110].
Workflow for Integrated EA-ML-Experimental Benchmarking
The following diagram visualizes a robust benchmarking workflow that integrates EA, ML, and experimental validation to fully assess a predictive model.
Diagram 1: Integrated EA-ML-Experimental Benchmarking Workflow. This workflow evaluates models on static metrics, dynamic simulation stability, and ultimately, experimental agreement.
Experimental Protocols for Validation
To concretely validate and probe the limitations of EA/ML predictions, specific experimental protocols are essential.
Protocol 1: Stopped-Flow Kinetics for Folding Pathway Analysis
This protocol is used to characterize the folding mechanism and identify intermediates, as employed in the study of metamorphic proteins B4 and Sb3 [109].
- Objective: To determine whether a protein follows a two-state or three-state folding mechanism and to characterize the stability of any intermediate states.
- Procedure:
- Rapid Dilution: The purified protein is rapidly diluted from a high concentration of denaturant (e.g., 6 M Guanidine Hydrochloride, GdnHCl) into a series of solutions with varying lower concentrations of GdnHCl.
- Fluorescence Monitoring: The time-dependent change in intrinsic tryptophan fluorescence, which reports on the burial of aromatic residues during folding, is monitored.
- Chevron Plot Analysis: The logarithm of the observed folding/unfolding rate constants ((k_{obs})) is plotted against the denaturant concentration. A linear "V-shaped" chevron plot indicates a two-state mechanism. A non-linear "roll-over" in the refolding arm indicates the accumulation of a folding intermediate (three-state mechanism) [109].
- Perturbation Studies: The experiment is repeated under different conditions (e.g., pH, stabilizing salts) to probe the stability and characteristics of the transition state and any intermediates.
Protocol 2: Deep Mutational Scanning (DMS) for Epistasis and Stability
This high-throughput method, as applied to the GnRHR membrane protein, assesses how mutations impact function and expression on a large scale [111].
- Objective: To quantify the functional effects of hundreds to thousands of single-point mutations in a protein, revealing constraints on stability and folding.
- Procedure:
- Library Construction: A comprehensive library of mutant genes is created via site-directed mutagenesis.
- Functional Selection: The mutant library is expressed in a cellular system (e.g., mammalian cells). For membrane proteins like GPCRs, plasma membrane expression (PME) is often used as a proxy for proper folding [111].
- Sorting and Sequencing: Cells are sorted based on a functional readout (e.g., surface expression via fluorescence). High-throughput DNA sequencing of sorted populations quantifies the enrichment or depletion of each mutation.
- Epistasis Analysis: The effect of each mutation is measured in different genetic backgrounds (e.g., wild-type vs. a destabilizing mutant). Non-additive effects (epistasis) reveal how mutations interact, providing insights into folding pathways and energetic constraints [111].
Table 2: Key Research Reagents and Resources for Protein Folding Research
| Item | Function and Application |
|---|---|
| Stopped-Flow Spectrofluorometer | Apparatus for rapid mixing (<1 ms) of reagents to monitor fast kinetic events like protein folding. |
| Guanidine Hydrochloride (GdnHCl) | Chemical denaturant used to unfold proteins and create energy landscapes for folding studies via chevron plots [109]. |
| Fluorescent Tags (e.g., SNAP-tag) | Used for site-specific labeling of proteins for detection and quantification, crucial for assays like Plasma Membrane Expression (PME) in DMS [111]. |
| Fluorescence-Activated Cell Sorter (FACS) | Instrument for high-throughput sorting and analysis of cells based on fluorescent labels, enabling Deep Mutational Scanning [111]. |
| AlphaFold Database (AFDB) | Repository of over 214 million pre-computed AF2 protein structure models for initial hypothesis generation [40]. |
| ColabFold | Accessible protein structure prediction platform combining MMseq2 for fast MSA generation and AlphaFold2, often via Google Colab [40]. |
| Protein Data Bank (PDB) | International repository for experimentally determined 3D structures of proteins, serving as the ground truth for benchmarking predictions [40]. |
| Foldseck | Tool for rapid comparison of protein structures and large-scale searches of structural databases [40]. |
The relationship between Evolutionary Analysis and Machine Learning in protein folding is one of deep synergy tempered by critical divergence. While ML has masterfully leveraged the principles of EA to achieve stunning predictive accuracy, benchmarking against experimental reality consistently reveals gaps in dynamic representation, conformational diversity, and the handling of atypical folds. For researchers in academia and drug discovery, the path forward requires a disciplined, integrated approach. This involves leveraging standardized benchmarking suites like MLIPAudit, employing rigorous experimental protocols to probe folding dynamics and mutational effects, and maintaining a clear understanding that even the most advanced ML predictions are computational hypotheses. They are powerful starting points that must be validated and refined through empirical evidence, ensuring that the convergence of EA and ML truly illuminates, rather than obscures, the complex reality of protein folding.
Conclusion
The benchmarking of Evolutionary Analysis and Machine Learning reveals that these are not mutually exclusive but powerfully complementary paradigms for protein folding. While ML models like AlphaFold have achieved unprecedented accuracy for single-fold prediction, EA methods remain crucial for identifying evolutionarily selected phenomena like fold-switching, which current ML models systematically miss. The future of the field lies in the integration of these approaches—using generative AI to explore the vast, uncharted protein universe and leveraging evolutionary insights to constrain and validate these designs. For biomedical research, this synergy promises to accelerate the discovery of novel therapeutic targets, enable the de novo design of high-affinity drugs and biologics, and ultimately usher in a new era of personalized medicine. Overcoming current limitations, such as accurately modeling protein-ligand interactions and dynamic conformational changes, will be the next frontier.
References
- 1. and de novo Protein design for biotechnological... folding protein [https://pubmed.ncbi.nlm.nih.gov/24268901/]
- 2. A master of protein origami | Hub [https://hub.jhu.edu/magazine/2025/summer/proteins-folding-unfolding-aging/]
- 3. DIProT: A deep learning based interactive toolkit for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10876589/]
- 4. AI in Biology: What Difference Did the Rise of the Machines ... [https://mindmatters.ai/2025/03/ai-in-biology-what-difference-did-the-rise-of-the-machines-make/]
- 5. Protein folding: bringing theory and experiment closer ... [https://www.sciencedirect.com/science/article/abs/pii/S0959440X03000071]
- 6. Practical Approaches to Protein Folding and Assembly [https://pmc.ncbi.nlm.nih.gov/articles/PMC2778058/]
- 7. Evaluation of machine learning-assisted directed evolution ... [https://www.sciencedirect.com/science/article/abs/pii/S2405471225002200]
- 8. Benchmarking Machine Learning Methods for Protein ... [https://310.ai/blog/benchmarking-machine-learning-methods-for-protein-folding-a-comparative-study-of-esmfold-omegafold-and-alphafold]
- 9. Highly accurate protein structure prediction with AlphaFold [https://www.nature.com/articles/s41586-021-03819-2]
- 10. AlphaFold2: A high-level overview | AlphaFold [https://www.ebi.ac.uk/training/online/courses/alphafold/inputs-and-outputs/a-high-level-overview/]
- 11. Coevolutionary signals in multiple sequence alignments ... [https://www.nature.com/articles/s41598-025-24298-9]
- 12. Comparing co-evolution methods and their application to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5860252/]
- 13. The functional importance of co-evolving residues in proteins [https://pmc.ncbi.nlm.nih.gov/articles/PMC11113390/]
- 14. A method for multiple-sequence-alignment-free protein ... [https://www.nature.com/articles/s42256-023-00721-6]
- 15. Novel machine learning approaches revolutionize protein ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10570143/]
- 16. The Role of AI-Driven De Novo Protein Design in the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12467925/]
- 17. Deep learning techniques have significantly impacted ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8222070/]
- 18. Artificial intelligence methods for protein folding and design [https://www.sciencedirect.com/science/article/abs/pii/S0959440X25000843]
- 19. A comprehensive application of FiveFold for conformation ... [https://www.nature.com/articles/s41598-025-17022-0]
- 20. SimpleFold: Folding Proteins is Simpler than You Think [https://machinelearning.apple.com/research/simplefold]
- 21. The Protein Folding Problem: The day AI unlocked a secret of ... [https://medicine.iu.edu/blogs/research-updates/the-protein-folding-problem-the-day-ai-unlocked-a-secret-of-life]
- 22. An iterative deep learning-guided algorithm for directed ... [https://www.sciencedirect.com/science/article/pii/S2589004225015858]
- 23. Protein structure prediction using the evolutionary ... [https://pubmed.ncbi.nlm.nih.gov/36780132/]
- 24. Evolutionary selection of proteins with two folds [https://www.nature.com/articles/s41467-023-41237-2]
- 25. Assessing scoring metrics for AlphaFold2 and AlphaFold3 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12516916/]
- 26. Beyond static structures: protein dynamic conformations ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12262120/]
- 27. State of the Art of Protein Structure Prediction as of 2025 ... [https://medium.com/advances-in-biological-science/state-of-the-art-of-protein-structure-prediction-as-of-2025-as-evaluated-by-casp16-0c423636bc97]
- 28. The genetic architecture of protein stability [https://www.nature.com/articles/s41586-024-07966-0]
- 29. Protein folding: the endgame [https://pubmed.ncbi.nlm.nih.gov/9242917/]
- 30. The Thermodynamics of Intelligence: Coherence Geometry ... [https://zenodo.org/records/15368797]
- 31. New class of protein misfolding simulated in high definition [https://science.psu.edu/news/Obrien8-2025]
- 32. Signatures of adaptation in myopia-related genes on the ... [https://jphysiolanthropol.biomedcentral.com/articles/10.1186/s40101-023-00341-4]
- 33. Integrative Transcriptome and Proteome Analyses ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10584019/]
- 34. Proteomic insights into myopia development: Differential ... [https://www.sciencedirect.com/science/article/pii/S2949723X24000126]
- 35. A large-scale benchmark study of tools for the classification ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9757047/]
- 36. AlphaFold is five years old — these charts show how it ... [https://www.nature.com/articles/d41586-025-03886-9]
- 37. AlphaFold: Five Years of Impact [https://deepmind.google/blog/alphafold-five-years-of-impact/]
- 38. Distinguishing features of fold‐switching proteins - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9951197/]
- 39. Evolutionary selection of proteins with two folds - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10482954/]
- 40. Emerging frontiers in protein structure prediction following ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11999738/]
- 41. Critical assessment of AI-based protein structure prediction [https://www.sciencedirect.com/science/article/pii/S2950363925000353]
- 42. 2025 Guide to AI-Driven Protein Design & Machine Learning ... [https://nanohelix.ai/blog/ai-protein-design-guide-2025]
- 43. The Illustrated AlphaFold | Elana Simon [https://elanapearl.github.io/blog/2024/the-illustrated-alphafold/]
- 44. AlphaFold [https://en.wikipedia.org/wiki/AlphaFold]
- 45. Meta's ESMfold: the rival of AlpahFold2 | by Salvatore Raieli [https://salvatore-raieli.medium.com/metas-esmfold-the-rival-of-alpahfold2-2223b67f6021]
- 46. OmegaFold: High-resolution de novo Structure Prediction ... [https://hpc.nih.gov/apps/OmegaFold.html]
- 47. OmegaFold: Deep Learning Paradigm for Universal Protein ... [https://sciety-labs.elifesciences.org/articles/by?article_doi=10.21203/rs.3.rs-7514283/v1]
- 48. OmegaFold [https://cosmic-cryoem.org/tools/omegafold/]
- 49. PSBench: a large-scale benchmark for estimating the ... [https://arxiv.org/html/2505.22674v1]
- 50. Generative artificial intelligence for de novo protein design [https://www.sciencedirect.com/science/article/pii/S0959440X24000216]
- 51. De novo design of protein structure and function with ... [https://www.nature.com/articles/s41586-023-06415-8]
- 52. An iterative deep learning-guided algorithm for directed ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12415074/]
- 53. A Benchmark for De novo Protein Design from Function [https://arxiv.org/html/2505.20346v1]
- 54. PDFBench: A Benchmark for De novo Protein Design from ... [https://openreview.net/forum?id=4blGqgStVr]
- 55. Severe deviation in protein fold prediction by advanced AI [https://www.nature.com/articles/s41598-025-89516-w]
- 56. Decoding the limits of deep learning in molecular docking for ... [https://pubs.rsc.org/en/content/articlehtml/2025/sc/d5sc05395a]
- 57. Kipu Quantum at AI in Drug Discovery 2025 [https://kipu-quantum.com/knowledge-hub/blog/kipu-quantum-at-ai-in-drug-discovery-2025/]
- 58. Navigating the landscape of protein folding and proteostasis [https://www.nature.com/articles/s41392-025-02439-w]
- 59. Top 5 Drug Discovery Trends 2025 Driving Breakthroughs [https://www.pelagobio.com/cetsa-drug-discovery-resources/blog/drug-discovery-trends-2025/]
- 60. Folding the Future: How AI is Reshaping Protein Engineering [https://www.synbiobeta.com/read/folding-the-future-how-ai-is-reshaping-protein-engineering]
- 61. Current status and emerging frontiers in enzyme engineering [https://www.sciencedirect.com/science/article/pii/S2405844024087048]
- 62. Benchmarking inverse folding models for antibody CDR ... [https://pubmed.ncbi.nlm.nih.gov/40465588/]
- 63. Benchmarking inverse folding models for antibody CDR ... [https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0324566]
- 64. Accelerated enzyme engineering by machine-learning guided ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11747319/]
- 65. Protein engineering approaches for antibody fragments [https://pmc.ncbi.nlm.nih.gov/articles/PMC6426644/]
- 66. Synthetic Biology Technologies for Antibody Discovery [https://isogenica.com/synthetic-biology-technologies-for-antibody-discovery/]
- 67. AlphaFold2 fails to predict protein fold switching - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9134877/]
- 68. The EVcouplings Python framework for coevolutionary ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6499242/]
- 69. Large-scale discovery of protein interactions at residue ... [https://www.nature.com/articles/s41467-021-21636-z]
- 70. AlphaFold2 fails to predict protein fold switching - PubMed - NIH [https://pubmed.ncbi.nlm.nih.gov/35634782/]
- 71. How accurate are AlphaFold 2 structure predictions? [https://www.ebi.ac.uk/training/online/courses/alphafold/validation-and-impact/how-accurate-are-alphafold-structure-predictions/]
- 72. Evolutionary selection of proteins with two folds - PubMed - NIH [https://pubmed.ncbi.nlm.nih.gov/37673981/]
- 73. Protein–Ligand Docking in the Machine-Learning Era - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9323102/]
- 74. Benchmarking AlphaFold-enabled molecular docking ... [https://pubmed.ncbi.nlm.nih.gov/36065847/]
- 75. Artificial intelligencemethods for protein folding and design [https://pubmed.ncbi.nlm.nih.gov/40505455/]
- 76. Benchmarking different docking protocols for predicting the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10945300/]
- 77. Benchmarking of different molecular docking methods for ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-018-2449-y]
- 78. Efficient Inference, Training, and Fine-tuning of Protein ... [https://www.sciencedirect.com/science/article/pii/S2589004225017560]
- 79. High-accuracy protein complex structure modeling based ... [https://www.nature.com/articles/s41467-025-65090-7]
- 80. SimpleFold: Folding Proteins is Simpler than You Think [https://arxiv.org/abs/2509.18480]
- 81. Evolutionary algorithms simulating molecular evolution [https://pmc.ncbi.nlm.nih.gov/articles/PMC11317538/]
- 82. What Have We Learned from Design of Function in Large ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10521758/]
- 83. Accurate structure prediction of biomolecular interactions ... [https://www.nature.com/articles/s41586-024-07487-w]
- 84. Current successes and remaining challenges in protein ... [https://www.frontiersin.org/journals/bioinformatics/articles/10.3389/fbinf.2023.1222182/full]
- 85. Recent advances and challenges in protein complex ... [https://www.sciencedirect.com/science/article/pii/S2001037024001363]
- 86. Advances of computational protein design: Principles ... [https://www.sciencedirect.com/science/article/abs/pii/S0734975025001429]
- 87. pLDDT: Understanding local confidence | AlphaFold [https://www.ebi.ac.uk/training/online/courses/alphafold/inputs-and-outputs/evaluating-alphafolds-predicted-structures-using-confidence-scores/plddt-understanding-local-confidence/]
- 88. Flexibility or uncertainty? A critical assessment of ... [https://www.sciencedirect.com/science/article/abs/pii/S0969212625003442]
- 89. Optimizing and Benchmarking Parallelism and Memory ... [https://medium.com/@co232/optimizing-and-benchmarking-parallelism-and-memory-consumption-for-alphafold-and-fastfold-a1acbc3d92fe]
- 90. New benchmark metrics for protein-protein docking methods [https://pmc.ncbi.nlm.nih.gov/articles/PMC3076516/]
- 91. Reliable protein–protein docking with AlphaFold, Rosetta, ... [https://elifesciences.org/articles/94029]
- 92. Unlocking the power of AI models: exploring protein folding ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11377126/]
- 93. An end-to-end framework for the prediction of protein ... [https://www.nature.com/articles/s41467-024-51776-x]
- 94. Probing Functional Allosteric States and Conformational Ensembles of... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12103074/]
- 95. Balancing Speed and Precision in Protein Folding: A Comparison of... [https://openreview.net/forum?id=eRE68eRo7t]
- 96. A Comparison of AlphaFold2, ESMFold, and OmegaFold [https://icml.cc/virtual/2025/51245]
- 97. Investigating whether deep learning models for co-folding ... [https://www.nature.com/articles/s41467-025-63947-5]
- 98. Structural Biology in the AlphaFold Era: How Far Is Artificial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12109453/]
- 99. Recent Progress of Protein Tertiary Structure Prediction - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10893003/]
- 100. Decoding Protein Structures: From AlphaFold to Beyond [https://www.recursion.com/news/demystifying-protein-structure-prediction-models-alphafold-rosettafold-esmfold-and-beyond]
- 101. Are protein language models the new universal key? [https://www.sciencedirect.com/science/article/pii/S0959440X25000156]
- 102. Use OmegaFold Online [https://neurosnap.ai/service/OmegaFold]
- 103. Improving deep learning protein monomer and complex ... [https://www.nature.com/articles/s41592-023-02130-4]
- 104. Evolutionary selection of proteins with two folds [https://www.nature.com/articles/s41467-023-41237-2?error=cookies_not_supported&code=565309ac-cbae-4647-8bcc-663306334efd]
- 105. Benchmarking AlphaFold2 on Peptide Structure Prediction [https://pmc.ncbi.nlm.nih.gov/articles/PMC9883802/]
- 106. GitHub - ncbi/ dual _ fold _ coevolution [https://github.com/ncbi/dual_fold_coevolution]
- 107. Bridging prediction and reality: Comprehensive analysis of ... [https://www.sciencedirect.com/science/article/pii/S2001037025001734]
- 108. Severe deviation in protein prediction by advanced AI: a case study fold [https://pubmed.ncbi.nlm.nih.gov/39922965/]
- 109. Topological determinants in protein dynamics: a comparative... folding [https://biologydirect.biomedcentral.com/articles/10.1186/s13062-025-00642-x]
- 110. MLIPAudit: A benchmarking tool for Machine Learned ... [https://arxiv.org/html/2511.20487v1]
- 111. -Mediated Epistasis Among Unstable Membrane... Divergent Folding [https://elifesciences.org/reviewed-preprints/92406]