Molecular Dynamics Simulation Box Size Optimization: Balancing Accuracy and Computational Efficiency
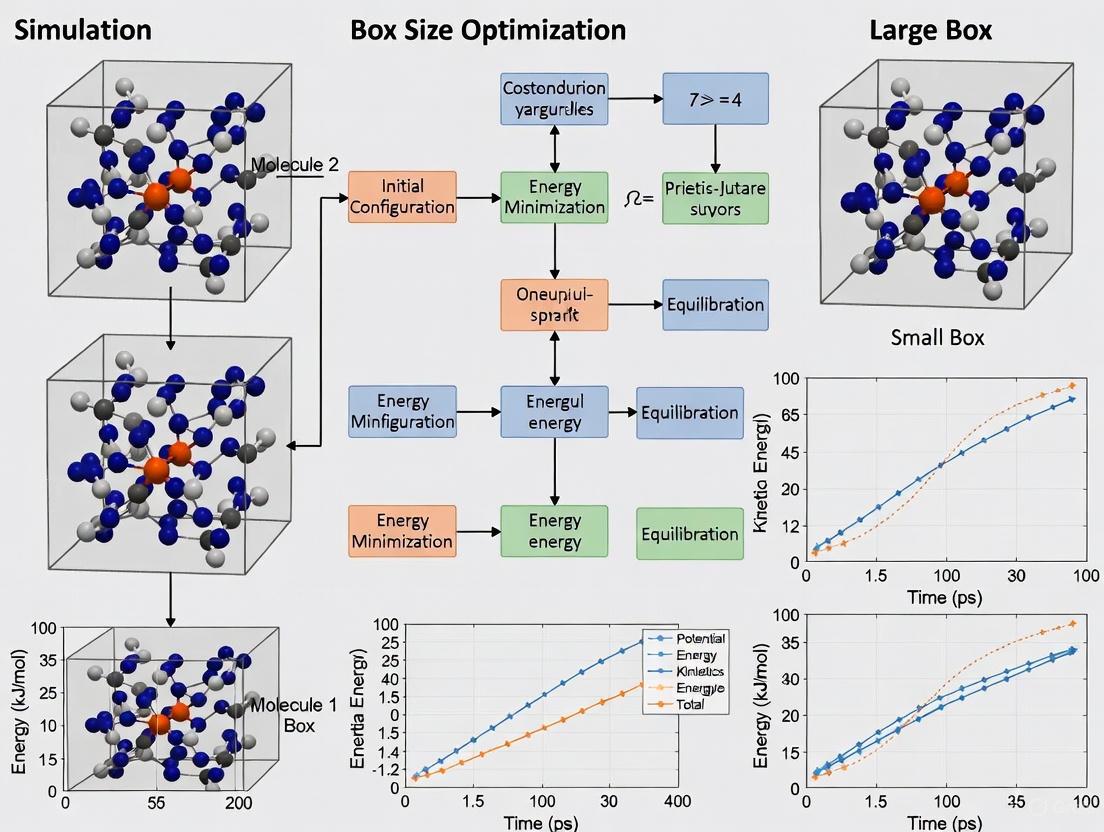
This comprehensive review examines molecular dynamics simulation box size optimization strategies for biomedical and drug development applications.
Molecular Dynamics Simulation Box Size Optimization: Balancing Accuracy and Computational Efficiency
Abstract
This comprehensive review examines molecular dynamics simulation box size optimization strategies for biomedical and drug development applications. Covering foundational principles, methodological approaches, troubleshooting techniques, and validation protocols, we provide researchers with evidence-based guidance for selecting appropriate simulation volumes. The article addresses critical considerations including periodic boundary condition artifacts, computational efficiency trade-offs, and statistical validation methods, with specific applications to protein folding, peptide dynamics, and biomolecular interactions. By synthesizing current research findings and best practices, this work serves as a practical resource for optimizing simulation box parameters to ensure physically meaningful results while maximizing computational resources in pharmaceutical research.
Fundamental Principles of Simulation Box Design in Molecular Dynamics
Frequently Asked Questions (FAQs)
1. What are Periodic Boundary Conditions (PBC) and why are they used in Molecular Dynamics (MD) simulations? Periodic Boundary Conditions (PBC) are a computational method used to model a large, bulk system by simulating a small, representative part of it. In this setup, the central simulation box is surrounded by identical copies of itself in all directions. This creates an infinite, periodic lattice that eliminates the artificial surfaces and edge effects that would otherwise exist in a finite box, allowing researchers to more accurately model the behavior of a system within a macroscopic environment [1] [2]. This approach is efficient because it enables the study of bulk properties without the computational expense of simulating a truly immense number of particles [1].
2. What are the common artifacts or errors associated with an incorrectly chosen simulation box size? An improperly small box size can introduce several artifacts due to the unnatural proximity of periodic images:
- Artificial Correlations: The simulated molecules may interact with their own periodic images, leading to unrealistic forces and corrupting the simulation results [2]. One study notes that artifacts are possible when periodicity is combined with Ewald summation methods for long-range electrostatics [2].
- Loss of Accuracy: For proteins, a box that is too small can constrain large-scale dynamics, such as domain motions, and prevent the correct reproduction of experimental data, like neutron scattering results [2].
- Altered Thermodynamics and Kinetics: While some reported box-size effects on thermodynamics and kinetics may vanish with sufficient sampling [3], using an extremely small box (e.g., with only ~0.5 nm between the solute and box edge) definitively introduces artifacts. In such cases, the solvation shells of periodic images interact, and water screening of electrostatic interactions becomes insufficient [3].
3. What is the minimum recommended distance between a solute (like a protein) and the edge of the simulation box? Research on protein crystallization solutions has established a practical minimum distance. A study on lysozyme oligomers found that a minimum distance of 1 nm between the protein atoms and the box face was sufficient to correctly reproduce the relative stability of different oligomers (e.g., stable dimers vs. unstable hexamers) without introducing significant artifacts [4]. Reducing the box size further, such that this offset is less than 1 nm, can lead to serious artifacts because protein copies in neighboring virtual boxes would exert an unrealistically large influence on each other [4].
4. How does the choice of boundary conditions impact the calculation of electrostatic interactions? The treatment of electrostatics is tightly linked to the use of PBC. The standard method is the Particle Mesh Ewald (PME) algorithm, which accurately accounts for long-range electrostatic interactions within a periodic system [3]. However, caution is required:
- In Vacuum: Using PBC/PME to simulate a single, charged molecule in a vacuum is problematic. It creates a periodic system with infinite charge and can lead to severe simulation artifacts, as the lack of a screening solvent means periodic images interact strongly and unrealistically [3].
- In Solution: Water molecules effectively screen electrostatic interactions. With a sufficiently large box (maintaining the recommended 1 nm solute-box distance), PME provides accurate results, and thermodynamic properties remain independent of box size [3].
5. Are there alternatives to standard cubic PBC for simulating elongated systems? Yes, Asymmetric Periodic Boundary Conditions (APBC) can be used for systems like infinitely long polymers, including DNA. In APBC, the simulation box is a cuboid where the periodicity along one axis (e.g., the z-axis, along the polymer chain) is dictated by the intrinsic periodicity of the molecule itself. The box dimensions in the perpendicular directions (x and y) are chosen to provide adequate solvation, similar to a standard simulation. This approach allows for the efficient study of sequence-dependent properties and the ion atmosphere around elongated molecules without the computational cost of simulating a very long chain in a large cubic box [5].
Troubleshooting Guides
Issue 1: Unrealistic System Behavior Due to Small Box Size
Problem: Your simulation shows unstable protein oligomers, unphysical forces, or constrained domain motions that are not observed experimentally. Solution: Verify and adjust your simulation box size to ensure sufficient spacing between periodic images. Methodology:
- Measure the Offset: Calculate the minimum distance between any atom of your solute (e.g., a protein) and the face of the simulation box.
- Apply the 1 nm Rule: Ensure this minimum distance is at least 1 nm [4]. The following table summarizes findings from a systematic investigation on lysozyme oligomers:
Table 1: Stability of Lysozyme Oligomers in Different Box Sizes [4]
| Minimum Solute-Box Face Distance (Offset) | Dimer Stability | Hexamer Stability | Conclusion |
|---|---|---|---|
| 1.0 nm | Stable | Unstable | Results are correct |
| 1.5 nm | Stable | Unstable | Results are correct |
| 2.0 nm | Stable | Unstable | Results are correct |
| 2.5 nm | Stable | Unstable | Results are correct |
| 3.0 nm | Stable | Unstable | Results are correct |
- Re-equilibrate: After increasing the box size, repeat the energy minimization and system equilibration steps (NVT and NPT ensembles) before proceeding with production simulation.
Issue 2: Poor Convergence of Thermodynamic Properties
Problem: Calculated properties, such as solvation free energy, show a strong dependence on the simulation box size and do not converge. Solution: Distinguish between genuine box-size artifacts and a simple lack of statistical sampling. Methodology:
- Ensure a Minimally Sufficient Box: First, confirm your box meets the 1 nm minimum distance rule to eliminate obvious artifacts [4] [3].
- Perform Multiple Repeats: If the box size is sufficient, the apparent dependence is likely a sampling issue. Conduct multiple independent simulation runs (repeats) for the property of interest at a single box size.
- Calculate Statistical Uncertainty: Compute the average and standard deviation (or confidence intervals) across the repeats. A study on hydration free energy showed that apparent trends from single simulations were statistically insignificant; the true value was box-size independent once a large number of repeats (N=20) were analyzed [3].
- Use Enhanced Sampling: For challenging calculations like protein solvation free energy, employ advanced sampling techniques such as Hamiltonian Replica Exchange to improve phase space exploration and ensure convergence [3].
Issue 3: Simulating a Solvated Polymer or Filament
Problem: You need to simulate an effectively infinite polymer (like DNA) and want to avoid the cost of a very large solvated sphere or an artificially short chain. Solution: Implement Asymmetric Periodic Boundary Conditions (APBC). Methodology:
- System Setup: Position the polymer (e.g., a 10-base-pair DNA segment) so its long axis is parallel to the z-direction of the box [5].
- Define Box Dimensions: Set the z-dimension (Lz) to match the intrinsic periodicity of the molecule (e.g., the length of one or more helical pitches). The x and y dimensions (Lx, Ly) can be chosen independently to provide a sufficient solvation shell, typically using the 1 nm rule [5].
- Apply an Asymmetric Barostat: In the NpT ensemble, use a barostat that allows the x and y dimensions to fluctuate independently of the fixed z-dimension to maintain the correct molecular periodicity [5].
- Analysis Consideration: Be aware that global properties like the end-to-end distance of the polymer will be constrained by the periodicity. Instead, focus on local properties (e.g., local bending angles) to extract parameters such as the persistence length [5].
The following workflow diagram summarizes the decision process for setting up a simulation with PBC:
Table 2: Key Software and Force Fields for MD Simulations with PBC
| Item Name | Function / Application | Relevant Context from Search Results |
|---|---|---|
| GROMACS | A high-performance MD simulation package. | Used in recent studies to investigate box size effects and protein oligomer stability [4] [3]. |
| AMBER ff99SB-ILDN | A force field for simulating proteins and nucleic acids. | Employed in the study determining the minimum acceptable box size for lysozyme simulations [4]. |
| Particle Mesh Ewald (PME) | An algorithm for handling long-range electrostatic interactions under PBC. | Described as the standard method for electrostatics in solvated, periodic systems [3]. |
| TIP4P-Ew | A four-site water model for explicit solvation. | Used to fill the simulation box in the lysozyme oligomer stability study [4]. |
| LINCS | An algorithm to constrain bond lengths, allowing for longer simulation time steps. | Used to keep the bond lengths of lysozyme oligomers constant during dynamics [4]. |
Box Size Effects on Thermodynamic Properties and System Behavior
Frequently Asked Questions (FAQs)
FAQ 1: Does simulation box size significantly affect thermodynamic properties like solvation free energy? For sufficiently large boxes with adequate sampling, the simulation box size has only a minimal or non-existent effect on thermodynamic properties such as solvation free energy. Apparent effects often disappear when a sufficient number of simulation repeats are performed, indicating that observed trends from single or few realizations are likely statistical artifacts rather than a true physical phenomenon [3]. For example, calculations of hydration free energy for a small molecule (anthracene) and a protein (GB) showed no statistically significant trend across boxes ranging from 473 to 5334 water molecules when based on 20 repeats per box size [3].
FAQ 2: What is the minimum acceptable distance between the solute and the box edge? A minimum distance (offset) of 1 nanometer between the solute atoms and the box face is generally acceptable for protein systems to avoid major artifacts [3] [4]. One study on lysozyme oligomers found that a 1 nm offset was the smallest permissible size that still yielded correct stability trends between dimers and hexamers [4]. However, reducing this distance further (e.g., to ~0.5 nm) introduces artifacts because water screening becomes insufficient and solvation shells of periodic images can interact [3].
FAQ 3: Can a smaller box size affect the kinetics of a biomolecular process? Similar to thermodynamics, the kinetics of biomolecular processes (e.g., protein conformational transition rates) are not significantly affected by the simulation box size, provided that the box is sufficiently large and adequate sampling is performed. The primary concern is ensuring that the box size does not lead to spurious interactions between periodic images, which is mitigated by maintaining the recommended minimum offset [3].
FAQ 4: Why do some studies report a box size effect while others do not? Discrepancies often arise from differences in sampling and statistics. Studies reporting box size effects may be based on insufficient sampling, where conclusions are drawn from a small number of trajectory replicates that exhibit random statistical fluctuations [3]. Other factors include the use of different force fields, treatment of long-range electrostatics, solute restraints, and equilibration protocols [3]. When extensive sampling and multiple replicates are used, the box size effect typically vanishes [3].
FAQ 5: How does the treatment of electrostatics interact with box size? The proper treatment of long-range electrostatics is crucial. The Particle-Mesh Ewald (PME) method is standard. In vacuum simulations, a small box size can strongly alter the protein's environment as distances between periodic images decrease, creating a significant box size dependence. In explicit solvent, water molecules screen these electrostatic interactions, making properties independent of box size in sufficiently large boxes [3]. If solvent screening is reduced, a box size dependence can re-emerge [3].
Troubleshooting Guides
Problem 1: Observed thermodynamic property drifts with changing box size.
- Potential Cause: Inadequate sampling and lack of uncertainty quantification.
- Solution:
- Increase the number of independent simulation repeats (e.g., N=20) for each box size [3].
- Use enhanced sampling techniques (e.g., Hamiltonian replica exchange) to improve phase space exploration [3].
- Always calculate and report confidence intervals (e.g., standard error) for your estimates. An observable with a large confidence interval indicates the result is not statistically significant, and any perceived trend is likely anecdotal [3].
Problem 2: Simulation results in vacuum are highly dependent on box size.
- Potential Cause: In non-solvated (vacuum) systems with periodic boundaries, using PME for electrostatics leads to artificial interactions between periodic images as the box shrinks.
- Solution:
- For vacuum calculations, use an infinitely large non-periodic box as a reference, which can be approximated by a very large box or by applying the appropriate analytical corrections [3].
- Alternatively, perform the relevant thermodynamic cycle in explicit solvent, where screening is present, and subtract the vacuum component calculated in a very large box [3].
Problem 3: Unstable protein oligomers or unexpected unfolding in crystallization solutions.
- Potential Cause: The simulation box may be too small, leading to artificial interactions that destabilize the native state.
- Solution:
- Ensure the minimum distance between the protein surface and the box edge is at least 1 nm [4].
- Validate your box size by testing known system properties. For example, in a lysozyme crystallization solution, a correct simulation should show stable dimers but unstable hexamers. If a hexamer remains stable in a small box, the box size is likely causing artifacts [4].
Experimental Protocols & Data
Protocol 1: Assessing Box Size Dependence for Solvation Free Energy
This protocol is adapted from studies on small molecules and the GB protein [3].
1. System Setup:
- Solute Preparation: Obtain the initial coordinates of your solute (e.g., a small molecule or protein).
- Box Size Selection: Create multiple simulation boxes where the solute is solvated. Systematically vary the box size. A recommended range is from the minimally acceptable size (e.g., >2x the non-bonded cut-off) to a very large size (e.g., >5000 water molecules) [3].
- Solvation: Solvate the solute with water models (e.g., TIP4P-Ew) and add ions to neutralize the system and match experimental conditions [4].
2. Simulation Parameters:
- Force Field: Choose an appropriate force field (e.g., Amber ff99SB-ILDN for proteins) [4].
- Electrostatics: Use the Particle-Mesh Ewald (PME) method for long-range electrostatics [3] [4].
- Ensembles: Use the NPT ensemble for equilibration and production runs to maintain constant pressure and temperature [6].
- Thermostat/Barostat: Use a thermostat (e.g., V-rescale) and a barostat (e.g., Parrinello-Rahman) [4].
3. Enhanced Sampling & Sampling Amount:
- For challenging calculations like protein solvation, use Hamiltonian replica exchange to ensure proper equilibration and phase space overlap between alchemical windows [3].
- Invest significant sampling time (e.g., >1 μs aggregate time per box size) and perform multiple independent repeats (e.g., N=20) to gather statistics [3].
4. Analysis:
- Calculate the free energy for each box size and repeat using a method like the alchemical pathway [3].
- Plot the mean estimated free energy for each box size with error bars (e.g., standard deviation or 95% confidence interval) across the repeats.
- A statistically significant box size effect is absent if the confidence intervals for different box sizes overlap substantially and no trend is visible.
Protocol 2: Determining Minimum Box Size for Protein Oligomers
This protocol is adapted from a study on lysozyme oligomers [4].
1. System Preparation:
- Oligomer Construction: Build models of the protein oligomers (e.g., dimer and hexamer) from a known crystal structure.
- Protonation: Set the protonation states of amino acid residues to match the experimental pH (e.g., pH 4.5) [4].
- Box Creation: Place the oligomer in the center of a cubic simulation box. Create boxes with varying offsets (e.g., 1.0, 1.5, 2.0, 2.5, 3.0 nm) [4].
2. Simulation Run:
- Energy Minimization: Use the steepest descent algorithm until the maximum force is below a threshold (e.g., 1000 kJ/(mol·nm)) [4].
- Equilibration:
- Equilibrate in the NVT ensemble for 100 ps.
- Equilibrate in the NPT ensemble for 100 ps.
- Maintain the target temperature (e.g., 10°C) using a thermostat [4].
- Production MD: Run long-scale production simulations (e.g., 700 ns to 1 μs) for each box size [4].
3. Analysis and Validation:
- Calculate stability metrics for each oligomer in each box size:
- Root-mean-square fluctuation (RMSF)
- Root-mean-square deviation (RMSD)
- Radius of gyration (Rg)
- The minimum acceptable box size is the smallest one where the known stability trends from experiment are reproduced. For example, in lysozyme solutions, the dimer should be more stable than the hexamer [4].
Quantitative Data on Box Size and Protein Stability
The table below summarizes data from a molecular dynamics study of lysozyme dimers and hexamers in boxes of different sizes. The stability was judged by the ability of the simulation to reproduce the experimental observation that dimers are stable while hexamers are not [4].
Table 1: Stability of Lysozyme Oligomers in Different Simulation Box Sizes
| Minimum Protein-Box Edge Offset (nm) | Box Edge Length for Dimer (nm) | Box Edge Length for Hexamer (nm) | Dimer Stability (over 1 μs) | Hexamer Stability (over 700 ns) | Conclusion |
|---|---|---|---|---|---|
| 1.0 | 8.6 | 10.9 | Stable | Unstable | Acceptable [4] |
| 1.5 | 9.7 | 11.9 | Stable | Unstable | Acceptable [4] |
| 2.0 | 10.6 | 12.9 | Stable | Unstable | Acceptable [4] |
| 2.5 | 11.6 | 13.9 | Stable | Unstable | Acceptable [4] |
| 3.0 | 12.6 | 14.9 | Stable | Unstable | Acceptable [4] |
The Scientist's Toolkit
Table 2: Essential Research Reagents and Software for Box Size Studies
| Item Name | Function / Relevance in Box Size Studies |
|---|---|
| GROMACS | A molecular dynamics simulation package used for setting up, running, and analyzing simulations. It was used in key studies to assess box size effects [3] [4]. |
| AMBER (ff99SB-ILDN force field) | A force field and software suite. The ff99SB-ILDN force field is commonly used for simulating proteins and was applied in the lysozyme oligomer stability study [4]. |
| Particle-Mesh Ewald (PME) | An algorithm for handling long-range electrostatic interactions in simulations with periodic boundary conditions. Its proper use is critical for avoiding box-size artifacts [3] [4]. |
| TIP4P-Ew Water Model | A four-site water model frequently used in MD simulations. It was used to solvate the lysozyme oligomers in the box size study [4]. |
| Hamiltonian Replica Exchange | An enhanced sampling method that improves conformational sampling by exchanging states between replicas with different Hamiltonians. It is crucial for achieving converged solvation free energies of proteins [3]. |
| Analysis Tools | |
gmx rmsf, gmx rms, gmx gyrate |
GROMACS tools for calculating Root-Mean-Square Fluctuation (RMSF), Root-Mean-Square Deviation (RMSD), and Radius of Gyration (Rg), respectively. These are standard metrics for assessing protein stability and convergence [4]. |
Workflow Diagram
Minimum Image Convention and Cutoff Radius Relationships
Frequently Asked Questions (FAQs)
1. What is the fundamental relationship between the cutoff radius and the simulation box size under the Minimum Image Convention (MIC)?
The fundamental rule is that the cutoff radius ((R_c)) must be less than half the length of the shortest box vector [7] [8]. Formally, this is expressed as:
(R_c < \frac{1}{2} \min(\|\mathbf{a}\|, \|\mathbf{b}\|, \|\mathbf{c}\|))
This ensures that a particle interacts with, at most, the closest periodic image of any other particle in the system, preventing unphysical interactions where a single particle would interact with both another particle and its own periodic image simultaneously [7] [9].
2. What happens if I violate this relationship, for example, by setting a cutoff radius that is too large?
Violating this rule introduces significant artifacts, including [10] [7] [8]:
- Unphysical Forces and Energies: A single particle may interact with the same neighbor more than once (e.g., with the "real" particle and its image), leading to grossly overestimated interaction energies and incorrect forces.
- Energy Non-Conservation: As particles move, their interactions can discontinuously "switch" from one image to another, causing a failure in energy conservation that is critical for molecular dynamics simulations [11].
- Structural Distortions: The resulting incorrect forces can distort the simulated liquid or molecular structure, leading to unreliable results.
3. My simulation involves a macromolecule, like a protein. Is the half-box-length rule sufficient?
For simulations of macromolecules in solvent, a more stringent rule applies. The box must be large enough so that the cutoff radius does not exceed half the shortest box vector, and the solvent layer around the macromolecule must be thick enough to prevent a single solvent molecule from "seeing" both sides of the macromolecule [7]. A common recommendation is to have at least 1 nm of solvent between the surface of the solute and the edge of the box in all directions [12]. Compromising on this to save computational cost is common but introduces approximation errors [7].
4. Are the requirements different for non-cubic simulation boxes, like a rhombic dodecahedron?
The core principle remains the same: (R_c) must be smaller than half the shortest distance between any two faces of the periodic box [7]. The strength of using non-cubic boxes like the rhombic dodecahedron or truncated octahedron is that they are more spherical than a cube. This allows for a smaller box volume for the same cutoff radius, saving significant computational time by reducing the number of solvent molecules required [7] [8].
5. How does the choice of algorithm for searching particle neighbors affect the cutoff restriction?
The required relationship can become more restrictive depending on the search algorithm. While the fundamental MIC rule is (R_c < \frac{1}{2} \min(\|\mathbf{a}\|, \|\mathbf{b}\|, \|\mathbf{c}\|)), some algorithms impose tighter bounds for efficiency [7]:
- For grid search: (Rc < \min(ax, by, cz))
- For simple search: (Rc < \frac{1}{2} \min(ax, by, cz))
Always consult your specific molecular dynamics software documentation for its exact requirements.
Troubleshooting Guides
Problem 1: Energy Drift or "Blowing Up" of the Simulation
Symptoms: The total energy of the system is not conserved but steadily increases (or decreases) over time. The simulation may eventually crash as atoms acquire unrealistically high velocities.
Potential Causes and Solutions:
| Cause | Diagnostic Check | Solution |
|---|---|---|
| Cutoff radius is too large, violating the MIC. | Verify that your set (R_c) is less than half the shortest box side length. Check your simulation log files for warnings. | Reduce (R_c) to comply with the rule, or increase the box size. |
| Forces are discontinuous when particles move across the cutoff boundary. | Check if your potential function is smoothly truncated to zero at (R_c). A sharp cutoff causes force discontinuities. | Apply a switching function or potential shift to ensure the potential and force smoothly go to zero at the cutoff [11]. |
| Incorrect treatment of long-range interactions like electrostatics. | This problem persists even with a correct short-range cutoff. | For electrostatic interactions, use a dedicated long-range method like Particle Mesh Ewald (PME) instead of a simple cutoff [7] [12]. |
Problem 2: Unphysical Structural Artifacts
Symptoms: The radial distribution function (g(r)) shows strange peaks or dips, or the structure of a liquid or solvent shell around a protein looks unnatural.
Potential Causes and Solutions:
| Cause | Diagnostic Check | Solution |
|---|---|---|
| The simulation box is too small, causing the macromolecule to interact with its own periodic image. | Measure the distance between the solute and its closest periodic image. If it is less than (2 \times R_c), the box is too small. | Increase the box size to ensure a sufficient solvent buffer. A minimum of 1 nm is a good starting point [12]. |
| The cutoff radius is too small to capture essential interactions. | Check if your (R_c) is large enough for the potential to have decayed sufficiently. For Lennard-Jones, common cutoffs are 2.5σ/σ. | Increase (R_c) while ensuring the MIC is still respected. This will require a larger simulation box. |
Experimental Protocols for Box Size and Cutoff Optimization
Protocol 1: Validating MIC Compliance for a Given System
This protocol outlines the steps to ensure a new simulation setup adheres to the Minimum Image Convention.
1. Determine Shortest Box Vector: * For a cubic box of side length (L), the shortest vector is (L). * For a rectangular box, it is the smallest of the three side lengths. * For a triclinic box, use the norm (length) of each box vector ((\mathbf{a}, \mathbf{b}, \mathbf{c})). The shortest norm is the limiting factor [7].
2. Calculate Maximum Allowable Cutoff: * Apply the rule: (R_{c}^{max} = \frac{1}{2} \times \text{shortest box vector}).
3. Set Operational Cutoff: * Choose a cutoff radius (Rc) that is less than (R{c}^{max}). A safety margin of 0.1 nm is often advisable to account for numerical rounding and finite integration timesteps.
4. Verify Solvent Buffer (for solvated systems):
* Measure the maximum diameter of your solute (e.g., protein) in each dimension.
* Calculate the available solvent buffer: (box_length - solute_diameter) / 2.
* Ensure the solvent buffer is greater than your chosen (R_c). If not, you must increase the box size.
Protocol 2: Systematic Investigation of Cutoff Artifacts
This experiment quantifies the impact of the cutoff radius on simulation stability and physical properties.
Objective: To determine a cutoff radius that provides a physically accurate representation of the bulk system while being computationally efficient.
Methodology:
- System Preparation: Create a well-equilibrated system (e.g., a box of water or a solvated protein) with a fixed, sufficiently large box size.
- Parameter Variation: Run a series of short simulations using identical starting coordinates and velocities but varying the cutoff radius (Rc). Ensure all values both obey and deliberately violate the MIC (e.g., from 0.4 × (R{c}^{max}) to 0.6 × (R_{c}^{max})).
- Data Collection: For each simulation, track:
- Total energy conservation (drift over time).
- Potential energy.
- Radial distribution function, (g(r)).
- Density of the system.
Expected Outcome: Simulations with (Rc > 0.5 \times \text{box length}) will show clear energy drift and unphysical structures. The goal is to identify a value of (Rc) below which these properties converge to a stable, artifact-free result. The following table summarizes a hypothetical investigation for a Lennard-Jones fluid in a 4.0 nm cubic box:
| Cutoff Radius (nm) | Ratio (R_c / L) | MIC Compliant? | Energy Drift (kJ/mol/ps) | g(r) First Peak Height | Conclusion |
|---|---|---|---|---|---|
| 1.8 | 0.45 | Yes | Low (~0.01) | 3.05 | Accurate and stable |
| 1.95 | 0.487 | Yes | Low (~0.01) | 3.04 | Accurate and stable |
| 2.1 | 0.525 | No | High (>1.0) | 2.81 | Unphysical, unstable |
Data Presentation
Table 1: Common Simulation Box Types and Their Properties for MIC
| Box Type | Image Distance | Box Volume (relative to cube) | Typical Use Case | MIC Restriction |
|---|---|---|---|---|
| Cubic | (L) | (L^3) (1.000) | General use, crystals | (R_c < \frac{1}{2}L) |
| Rhombic Dodecahedron (xy-square) | (L) | (0.707~L^3) | Spherical solutes (proteins) | (R_c < \frac{1}{2} \times \text{shortest vector}) |
| Truncated Octahedron | (L) | (0.770~L^3) | Spherical solutes (proteins) | (R_c < \frac{1}{2} \times \text{shortest vector}) |
Table comparing common periodic box shapes. The rhombic dodecahedron and truncated octahedron are more efficient for simulating spherical molecules as they require ~23-30% fewer solvent molecules for the same image distance [7].
Mandatory Visualization
The Scientist's Toolkit
Research Reagent Solutions
| Item | Function in MIC/Cutoff Context |
|---|---|
| Periodic Simulation Box | The fundamental "container" that defines the unit cell of the periodic system. Its dimensions directly dictate the maximum allowable cutoff radius [7] [8]. |
| Cutoff Radius ((R_c)) | The distance beyond which non-bonded interactions are truncated. It is the key variable that must be optimized for physical accuracy and MIC compliance [7] [13]. |
| Short-Ranged Potential | A potential, like the Lennard-Jones, whose value decays to zero within a finite distance, making it compatible with truncation at (R_c) [13]. |
| Long-Range Electrostatic Solver (e.g., PME) | A method required for handling electrostatic interactions, which decay slowly and cannot be accurately treated with a simple cutoff without introducing severe artifacts [7] [12]. |
| Neighbor List | A computational list of particles within an extended radius ((R{list} > Rc)) that is updated periodically. This drastically reduces the number of pairwise distance calculations needed each step while maintaining accuracy [11] [9]. |
Frequently Asked Questions (FAQs)
1. Does simulation box size meaningfully affect the thermodynamics and kinetics of my biomolecular system? For a wide range of biomolecular systems, current evidence suggests that with sufficient statistical sampling, the simulation box size has a minimal effect on both thermodynamics and kinetics, provided a minimum distance is maintained between the solute and the box edge. Early studies that reported strong box-size dependencies have, upon re-examination with more replicates, largely failed to demonstrate statistically significant effects. The apparent effects often vanish when adequate sampling is performed and uncertainties are properly accounted for [3] [14].
2. What is the minimum acceptable distance to maintain between my solute and the box edge? A minimum offset of 1.0 nm is generally acceptable for many protein systems, such as lysozyme oligomers in a crystallization solution. This distance ensures that the protein copies in periodic images do not exert an unrealistic influence on one another. While larger boxes (e.g., with a 1.5-2.0 nm offset) may be used, they do not necessarily provide more accurate results for stability assessments but come with a significant computational cost increase [4].
3. Can a larger box size change the hydrophobic effect or water properties in my simulation? Analyses suggest that reported changes in water properties (e.g., radial distribution functions, diffusion constants) with box size are often explainable by a dilution effect. In a larger box, there is a higher proportion of bulk-like water to protein-bound water, which changes the system-wide average. When analyzed properly (e.g., by keeping the analysis volume constant), no significant box-size effect on the inherent hydrophobic effect or local water structure is observed [14].
4. I see conflicting results in the literature about box size effects. Why is that? Many apparent conflicts arise from insufficient statistics. Single or a handful of simulation replicates are prone to substantial statistical fluctuations, which can create illusory trends. Studies that perform many replicates (e.g., 20 per condition) typically find that these box-size dependencies are not statistically significant [3] [14]. Differences in simulation setup details (e.g., protonation states, treatment of electrostatic interactions, force fields) can also contribute to disparate findings [14].
Troubleshooting Guides
Issue: Unexpected Instability or Conformational Changes
Problem: Your protein becomes unstable or undergoes a conformational change in a small simulation box, but you suspect it might be an artifact.
Diagnosis Steps:
- Check Box Size: Verify that the minimum distance between any protein atom and the box edge is at least 1.0 nm [4].
- Run Multiple Replicas: A single simulation can be misleading. The initial observation might be a statistical outlier. Run at least 3-5 replicas for each condition to assess variability [14].
- Compare to Larger Box: Run a single, shorter simulation in a significantly larger box (e.g., with a 2.0 nm offset). If the same instability is observed, it is less likely to be a box-size artifact [3].
Solution: If instability disappears in a larger box and with multiple replicas, increase your box size to maintain at least a 1.0 nm offset and always plan for multiple simulation replicates.
Issue: Artifacts from Overly Small Box Sizes
Problem: The simulation box is so small that the solute's periodic images are interacting directly.
Symptoms:
- Abnormal structural fluctuations (very high RMSD/RMSF).
- disruption of expected thermodynamic stability (e.g., a protein hexamer disassembling when it should be stable, or a dimer destabilizing) [4].
- Corrupted solvation shell and unrealistic electrostatic interactions due to insufficient screening [3].
Solution: Always ensure the box size is larger than twice the non-bonded cut-off radius in all dimensions. The recommended minimum offset of 1.0 nm already satisfies this condition for common cut-off settings [3] [4].
Issue: High Computational Cost with Large Boxes
Problem: Your simulation is running very slowly because the box size leads to a very large number of particles.
Diagnosis: The computational cost of molecular dynamics scales with the number of atoms. A larger box with more solvent atoms increases the cost of force calculations and communication in parallel runs [15].
Optimization Strategies:
- Justify Box Size: Use the minimum acceptable offset of 1.0 nm unless your specific system requires a larger one [4].
- Optimize Parallelization: Use efficient parallelization schemes in GROMACS, such as domain decomposition, and consider offloading the non-bonded calculations to a GPU [15].
- Check Performance Settings: Ensure you are using an optimized version of your MD engine compiled with the correct SIMD instructions (e.g., AVX2) for your CPU architecture [15].
Experimental Data and Protocols
Table 1: Summary of findings from key molecular dynamics studies on box size effects.
| System Studied | Box Sizes (Edge Length or Offset) | Key Finding | Reference |
|---|---|---|---|
| Lysozyme Dimer/Hexamer | 1.0, 1.5, 2.0, 2.5, 3.0 nm offset | A 1.0 nm offset is the minimum permissible for correct stability ranking of oligomers. | [4] |
| Hemoglobin T-to-R Transition | 9, 12, 15 nm cubes | No significant box size effect on kinetics or thermodynamics was found with sufficient (10-20) replicates. | [14] |
| Hydration Free Energy (Anthracene) | 473 to 5334 water molecules | No trend with box size was observed when using 20 repeats per size. | [3] |
| DNA Dodecamer | 5, 10, 15 Å water layers | The smallest box (5 Å layer) showed no disadvantage compared to larger boxes. | [16] |
| Solvation Free Energy (GB Protein) | Varied, >1.0 nm offset | No box size dependence in solvated systems, but a dependence emerges in vacuum simulations. | [3] |
Quantitative Impact of Box Size on Computational Cost
Table 2: Illustrative computational cost as a function of simulation box size. Cost is influenced by the number of atoms and simulation parameters.
| System Description | Approx. Number of Atoms | Relative Computational Cost per ns | Key Considerations |
|---|---|---|---|
| Small Protein, 1.0 nm offset | ~50,000 | 1.0x (Baseline) | Optimal balance of cost and accuracy for many systems. |
| Same Protein, 1.5 nm offset | ~100,000 | ~2.5x | Significant cost increase for potentially minimal accuracy gain. |
| Same Protein, 2.0 nm offset | ~175,000 | ~5x | Often unnecessary; reserve for testing specific artifacts. |
Detailed Protocol: Establishing Minimum Box Size for a Protein System
This protocol, based on the methodology from [4], provides a systematic way to determine a sufficient box size for your protein.
Title: Workflow for Determining Minimum Simulation Box Size
Procedure:
System Preparation:
- Select your protein of interest and prepare its structure (e.g., assign protonation states appropriate for the pH).
- Generate models for different oligomeric states known to be relevant. For instance, if studying crystallization, use oligomers known to be stable (dimers) and unstable (hexamers) in solution as a reference [4].
Box Creation and Solvation:
- For each oligomer, create a series of cubic simulation boxes where the minimum distance between the protein atoms and the box face (the "offset") is systematically varied. A recommended range is 1.0, 1.5, and 2.0 nm [4].
- Solvate the system with a water model (e.g., TIP4P-Ew) and add ions to neutralize the system and match the experimental precipitant or salt concentration.
Simulation and Equilibration:
- Use an MD package like GROMACS [4] and a modern force field (e.g., Amber ff99SB-ILDN).
- Perform standard energy minimization and equilibration in NVT and NPT ensembles.
- Run production simulations. For stability assessments, extended times are needed; a duration of 700 ns to 1 μs is often necessary to observe stability differences [4].
Stability Analysis:
- Calculate standard stability metrics from the trajectories:
- Root-mean-square deviation (RMSD) of the oligomer backbone relative to the starting structure.
- Root-mean-square fluctuation (RMSF) of individual alpha-carbon atoms.
- Radius of gyration (Rg) of the oligomer.
- The key is to compare the relative stability between different oligomeric states (e.g., dimer vs. hexamer). The correct, physically meaningful stability ranking (e.g., dimer more stable than hexamer) should be reproduced [4].
- Calculate standard stability metrics from the trajectories:
Interpretation:
- The smallest box size that correctly reproduces the expected stability ranking of the oligomers, and for which the calculated stability metrics do not show signs of major artifactual instability, can be considered the minimum acceptable box size for future studies of that system.
The Scientist's Toolkit
Table 3: Essential software and analysis tools for molecular dynamics simulations.
| Tool Name | Type | Primary Function | Application Note |
|---|---|---|---|
| GROMACS [15] [4] | MD Software Suite | High-performance MD simulation engine. | Highly optimized for CPUs and GPUs. Uses domain decomposition and particle-mesh Ewald (PME) for efficient parallelization [15]. |
| Particle-Mesh Ewald (PME) [3] [4] | Algorithm | Accurate calculation of long-range electrostatic interactions in periodic systems. | Essential for realism. Requires a sufficiently large box so that water can properly screen interactions between periodic images [3]. |
| geomeTRIC [17] | Optimization Library | Molecular geometry optimization using internal coordinates. | Can be used for optimizing structures with neural network potentials; performance depends on the potential used. |
| Sella [17] | Optimization Library | Transition-state and minimum optimization. | Another optimizer for NNPs; using internal coordinates can significantly improve performance and success rate [17]. |
#1 Core Concepts and Selection Guide
What are the common simulation box geometries in molecular dynamics, and how do I choose?
In molecular dynamics (MD), a simulation box with Periodic Boundary Conditions (PBCs) is used to model a bulk system by effectively creating an infinite lattice of identical images of the primary box. The most common geometries are cubic, rectangular (orthorhombic), and triclinic.
The following table compares the key characteristics of the primary box shapes:
| Box Geometry | Typical Use Case | Key Characteristics | Shape in Real Space | Common Force Field Cut-off Requirement |
|---|---|---|---|---|
| Cubic | Isotropic systems (e.g., simple liquids, spherical proteins in solution) | All box vectors are equal in length and mutually perpendicular. Simplest to implement. | Cube | Minimum box length ≥ 2 × R~c~ |
| Rectangular (Orthorhombic) | Solids or biomolecules with different dimensions | Box vectors are perpendicular but of different lengths. | Rectangular Prism | Minimum box dimension ≥ 2 × R~c~ |
| Triclinic (General) | Complex systems, efficient packing for spherical molecules | Three box vectors of different lengths and non-orthogonal angles. Most general form. | Parallelepiped | Minimum image distance ≥ 2 × R~c~ |
| Rhombic Dodecahedron | Efficient simulation of spherical molecules (e.g., globular proteins) | A specific type of triclinic box that is nearly spherical, minimizing the number of solvent atoms needed. | Rhombic Dodecahedron | Minimum image distance ≥ 2 × R~c~ |
The choice of box geometry significantly impacts computational efficiency and the physical accuracy of your simulation. A box that is too small can lead to artifacts where a particle interacts with its own periodic image, which is unphysical [8] [4]. The minimum image criterion must be satisfied, which generally requires that the box dimensions are larger than twice the non-bonded interaction cut-off distance (R~c~) [8]. For a cubic box, this means all sides must be longer than 2R~c~.
For simulating a single globular protein in solution, a rhombic dodecahedron (a specific triclinic box) is often the most efficient choice because its near-spherical shape minimizes the number of required water molecules, reducing computational cost while maintaining a sufficient distance between the protein and its images [8] [18].
#2 Troubleshooting Common Box Geometry Problems
Why does LAMMPS warn that my "Triclinic box skew is large," and how can I fix it?
This warning indicates that the angles of your triclinic box deviate significantly from 90 degrees, which can cause simulations to run inefficiently [19]. While the simulation may still be valid, performance can suffer.
Solutions:
- Remap to an Orthogonal Box: If your system allows, the most straightforward solution is to convert the skewed box into an orthogonal (rectangular) one. This can often be achieved by replicating the unit cell to create a supercell and then remapping the atomic coordinates [19]. This process also helps avoid finite size effects by creating a larger simulation box.
- Increase System Size: The warning is often more critical for very small systems. For "typical" systems with thousands of atoms, the performance impact might be negligible. However, if your box is small, increasing its size can mitigate both the skew warning and potential finite-size artifacts [19].
- Check and Remap Coordinates: As a standard solution, you can remap the coordinates by cyclically swapping the axes (e.g., z → x, x → y, y → z). This can be done by modifying the input file and the "Atoms" coordinate section, a task easily accomplished with command-line tools like
awk[19].
My protein in a triclinic box is rotating and getting too close to its periodic image. Is this a problem?
Yes, this is a significant problem. If your protein interacts with its periodic image, it will introduce severe artifacts and invalidate your simulation results [18].
Solutions:
- Check the Minimum Distance: Use analysis tools specific to your MD software to monitor the distance between periodic images of your protein. In GROMACS, the command
gmx mindist -piis designed for this purpose [18]. - Increase the Box Size: The solution is to increase the distance between the protein and the box boundary. When setting up the simulation, ensure the minimum distance from any protein atom to the box face is at least 1.2 nm [18]. This provides a buffer larger than the typical van der Waals cut-off (e.g., 1.0 nm), preventing direct interactions between the protein and its image.
- Use a Rhombic Dodecahedron: For simulating globular proteins in solution, using a
-bt dodecahedronbox in GROMACS (which creates a rhombic dodecahedron) is highly recommended. This shape is more spherical and is the most efficient for enclosing a globular object, minimizing solvent count while maintaining a safe distance to the boundary [18].
My simulation box keeps shrinking during an NPT simulation. What is wrong?
A steadily shrinking box in an isothermal-isobaric (NPT) ensemble simulation typically indicates that the system is collapsing inward because the internal pressure is lower than the target pressure of the barostat.
Potential Causes and Solutions:
- Incorrect Initial Density: The initial configuration may have the molecules placed too far apart, in a low-density state (like a gas phase). The barostat then correctly compresses the system to reach the target density of a liquid or solid phase [20].
- Inaccurate Force Field Topology: This is a common issue. If the interatomic attractions are too strong, or if the intramolecular or intermolecular repulsions are too weak, the molecules will collapse into an artificially dense state. You must refine your topology [20].
- For large molecules: Do not parametrize the entire large molecule as a single unit. Instead, break it down into smaller, representative model compounds, parametrize these pieces, and then link them together. This avoids inconsistent charge assignments and other parametrization errors [20].
- Check existing literature: Often, topologies for common molecules (like Tween 80) have already been published. Using these validated parameters can save time and ensure accuracy [20].
- Missing Solvent: If you are trying to simulate a liquid without explicit solvent molecules (e.g., in vacuum), the molecules will coalesce into a droplet, and the box will shrink accordingly. For condensed phase simulations, you must include the appropriate solvent [20].
#3 Experimental Protocols and Best Practices
Protocol: Determining the Minimum Acceptable Simulation Box Size
This protocol is based on a study that established a minimum box size by comparing the stability of lysozyme oligomers, a method that can be adapted for validating box sizes in other protein systems [4].
1. System Preparation:
- Oligomer Models: Obtain or create structural models of the protein oligomers you wish to study (e.g., a stable dimer and an unstable hexamer). These can be isolated from a known crystal structure using visualization software like PyMOL [4].
- Solution Conditions: Set up the protonation states of amino acid residues to match the experimental pH of the solution you are modeling. Use a tool like the PROPKA server [4].
- Simulation Boxes: Place the oligomers at the center of cubic simulation boxes. Create a series of boxes where the minimum distance between the protein atoms and the box face (the "offset") varies systematically (e.g., 1.0 nm, 1.5 nm, 2.0 nm, etc.) [4].
2. Simulation Parameters (Based on GROMACS):
- Force Field: Use an appropriate protein force field (e.g., Amber ff99SB-ILDN) [4].
- Water and Ions: Solvate the system with a water model (e.g., TIP4P-Ew) and add ions to match the experimental precipitant or salt concentration [4].
- Ensemble: Use the NPT ensemble to maintain constant temperature (with a thermostat like V-rescale) and pressure (with a barostat like Parrinello-Rahman). A temperature of 10°C was used in the reference study [4].
- Electrostatics: Use the Particle Mesh Ewald (PME) algorithm for long-range electrostatic interactions. Set the non-bonded interaction cut-off to 1.0 nm [4].
- Constraint Algorithm: Use the LINCS algorithm to constrain bond lengths [4].
3. Analysis and Validation:
- Run simulations for a sufficiently long time to observe stability or dissociation (e.g., 700 ns to 1 μs) [4].
- Calculate the following metrics for each box size:
- Root-mean-square fluctuation (RMSF) of Cα atoms.
- Root-mean-square deviation (RMSD) of Cα atoms from the original structure.
- Radius of gyration (Rg).
- Validation Criterion: The simulation box size is considered sufficient if the results correctly reflect the known experimental stability of the oligomers. For example, if the dimer is known to be stable and the hexamer unstable, the correct box size is the smallest one for which the dimer remains stable and the hexamer becomes unstable during the simulation [4]. The study concluded that a 1.0 nm offset was the minimum permissible for studying lysozyme crystallization solutions [4].
The Scientist's Toolkit: Essential Research Reagents and Software
The following table details key materials and software tools used in the setup and analysis of MD simulations, as referenced in the protocols.
| Item Name | Function/Brief Explanation | Example Use Case |
|---|---|---|
| GROMACS | A fast, free, and widely-used software package for performing MD simulations and analysis. | Primary engine for running simulations and calculating properties like RMSD and RMSF [21] [4]. |
| AMBER | A suite of biomolecular simulation programs, including the tleap program for system building. |
Used for generating initial topologies and system parameters for organic molecules [20]. |
| ACPYPE | A tool for converting molecular topology files from AMBER to GROMACS format. | Essential when using ligands or molecules parametrized with AMBER/GAFF in a GROMACS simulation [20]. |
| PyMOL | A powerful molecular visualization system. | Used to isolate oligomers from a larger crystal structure for simulation [4]. |
| PROPKA | A web server for predicting the pK~a~ values of ionizable residues in proteins. | Critical for determining the correct protonation states of amino acids at a specific pH during system setup [4]. |
| Bio3D (R Package) | An R package for the analysis of biomolecular structure, sequence, and simulation data. | Used to analyze MD trajectories and calculate dynamic cross-correlation matrices (DCCM) [21]. |
| GAFF (General Amber Force Field) | A force field designed for organic molecules, often used for drug-like molecules. | Parametrizing small molecule ligands or surfactants (e.g., Tween 80) for simulation with proteins [20]. |
| TIP4P-Ew Water Model | A 4-site rigid water model parameterized for use with Ewald summation techniques. | Used to solvate the simulation box in explicit solvent simulations [4]. |
#4 Reference Data Tables
Quantitative Results from Lysozyme Box Size Study
The following data is adapted from a study that determined the minimum box size by simulating lysozyme dimers and hexamers in boxes of different sizes [4]. The "Offset" is the minimum distance between the protein atoms and the box face.
Table 1: System Dimensions and Stability Metrics for Lysozyme Oligomers
| Oligomer | Box Offset (nm) | Box Edge Length (nm) | Maximum RMSF (nm) | Maximum RMSD (nm) | Rg Variation (nm) |
|---|---|---|---|---|---|
| Dimer | 1.0 | 8.6 | 0.5 | 0.9 | 0.2 |
| Dimer | 1.5 | 9.7 | 0.6 | 1.0 | 0.2 |
| Dimer | 2.0 | 10.6 | 0.5 | 0.8 | 0.2 |
| Dimer | 2.5 | 11.6 | 0.7 | 1.0 | 0.2 |
| Dimer | 3.0 | 12.6 | 0.5 | 0.8 | 0.2 |
| Hexamer | 1.0 | 10.9 | 1.6 | 1.5 | 1.0 |
| Hexamer | 1.5 | 11.9 | 1.8 | 1.5 | 0.9 |
| Hexamer | 2.0 | 12.9 | 1.4 | 1.4 | 0.8 |
| Hexamer | 2.5 | 13.9 | 1.2 | 1.2 | 0.8 |
| Hexamer | 3.0 | 14.9 | 1.7 | 1.6 | 1.0 |
Interpretation: The dimer showed relatively stable RMSF, RMSD, and Rg values across all box sizes, indicating its inherent stability was not an artifact of the box size. In contrast, the hexamer showed significantly larger fluctuations and deformations (high RMSF, RMSD, and Rg variation) in all boxes, correctly reflecting its instability in solution. This confirmed that even the smallest box (1.0 nm offset) was sufficient to yield correct qualitative behavior for this system [4].
Practical Strategies for Box Size Selection and Optimization
Frequently Asked Questions (FAQs)
What are protein-surface offset standards and why are they important in Molecular Dynamics (MD) simulations? Protein-surface offset standards refer to the established minimum distances that must be maintained between a protein and the boundaries of its MD simulation box. Adhering to these standards is critical for obtaining accurate, biologically relevant results. A sufficient offset prevents the protein from artificially interacting with its own periodic image, which can distort protein folding, dynamics, and interaction properties [22].
My simulation results show abnormal protein folding or dynamics. Could a small simulation box size be the cause? Yes, an undersized simulation box is a likely cause. If the distance between the protein surface and the box edge is too small, the protein can artificially interact with its periodic copies. This "periodic boundary artifact" can force the protein into unnatural conformations, stabilize non-native states, and generally corrupt the sampling of its true structural ensemble [22].
How do I calculate the minimum required distance between my protein and the simulation box edge? The minimum distance is not a single universal value but is determined by the force field's non-bonded cut-off distance. A common and safe standard is to set the box size so that the distance from any protein atom to the nearest box edge is at least twice the non-bonded cut-off radius used in your simulation. Furthermore, research suggests that for reliable thermodynamic and mechanical property prediction, the entire simulation system should contain a minimum number of atoms (e.g., ~15,000 atoms for an epoxy system) to ensure proper statistical sampling and avoid size effects [22].
Does the required offset change when simulating a protein in a cellular environment versus a simple solvent? The fundamental principle remains the same, but the complexity of the environment increases. In a crowded cellular environment, you must account for the size and distribution of other macromolecules (e.g., nucleic acids, ribosomes) in addition to the box boundaries. The goal is to prevent the protein from interacting with its periodic image or with periodic images of other crowders, which may require a larger overall system size [23].
Troubleshooting Guides
Issue: Unstable Protein Structure in Simulations
- Problem: The protein unfolds or adopts unrealistic conformations shortly after the simulation begins.
- Potential Cause: Artifactual interactions with periodic images due to an insufficient box size.
- Solution:
- Recalculate Box Dimensions: Ensure the box is large enough so that the protein-surface offset is at least 1.0 to 1.5 nm (10-15 Å) per side, which typically exceeds the recommended twice the cut-off distance.
- Verify with Visualization: Use visualization software like VMD [24] to inspect the initial simulation setup. Check that there is clear, empty space between the protein and all box edges.
- Monitor Distance During Runtime: Implement tools in your MD software (e.g.,
distancetools in LAMMPS [25] or GROMACS) to log the minimum protein-box edge distance throughout the simulation.
Issue: Inaccurate Thermodynamic or Mechanical Properties
- Problem: Predicted properties like density, elastic modulus, or glass transition temperature are imprecise or do not converge.
- Potential Cause: The simulation box is too small to statistically represent the amorphous nature of the material or environment, leading to size effects [22].
- Solution:
- Conduct a Size-Convergence Study: Follow the methodology outlined in research [22]. Build multiple independent system replicates of increasing size (e.g., from 5,000 to 40,000 atoms).
- Plot Property vs. System Size: Calculate your property of interest (e.g., density, modulus) for each system size.
- Identify the Convergence Point: The optimal system size is the smallest size at which the standard deviation of the property predictions becomes acceptably low and stable. Research on an epoxy system found 15,000 atoms to be a good balance between precision and computational cost [22].
Troubleshooting Workflow for Simulation Accuracy
Issue: Poor Performance in Binding Site or Protein-Surface Interaction Analysis
- Problem: Mapping of protein-binding hot spots or analysis of protein-surface interactions is unreliable.
- Potential Cause: Restricted protein dynamics or artifacial surface contacts due to a confined simulation box.
- Solution:
- Ensure Full Solvation Shell: The box must be large enough to accommodate not only the protein but also a full hydration shell around it. This is critical for accurate modeling of protein-water interactions, which directly influence conformational sampling and surface properties [26] [27].
- Use Advanced Mapping Techniques: If studying binding sites, employ computational fragment screening methods like FTMap or mixed-solvent MD (MSMD) [28]. These techniques rely on the protein's ability to sample its natural conformational landscape, which can be hindered by a small box.
- Validate with Multiple Box Sizes: Repeat the analysis in a significantly larger box to confirm the results are not dependent on the initial system size.
Experimental Protocols
Protocol 1: Determining Optimal Simulation System Size
This protocol is adapted from studies on system size optimization for molecular dynamics [22].
Objective: To empirically determine the smallest molecular system size that yields precise and converged thermodynamic properties for your specific protein or material system.
Materials & Software:
- MD Simulation Software (e.g., LAMMPS [25] [22], GROMACS, NAMD [24])
- Molecular Visualization Tool (e.g., VMD [24])
- Force Field parameters (e.g., AMBER [27], CHARMM [26])
Methodology:
- System Replicate Generation: Construct multiple independent simulation systems of your protein in solution with varying total atom counts. A recommended range is from ~5,000 to ~40,000 atoms [22].
- Equilibration: For each system replicate, perform a standard energy minimization and equilibration protocol in the NPT ensemble (constant Number of particles, Pressure, and Temperature) to relax the system to its stable density.
- Production Simulation: Run multiple independent production MD simulations for each system size. Ensure the simulation time is long enough for the property of interest to equilibrate.
- Property Calculation: Calculate the key physical properties (e.g., mass density, radius of gyration, elastic modulus) from each production trajectory.
- Statistical Analysis: For each system size, calculate the average value and standard deviation of each property across the independent replicates.
- Convergence Identification: Plot the average value and standard deviation of the properties against the total number of atoms in the system. The optimal system size is identified as the point where the standard deviation plateaus and further increases in size do not significantly improve precision [22].
Protocol 2: Incorporating Distance Restraints from Experimental Data
This protocol is based on methods using FRET-derived distances to guide MD simulations [23].
Objective: To incorporate experimental distance restraints (e.g., from FRET data) into MD simulations to bias the system toward a target conformational state.
Materials & Software:
- Experimental distance data (e.g., FRET pair separations) [23]
- MD Software with restraint capabilities (e.g., AMBER, GROMACS, LAMMPS)
- Initial and (if available) target protein structures.
Methodology:
- Restraint Selection: Identify the key amino acid pairs for which distance restraints will be applied. Studies show that enforcing only a small fraction of restraints (e.g., Nr/N ≲ 0.08, where N is the number of amino acids) can effectively induce conformational changes [23]. Selection methods can include:
- Force Constant Determination: Define an appropriate force constant for the harmonic restraints based on the uncertainty of the experimental distance measurements.
- Simulation with Restraints: Perform the MD simulation with the distance restraints applied to the selected atom pairs. The potential energy term is typically added to the standard force field.
- Validation: Monitor the Root-Mean-Square Deviation (RMSD) from the target experimental structure to assess the efficacy of the restraints in guiding the conformational change [23].
Data Presentation
This table summarizes findings from a study on an epoxy system, illustrating the convergence of property predictions with increasing system size.
| Number of Atoms | Density (g/cm³) | Std. Dev. (Density) | Elastic Modulus (GPa) | Std. Dev. (Modulus) | Simulation Time (ks) |
|---|---|---|---|---|---|
| 5,265 | ~1.20 | Low | Value | High | ~1,000 |
| 10,530 | ~1.20 | Low | Value | Medium | ~2,500 |
| 14,625 | ~1.20 | Low | Value | Medium | ~5,000 |
| 20,475 | ~1.20 | Low | Value | Low | ~10,000 |
| 31,590 | ~1.20 | Low | Value | Low | ~25,000 |
| 36,855 | ~1.20 | Low | Value | Low | ~40,000 |
This table compares different methods for selecting a minimal set of distance restraints to induce a protein conformational change.
| Restraint Selection Method | Requires Target Structure | Key Principle | Relative Efficacy (Cα RMSD Reduction) |
|---|---|---|---|
| Largest Change in Separation | Yes | Selects residue pairs with the largest distance change between initial and target states. | High |
| Linear Discriminant Analysis | Yes | Uses machine learning to identify discriminants between two conformational states. | High |
| Normal Mode Analysis | No | Selects residues involved in low-frequency collective motions. | Medium |
| Largest Cα Separation | No | Selects the most widely separated residue pairs in the initial structure. | Medium |
| Random Selection | No | Randomly selects a subset of residue pairs. | Low (Baseline) |
The Scientist's Toolkit: Essential Research Reagents & Software
| Item Name | Category | Function / Application |
|---|---|---|
| LAMMPS [25] [22] | MD Software | A highly versatile and open-source molecular dynamics simulator for modeling soft and solid-state materials. |
| AMBER1 [27] | Force Field / Software | A suite of biomolecular simulation programs and force fields. Refined versions (e.g., ff99SBws, ff03w-sc) improve the balance of protein-water interactions [27]. |
| VMD1 [24] | Visualization & Analysis | A molecular visualization program for displaying, animating, and analyzing large biomolecular systems. Essential for system setup and trajectory analysis. |
| CHARMM1 [26] | Force Field | An all-atom empirical force field for a wide range of molecules, widely used for simulating proteins and other biological macromolecules. |
| FTMap1 [28] | Computational Mapping | A computational algorithm for identifying binding hot spots on protein surfaces by exhaustively docking small molecular probes. |
| GROMACS1 [29] | MD Software | A high-performance MD software package for simulations of proteins, lipids, and nucleic acids, known for its extreme speed. |
| OpenMM1 [29] | MD Library | An open-source library for high-performance MD simulation, highly flexible and optimized for GPU acceleration. |
1 These items were listed as relevant examples in the search results [29] or other cited sources.
Step-by-Step Protocol for Initial Box Size Determination
Molecular dynamics (MD) simulations require careful selection of simulation box size to balance computational efficiency with result accuracy. An appropriately sized box minimizes artificial periodicity effects while conserving computational resources. This guide provides researchers and drug development professionals with practical protocols for determining optimal initial box sizes, troubleshooting common errors, and implementing best practices within molecular dynamics workflows.
Frequently Asked Questions (FAQs)
What is the fundamental purpose of a simulation box in molecular dynamics?
The simulation box defines the spatial boundaries for your molecular system and implements periodic boundary conditions (PBCs). PBCs eliminate surface effects by surrounding the primary box with identical image copies in all directions. As particles move out of one side of the box, they re-enter from the opposite side, maintaining a constant number of atoms. This approach models bulk conditions while simulating a finite system [8].
What are the critical factors to consider when determining box size?
- Minimum image criterion: Box dimensions must be at least twice the non-bonded interaction cutoff distance (typically 1.0-1.2 nm) to prevent atoms from interacting with their own images [8]
- System composition: Pure solutions, mixtures, and interfaces have different requirements [30]
- Sampling needs: Larger boxes may provide better statistics for certain properties [30]
- Computational resources: Larger boxes require more atoms and increased computation time [4]
What are the consequences of an improperly sized box?
- Artifactual interactions: Too-small boxes cause unnatural interactions between periodic images [4] [8]
- Domain decomposition errors: Incompatible with parallel processing, causing simulation crashes [30]
- Inaccurate results: Altered structural and dynamic properties, especially for biomolecules [4]
- System instability: Improper energy minimization and equilibration [30]
Quantitative Guidelines for Box Size Determination
Table 1: Recommended Minimum Offsets Between Molecule and Box Edge
| System Type | Minimum Offset (nm) | Reference |
|---|---|---|
| Proteins (general) | 1.0-1.2 | [4] |
| Protein crystallization solutions | 1.0 | [4] |
| Lysozyme dimers/hexamers | 1.0-1.5 | [4] |
| Water-chloroform interface with polymers | 1.0+ (see protocol) | [30] |
Table 2: Common Box Types and Their Applications
| Box Type | Typical Use Cases | Advantages | Disadvantages |
|---|---|---|---|
| Cubic/Cubic (orthorhombic) | General purpose, simple systems | Easy implementation | Less efficient for spherical molecules |
| Rhombic Dodecahedron | Spherical molecules, proteins | Minimizes number of solvent molecules | More complex setup |
| Truncated Octahedron | Proteins in solution | Efficient for spherical systems | Complex implementation |
| Hexagonal Prism | Membrane systems, interfaces | Accommodates elongated structures | Specific orientation requirements |
Experimental Protocols
Basic Protocol 1: Determining Initial Box Size for a Soluble Protein
Materials and Reagents
- Protein structure file (PDB format)
- MD software (GROMACS, NAMD, AMBER)
- Solvent model (TIP3P, TIP4P, SPC/E)
- Force field (AMBER, CHARMM, OPLS-AA)
Procedure
- Prepare molecular structure: Remove crystallographic water and ligands unless critical to study
- Generate topology: Use appropriate tools (e.g.,
pdb2gmxin GROMACS) [31] - Center molecule: Position the protein at the center of the coordinate system
- Calculate initial dimensions: Determine maximum molecular dimensions along x, y, and z axes
- Apply minimum offset: Add 1.0-1.5 nm buffer to each dimension [4]
- Select box type: Choose cubic for simplicity or rhombic dodecahedron for efficiency with spherical proteins
- Solvate system: Add solvent molecules to fill the box using tools like
solvate[31] - Verify box size: Ensure minimum atom-box face distance meets requirements
Basic Protocol 2: Setting Up a Biphasic System (Water-Chloroform Interface)
Materials and Reagents
- Polymer chains structure files
- Solvent models (water and chloroform)
- Force field with appropriate parameters for all components
Procedure
- Establish initial dimensions: Create rectangular box with appropriate x-y dimensions and extended z-dimension (e.g., 4×4×8 nm) [30]
- Separate phases: Place chloroform in lower region, water in upper region
- Equilibrate solvent system: Run NPT equilibration with semi-isotropic pressure coupling
- Add polymer: Insert polymer chains at the interface using tools like
gmx insert-molecules[30] - Stepwise equilibration: Equilibrate in stages, potentially adding components gradually
- Verify separation: Confirm maintenance of phase separation throughout equilibration
Figure 1: Workflow for basic box size determination for soluble proteins.
Troubleshooting Guide
Error: "There is no domain decomposition for X ranks..."
Problem: This GROMACS error occurs when the simulation box is too small to be divided across the requested number of parallel processors [30].
Solutions:
- Reduce MPI ranks: Change from multiple ranks to a single MPI rank
- Adjust thread count: Use
--ntasks=1with increased--cpus-per-task(e.g., 8-16 threads) - Increase box size: Expand box dimensions, particularly the smallest dimension
- Alternative: Use
-rddor-ddsoptions to adjust domain decomposition parameters (not recommended for very small systems) [30]
Error: "Pull group reference atom too far from group center"
Problem: Occurs when pull groups span distances larger than half the box size, particularly in umbrella sampling simulations [32].
Solutions:
- Increase box size: Ensure box dimension in pulling direction is at least twice the pull group size
- Specify reference atom: Designate a centrally located atom as the reference for pull group center of mass calculations [32]
- Check pulling direction: Ensure adequate box size in the specific dimension where pulling occurs
System Instability After Solvation
Problem: Energy minimization fails or system becomes unstable during equilibration.
Solutions:
- Verify box size: Ensure minimum distance from molecule to box face is at least 1.0 nm [4]
- Check solvent placement: Identify and correct overlapping atoms between solute and solvent
- Adjust minimization protocol: Use steepest descent before switching to conjugate gradient methods
- Review force field compatibility: Ensure all components (protein, ligands, solvent) use compatible parameters
The Scientist's Toolkit
Table 3: Essential Software Tools for Box Size Determination
| Tool Name | Function | Application Context |
|---|---|---|
GROMACS (editconf) |
Box generation and manipulation | General MD simulations |
| PACKMOL | Packing molecules in defined regions | Complex multicomponent systems |
| VMD | Visualization and measurement | Verification of box dimensions |
gmx solvate |
Solvation of simulation box | Adding water molecules |
gmx insert-molecules |
Inserting molecules into existing boxes | Adding ligands, ions, or polymers |
Advanced Considerations
Box Size Effects on Biomolecular Stability
Recent research demonstrates that insufficient box sizes can alter protein oligomer stability. In lysozyme solutions, a minimum 1.0 nm offset between protein atoms and box faces was necessary to maintain correct relative stability of dimers and hexamers observed experimentally. Smaller boxes artificially stabilized non-native hexamers, providing incorrect biological conclusions [4].
Efficient Equilibration Strategies
For complex systems like polymer-electrolyte membranes, advanced equilibration protocols can reduce computation time by 200-600% compared to conventional annealing methods. These approaches use targeted NVT and NPT ensembles with specific temperature and pressure coupling to achieve stable densities faster [6].
Figure 2: Relationship between box size and key simulation factors showing the trade-off between computational cost and result accuracy.
Proper simulation box size determination is a critical step in molecular dynamics setup that significantly influences simulation stability, computational efficiency, and result accuracy. By following the protocols outlined in this guide and adhering to the recommended minimum offsets, researchers can avoid common pitfalls and generate reliable simulation data. As systems increase in complexity, careful attention to box dimensions becomes increasingly important for producing biologically relevant results in drug development and biomolecular research.
Adaptive Box Resizing Techniques for Dynamic Systems
Molecular dynamics (MD) simulation is a critical tool for researchers and drug development professionals, enabling the prediction of thermo-mechanical properties of materials and the study of complex biological processes at the atomic level. A fundamental aspect of setting up accurate MD simulations is the selection of an appropriate periodic simulation box size. The simulation box must be large enough to minimize finite-size effects and accurately represent bulk material properties, yet computationally feasible to simulate within practical timeframes. Recent research demonstrates that meaningful simulation results, particularly for systems whose stability relies on long-range effects like the hydrophobic interaction, require surprisingly large solvent boxes. For instance, valid simulations of human hemoglobin require a box containing ten times more water molecules than the standard size, without which the tetramer undergoes an unphysical quaternary transition [33]. This technical support center provides guidance on diagnosing box size issues, quantitative data for planning simulations, and detailed protocols for optimizing your system size.
Troubleshooting Guides
Guide 1: Diagnosing an Inadequately Sized Simulation Box
Problem: Your simulation exhibits unphysical behavior, such as protein structural instability or abnormal material properties.
Steps to Diagnose:
- Check for Atoms Outside the Box: After trajectory processing (including re-centering and
nojumpcorrections), use tools likegmx checkto see if any atoms are reported as being outside the box. A small number of atoms may not be problematic, but consistent issues can indicate a box that is too tight [34]. - Analyze Structural Stability: For protein simulations, monitor the root-mean-square deviation (RMSD) of your protein's stable state (e.g., the T0 state of hemoglobin). A rapid, unphysical transition from this state suggests a lack of stabilizing forces, which can be a box size effect [33].
- Monitor Water Properties: Calculate the self-diffusion coefficient (D) of water molecules in your system. Compare this value to the expected value for your water model in a pure, large-scale water simulation. A significantly lower value indicates that the water structure and dynamics are perturbed by the proximity of the solute and its periodic images, a key sign of an undersized box [33].
- Check Density Profiles: Generate spatial density profiles for your solvent and solute. A non-uniform or lumpy density profile can indicate that the system is too small to accommodate natural fluctuations [22].
Guide 2: Resolving "Atoms Outside the Box" Errors
Problem: The gmx check utility reports atoms outside the simulation box even after using trjconv for re-centering and nojump unfolding.
Steps to Resolve:
- Don't Panic: A few atoms outside the box can "often occur and are normally not a problem" due to the nature of Periodic Boundary Conditions (PBCs) [34]. This may just change where the original copy of the atom appears.
- Clean the Trajectory: Use
gmx trjconvto thoroughly clean the trajectory from PBC effects. Ensure you are recentering the trajectory correctly, typically onto the protein or a main group of interest. - Verify Box Size Sufficiency: Use
gmx mindist(particularly with the-piflag) to check the minimum distance between your solute and the box boundary. The standard recommendation is at least five water molecules between any protein atom and the box boundary [33]. If this condition is not met, the box is likely too small. - Visualize: Load your trajectory into visualization software like VMD alongside the periodic box to visually confirm the system's behavior and the location of the reported atoms [34].
- Consider Re-running with a Larger Box: If analysis (as in Guide 1) confirms the box is too small, the most robust solution is to enlarge the simulation box and rerun the simulation.
Frequently Asked Questions (FAQs)
Q1: What is the standard rule of thumb for choosing an initial simulation box size? The conventional rule is that the periodic box should be large enough so that there is a minimum of five water molecules between any atom of the solute (e.g., protein) and the box boundary [33].
Q2: My simulation is computationally expensive. What is the optimal system size that balances precision and efficiency? The optimal size is system-dependent. A 2024 study on an epoxy resin (DGEBF/DETDA) found that a model size of ~15,000 atoms provided the best balance, offering precise predictions of mass density, elastic properties, strength, and thermal properties without sacrificing simulation efficiency [22]. However, for larger biomolecules, this size may be insufficient.
Q3: Why would a larger box size affect the stability of a protein's structure? Larger boxes are necessary for the accurate manifestation of long-range physical forces. For example, the stability of the unliganded human hemoglobin (T0) tetramer is governed by the hydrophobic effect. This effect arises from the disruption of the bulk water hydrogen bond network. In boxes that are too small, this network and the resulting "dewetting" phenomenon that stabilizes compact structures are not properly captured, leading to unphysical structural transitions [33].
Q4: Are there advanced simulation techniques that can help manage system size? Yes, methods like the Adaptive Resolution Simulation (AdResS) exist. This technique couples a high-resolution atomistic (AA) region around the molecule of interest with a surrounding coarse-grained (CG) region, allowing particles to move freely between them. This setup reduces the number of atoms that need to be simulated at full detail while aiming to reproduce the behavior of a full atomistic system, thereby saving computational cost [35].
Quantitative Data on System Size Effects
The following table summarizes key quantitative findings from a systematic study on the effect of MD model size on the predicted properties of an epoxy resin system. This data can serve as a benchmark for your own simulations [22].
Table 1: Impact of System Size on Precision and Simulation Time for an Epoxy Resin
| Number of Atoms | System Size (Monomers) | Average Density (g/cm³) | Standard Deviation of Density | Relative Simulation Time |
|---|---|---|---|---|
| 5,265 | 90/45 | 1.20 | Converged | Fastest |
| 10,530 | 180/90 | 1.20 | Converged | ... |
| 14,625 | 250/125 | 1.20 | Converged | ... |
| 20,475 | 350/175 | 1.20 | Converged | ... |
| 31,590 | 540/270 | 1.20 | Converged | ... |
| 36,855 | 630/315 | 1.20 | Converged | Slowest |
Conclusion from this study: For this specific epoxy system, a model size of 15,000 atoms was found to be optimal, providing precise predictions without the computational cost of larger systems [22].
Experimental Protocols
Protocol 1: Determining Optimal System Size for a New Protein
This protocol is based on the methodology used to validate human hemoglobin simulations [33].
1. Hypothesis: The simulation box must exceed a critical size to correctly capture the thermodynamic stability of the protein's native state.
2. System Setup:
- Molecular System: Prepare the initial structure of your protein (e.g., the T0 state of hemoglobin, PDB: 2DN2).
- Box Sizes: Construct a series of cubic simulation boxes of increasing size. For example, 75 Å, 90 Å, 120 Å, and 150 Å. Ensure protonation states of key residues (e.g., His146β in hemoglobin) are correct.
- Solvation and Ions: Solvate the protein in each box with a water model (e.g., TIP3P) and add ions (e.g., Na⁺ and Cl⁻) to achieve a physiological concentration (e.g., 0.15 mol/L).
- Replicates: Build multiple independent replicates (e.g., 5) for each system size to ensure statistical significance.
3. Simulation Parameters:
- Ensemble: Use the isothermal-isobaric (NPT) ensemble to maintain constant temperature and pressure.
- Thermostat/Barostat: Employ a thermostat like Nose-Hoover and a barostat.
- Force Field: Choose an appropriate force field (e.g., IFF, CHARMM, AMBER).
- Simulation Length: Run production simulations for a time sufficient to observe stability or transitions (e.g., 1 µs).
4. Data Analysis:
- Quaternary Structure Stability: Monitor the RMSD of the protein's quaternary structure from its starting configuration (e.g., T0 state). A stable simulation will maintain a low RMSD.
- Water Self-Diffusion Coefficient (D): Calculate D for water in each system. The value should match the expected bulk value for your water model in the largest box.
- Hydrogen Bond Analysis: Calculate the average number of hydrogen bonds per water molecule and its fluctuations. These should converge with increasing box size.
Protocol 2: A Systematic Approach for Polymer Resins
This protocol is adapted from a study seeking the optimal MD model size for an epoxy resin [22].
1. System Building:
- Monomer Mixing: Mix the polymer monomers (e.g., DGEBF and DETDA with a 2:1 molar ratio) in a periodic simulation box at a very low initial density (e.g., 0.111 g/cm³).
- Size Range: Create independent systems (replicates) across a range of atom counts (e.g., from ~5,000 to ~35,000 atoms).
2. Equilibration and Cross-Linking:
- Densification: Use a slow, staged process (e.g.,
fix/deformin LAMMPS) to gradually compress the low-density system to the target bulk density. - Annealing: Perform annealing simulations (e.g., heating from 27°C to 227°C and cooling back) to relax residual stresses.
- Cross-linking: Simulate cross-linking reactions using a protocol like REACTER in LAMMPS at elevated temperature, with defined cutoff distances and bond formation probabilities.
3. Property Prediction:
- Run simulations to predict key thermo-mechanical properties: mass density, elastic modulus, yield strength, and glass-transition temperature (Tg).
- Record the simulation time required for each system size.
4. Data Analysis:
- For each property and system size, calculate the mean and standard deviation across the replicates.
- Identify the smallest system size where the standard deviation of key properties has converged and the mean values are stable, while the simulation time remains acceptable.
Workflow Visualization
The following diagram illustrates the logical workflow for determining the optimal simulation box size, as described in the troubleshooting guides and experimental protocols.
The Scientist's Toolkit
Table 2: Essential Research Reagents and Software for MD System Setup
| Item Name | Type | Function / Application |
|---|---|---|
| LAMMPS | Software | A large-scale, massively parallel MD simulator used for building, cross-linking, and simulating material systems like polymers [22]. |
| GROMACS | Software | A high-performance MD software package primarily used for simulating proteins, lipids, and nucleic acids. Includes tools like gmx check, gmx trjconv, and gmx mindist [34]. |
| Interface Force Field (IFF) | Force Field | A force field used to describe atomic interactions, particularly for predicting physical, mechanical, and thermal properties of polymers and other materials [22]. |
| REACTER Protocol | Algorithm | A method implemented in LAMMPS to simulate chemical reactions, such as epoxy cross-linking, during an MD simulation based on predefined templates and cutoff distances [22]. |
| VMD | Software | A molecular visualization program used for displaying, animating, and analyzing large biomolecular systems. Essential for visually checking system setup and trajectory behavior [34]. |
| Nose-Hoover Thermostat/Barostat | Algorithm | A method used in MD simulations to control temperature and pressure, maintaining the desired thermodynamic ensemble (e.g., NPT) [22]. |
| AdResS (Adaptive Resolution Simulation) | Method | A simulation framework that couples a high-resolution atomistic region with a coarse-grained environment, reducing computational cost while aiming to reproduce full atomistic behavior [35]. |
Frequently Asked Questions (FAQs)
FAQ 1: What is the minimum acceptable simulation box size for studying protein oligomers in solution? The minimum acceptable box size is determined by the required distance between the protein atoms and the box edge. Research on lysozyme dimers and hexamers in crystallization solution found that a minimum offset of 1 nm is permissible. This distance prevents artifacts from periodic boundary conditions that could unrealistically influence protein copies in neighboring virtual boxes, while maintaining a balance between computational speed and result accuracy [4].
FAQ 2: I am getting a 'Residue not found in residue topology database' error in GROMACS. What should I do? This error means the force field you selected does not have a database entry for the residue 'XXX' in your structure file. Solutions include [36]:
- Check if the residue exists in the database under a different name and rename your residue accordingly.
- For terminal residues in AMBER force fields, ensure correct nomenclature (e.g., an N-terminal alanine should be 'NALA').
- If the residue is missing entirely, you cannot use
pdb2gmxautomatically. You will need to parameterize the molecule yourself, find a pre-existing topology file, or use another force field that has the parameters. - Use the
-ignhflag to allowpdb2gmxto ignore the hydrogen atoms in your file and add the correct ones itself.
FAQ 3: How can I increase the time step in my simulation of polymers? You can often increase the time step by imposing constraints on fast-moving bonds. For simulations of polymers and proteins, using constraint algorithms like SHAKE or P-LINCS on bond lengths and bond angles is common. Newer algorithms like ILVES-PC, which uses Newton's method, have been shown to solve constraint equations more accurately and efficiently, allowing for stable simulations with larger time steps [37].
FAQ 4: What does the 'Invalid order for directive' error mean in my GROMACS topology?
This error occurs when the directives in your .top or .itp files are in the incorrect sequence. The system topology has strict rules for the order of these directives. A common cause is placing a [ defaults ] or [ atomtypes ] directive after a [ moleculetype ] directive. The force field, including all atom types and parameters, must be fully defined before any molecules are declared [36].
Troubleshooting Common Errors
System Setup and Energy Minimization
| Error Message | Cause | Solution |
|---|---|---|
| 'Residue not found in residue topology database' [36] | The residue in the coordinate file does not match any entry in the selected force field's residue database. | 1. Check for naming discrepancies. 2. For terminal residues, use correct prefixes (e.g., NALA, CALA). 3. Manually provide a topology for the missing residue. |
| 'Long bonds and/or missing atoms' [36] | Atoms are missing from the initial coordinate file, causing pdb2gmx to place atoms incorrectly. |
Check the pdb2gmx output to identify the missing atom. Add the missing atoms to your PDB file using modeling software before processing. |
| 'Found a second defaults directive' [36] | The [ defaults ] directive appears more than once in the topology, which is invalid. |
Ensure the [ defaults ] directive appears only once, typically in the main force field file. Comment out or remove duplicate entries in included topology files. |
| 'Invalid order for directive defaults' [36] | The [ defaults ] section is not the first directive in the topology file. |
The [ defaults ] directive must be the very first in your topology. This is usually handled by the #include statement for the force field. |
| 'Out of memory when allocating' [36] | The system has run out of available memory (RAM) during a calculation or analysis. | 1. Reduce the number of atoms selected for analysis. 2. Shorten the trajectory being processed. 3. Check that your solvation step didn't create a massively oversized box. 4. Use a computer with more RAM. |
Production Run and Analysis
| Error Message | Cause | Solution |
|---|---|---|
| 'Atom index in position_restraints out of bounds' [36] | The atom indices in your position restraint file do not match the actual atom numbering for that specific molecule. | Ensure that position restraint files are included immediately after their corresponding [ moleculetype ] directive in the topology. The atom indices in the restraint file are local to that molecule. |
Quantitative Data for Simulation Box Sizing
The following data, derived from a study on lysozyme oligomers, provides guidance on minimum box sizes for protein simulations. The study established that a minimum offset of 1 nm was sufficient to correctly model the relative stability of dimers and hexamers [4].
Table: Minimum Box Size Validation for Lysozyme Oligomers [4]
| Oligomer | Minimum Offset (nm) | Box Edge Length (nm) | Simulation Time (ns) | Key Finding |
|---|---|---|---|---|
| Dimer | 1.0 | 8.6 | 1000 | Dimer was stable, as experimentally expected. |
| Dimer | 1.5 | 9.7 | 1000 | Dimer was stable, as experimentally expected. |
| Dimer | 2.0 | 10.6 | 1000 | Dimer was stable, as experimentally expected. |
| Hexamer | 1.0 | 10.9 | 700 | Hexamer was unstable, as experimentally expected. |
| Hexamer | 1.5 | 11.9 | 700 | Hexamer was unstable, as experimentally expected. |
| Hexamer | 2.0 | 12.9 | 700 | Hexamer was unstable, as experimentally expected. |
Experimental Protocol: Establishing Minimum Box Size
This protocol outlines the methodology used to determine the minimum acceptable simulation box size for protein crystallization solutions, based on the referenced lysozyme study [4].
Objective: To validate that a simulation box with a 1 nm offset between the protein and the box face produces physically accurate results for protein oligomer stability.
Workflow:
Detailed Steps:
System Preparation:
- Structure Source: Obtain initial coordinates for the protein oligomers (dimer and hexamer) from a relevant crystal structure (e.g., PDB ID: 6QWY for lysozyme) using a molecular visualization tool like PyMOL [4].
- Protonation: Assign protonation states of amino acid residues appropriate for the solution conditions (e.g., pH 4.5) using the PROPKA server [4].
- Box Creation: Center the oligomer in a cubic simulation box. Define multiple boxes where the minimum distance between any protein atom and the box face (the "offset") is 1.0 nm, 1.5 nm, 2.0 nm, etc. [4].
- Solvation and Ions: Solvate the system with an appropriate water model (e.g., TIP4P-Ew). Replace water molecules with ions to match the experimental precipitant concentration (e.g., 0.4 M NaCl). Add counterions as needed to neutralize the total system charge [4].
Simulation Parameters (Using GROMACS):
- Force Field: Amber ff99SB-ILDN [4].
- Energy Minimization: Use the steepest descent algorithm until the maximum force is below 1000 kJ/(mol·nm) [4].
- Equilibration:
- Production MD: Run for the required time (e.g., 1 μs for dimers, 700 ns for hexamers) with a 2 fs time step. Use the leap-frog integrator. Maintain temperature and pressure with the same thermostats and barostats. Constrain bond lengths using the LINCS algorithm. Calculate non-bonded interactions with a 1.0 nm cutoff and long-range electrostatics with the PME method [4].
Analysis:
- Process trajectories to remove periodic boundary effects and align structures (
gmx trjconv). - Calculate the following metrics for Cα atoms only [4]:
- Root-mean-square fluctuation (RMSF) using
gmx rmsf. - Root-mean-square deviation (RMSD) from the initial structure using
gmx rms. - Radius of gyration (Rg) using
gmx gyrate.
- Root-mean-square fluctuation (RMSF) using
- Validation Criterion: The simulation box size is considered sufficient if the relative stability of the oligomers matches experimental data (e.g., dimers remain stable while hexamers dissociate) [4].
- Process trajectories to remove periodic boundary effects and align structures (
The Scientist's Toolkit: Research Reagent Solutions
Table: Essential Materials and Software for Protein & Polymer MD Simulations
| Item | Function / Application |
|---|---|
| GROMACS | A versatile software package for performing MD simulations and analysis. It is highly optimized for speed and is widely used for biomolecular and polymer systems [4]. |
| AMBER ff99SB-ILDN | A force field specifically optimized for proteins and nucleic acids, providing accurate parameters for amino acid residues and their interactions [4]. |
| TIP4P-Ew | A four-site water model known for its accurate description of water properties under a wide range of conditions, making it suitable for detailed biomolecular simulations [4]. |
| LINCS | An algorithm used to constrain bond lengths in molecules, which allows for a longer integration time step (e.g., 2 fs) by eliminating the fastest vibrational motions [4]. |
| Particle Mesh Ewald (PME) | A standard method for accurately calculating long-range electrostatic interactions in periodic systems, which is critical for simulating charged proteins and polymers [4]. |
| ILVES-PC | A constraint algorithm for polymers and proteins that uses Newton's method. It can be integrated into GROMACS and offers superior speed and accuracy compared to SHAKE and P-LINCS, especially in parallel simulations [37]. |
| Specialized Polymer Force Fields | Custom parameterizations, such as those for poly(R-lactic acid) (R-PLA), are often required to accurately simulate the behavior of synthetic polymers and their conjugates with peptides [38]. |
Workflow Diagram: Simulation Box Size Optimization
This diagram illustrates the logical decision process for selecting and validating an appropriate simulation box size.
Troubleshooting Guide: Simulation Box Size and Shape
Frequently Asked Questions (FAQs)
Q1: Why does my simulation box keep shrinking during an NPT simulation?
A: This is typically caused by incorrect system topology or initial conditions that cause the molecules to condense. The system evolves in response to molecular attractions, causing inward collapse. In GROMACS, this often indicates that your force field parameters need refinement, particularly for complex molecules like surfactants or polymers. The solution involves verifying your topology generation process and ensuring initial coordinates are properly condensed for liquid-phase simulation [20].
Q2: How do I handle atoms reported as "outside the box" in trajectory analysis?
A: Atoms appearing "outside the box" are typically periodic boundary condition (PBC) artifacts, not actual errors. In GROMACS, use gmx trjconv with appropriate centering and PBC handling options (-pbc mol -center) to create continuous trajectories for analysis. For AMBER, use CPPTRAJ's center, unwrap, and autoimage commands to fix these artifacts [34] [39].
Q3: Can I dynamically change my box size during simulation to match a collapsing polymer?
A: Most MD packages don't support dynamic box resizing that adds/removes solvent molecules during a run. In LAMMPS, fix evaporate can slowly remove particles, but this generates disruptive phonons. A better approach is to run in chunks: when significant collapse occurs, restart with a smaller box, removing excess water, then re-equilibrate before continuing [40].
Q4: What causes segmentation faults when using large boxes and cut-offs?
A: In AMBER, segmentation faults occur when setting cut-offs too large (e.g., 450 in a 900Å box) due to 32-bit integer limitations and untested code paths for extreme length scales. The solution is to maintain standard cut-off values (e.g., 8-12Å) and ensure sufficient box padding (typically >10Å) between periodic images [41].
Q5: How can I change box dimensions to allow nanostructures to form properly?
A: For anisotropic structures, use anisotropic or triclinic pressure coupling. In LAMMPS, use fix box/relax aniso or tri to independently control box dimensions. In GROMACS, use pcoupl = anisotropic in your MDP file. This allows the box to change shape to accommodate asymmetric structures [42] [43].
Common Error Messages and Solutions
Table 1: Common Box-Related Errors and Solutions Across MD Packages
| Error Message | Software | Cause | Solution |
|---|---|---|---|
| "Atom index in position_restraints out of bounds" | GROMACS | Position restraint files included in wrong order | Ensure #include statements for position restraints immediately follow their respective molecule definitions [44] |
| "Segmentation fault" with large boxes | AMBER | 32-bit integer overflows with extreme system sizes | Reduce cut-off distance; ensure sufficient memory allocation [41] |
| "Pressure/forces becoming NaN or Inf" | LAMMPS | Close atomic contacts causing numerical overflow | Use delete_atoms overlap; run energy minimization; implement fix nve/limit [45] |
| "Unknown identifier in data file" | LAMMPS | Mistyped keywords in data file | Verify header syntax matches atom style; check data file consistency [45] |
| "Box tilting to extreme shapes" | LAMMPS | Excessive shear stress on liquid system | Verify external stress tensor doesn't exceed material yield stress; check system physical realism [43] |
Quantitative Data and Protocols
Box Size Optimization Parameters
Table 2: Key Parameters for Simulation Box Size Optimization
| Parameter | Recommended Value | Considerations | Software Implementation |
|---|---|---|---|
| Minimum box padding | 1.0-1.2 nm (10-12Å) from solute | Larger for charged systems; smaller for pure solvents | GROMACS: boxlen in gmx solvate; AMBER: buffer in tleap |
| Verlet buffer tolerance | 0.005 kJ/mol/ps per particle (default) | Determines pair-list update frequency | GROMACS: verlet-buffer-tolerance in MDP [46] |
| Maximum volume change per iteration | 0.001 (0.1%) | For stable box relaxation | LAMMPS: vmax 0.001 in fix box/relax [43] |
| Anisotropic pressure coupling | Semi-isotropic for membranes | Different x/y vs z pressures | GROMACS: tau_p and compressibility per direction |
| Neighbor list update frequency | Every 10-20 steps | Balance between performance and accuracy | GROMACS: nstlist; LAMMPS: neigh_modify [46] |
Box Resizing Experimental Protocol
Protocol 1: Manual Box Resizing for Polymer Collapse (LAMMPS/GROMACS)
- Initial Simulation: Run with extended polymer in large box until collapse begins
- Intermediate Analysis: Identify stable collapsed configuration and measure dimensions
- System Rebuilding:
- Preserve polymer coordinates
- Create new, smaller box with adequate padding (1.2-1.5× collapsed size)
- Remove waters outside new boundaries
- Add waters to fill new box if necessary
- Re-equilibration:
- Solvent energy minimization with polymer fixed
- Short NVT with position restraints on polymer
- Short NPT with decreasing restraints
- Production: Continue simulation with optimized box [40]
Protocol 2: Trajectory Fixing for PBC Artifacts (AMBER)
This protocol centers the system, unwraps coordinates across PBC, and re-images into a continuous trajectory [39].
Signaling Pathways and Workflows
Simulation Box Optimization Workflow
Box Relaxation Logic for Energy Minimization
The Scientist's Toolkit
Table 3: Essential Software Tools for Box Optimization Research
| Tool Name | Software Package | Primary Function | Key Commands/Options |
|---|---|---|---|
gmx energy |
GROMACS | Extract energy, volume, density timeseries | gmx energy -f simulation.edr -o volume.xvg |
fix box/relax |
LAMMPS | Box relaxation during minimization | fix 1 all box/relax iso 0.0 vmax 0.001 [43] |
CPPTRAJ |
AMBER | Trajectory analysis and PBC correction | center, image, autoimage, unwrap [39] |
gmx trjconv |
GROMACS | Trajectory conversion and PBC handling | -pbc mol -center -ur compact [34] |
gmx check |
GROMACS | Check trajectory for errors | gmx check -f trajectory.xtc [34] |
MDAnalysis |
Python | Trajectory analysis and manipulation | unwrap(), center_in_box(), fit_rot_trans() [39] |
MDAnalysis |
Python | Trajectory analysis and manipulation | unwrap(), center_in_box(), fit_rot_trans() [39] |
parmchk2 |
AMBER | Parameter checking for novel molecules | parmchk2 -i molecule.mol2 -f mol2 -o frcmod [20] |
Identifying and Resolving Box Size-Related Simulation Artifacts
Troubleshooting Guide: "Simulation Box Too Small" Errors
This guide helps researchers diagnose and resolve a common error in Molecular Dynamics (MD) simulations where the simulation box shrinks or triggers fatal errors, potentially compromising simulation stability and scientific results.
Why does my simulation box keep shrinking during the production run?
Problem: You observe a continuous decrease in simulation box volume and density over time during an NPT (constant Number of particles, Pressure, and Temperature) simulation. The molecules appear to be collapsing inward.
Diagnosis: This behavior indicates that your system is condensing. The primary causes are:
- Incorrect Initial Conditions: The molecules in the initial configuration were placed too far apart, creating an artificially low starting density. The barostat then correctly works to compress the system to its true equilibrium density [20].
- Inaccurate Force Field Topology: The attractions defined within and between molecules in your topology file cause them to collapse inward. If the volume and density stabilize at an incorrect value after a sufficiently long simulation, this strongly suggests that your molecular topology is incorrect and needs refinement [20].
Solution:
- Refine Your Topology: For large or complex molecules (e.g., lipids, surfactants, polymers), avoid parametrizing the entire molecule at once. Instead, break it down into smaller, representative chemical moieties and parametrize these pieces separately before linking them together. This ensures consistent charge assignment and geometry [20].
- Validate Against Literature: Use previously reported topology parameters for your molecule if available, as this can save significant time and effort [20].
- Check System Preparation: Ensure your initial system represents the physical state you want to simulate. To model a liquid droplet, start with a concentrated starting point, not individual molecules scattered in a vacuum [20].
How do I resolve the fatal error "Simulation box is too small" in HOOMD-blue?
Problem: Your simulation fails with an error similar to: "nlist: Simulation box is too small! Particles would be interacting with themselves." [47]
Diagnosis: This error is a direct consequence of the minimum image convention used with Periodic Boundary Conditions (PBC). It occurs when the simulation box is too small to prevent a particle from interacting with its own periodic image.
The specific rule is: The shortest box dimension (L_min) must be greater than twice the sum of your potential's cutoff distance (r_cut) and the neighbor list buffer (r_buff): 2 * (r_cut + r_buff) < L_min [47].
Solution:
- Increase Box Size: The most straightforward solution is to increase the size of your simulation box, which involves adding more particles (e.g., water and ions) to maintain the correct concentration [47].
- Reduce Cut-off Distance: If scientifically justified for your system, you can reduce the cut-off distance (
r_cut) of your non-bonded interactions [47]. - Adjust the Neighbor List Buffer: You can decrease the neighbor list buffer (
r_buff). Be aware that this will increase the frequency of neighbor list rebuilds, which may slightly impact performance. For small systems, this performance hit is usually negligible [47].
What is the minimum acceptable size for a simulation box?
Problem: You want to use the smallest possible box to save computational resources but are unsure how small you can go without introducing artifacts.
Diagnosis: A box that is too small can lead to artificial periodicity, where a particle interacts too strongly with its own images, distorting the results. A common threshold is to ensure the minimum distance between any protein atom and the box face is at least 1.0 nm to 1.5 nm [4].
Solution: A study on lysozyme oligomers established that an offset (the minimum distance between the protein atoms and the box face) of 1.0 nm was the minimum permissible for studying protein crystallization solutions without losing accuracy. Larger offsets (e.g., 1.5 nm or 2.0 nm) provide a greater safety margin and are often recommended [4].
Frequently Asked Questions (FAQs)
Q: How can I calculate the initial size of my simulation box? A: The box must be large enough to avoid self-interactions across the periodic boundary. For a simple system, a good starting point is to place your solute(s) in the center and ensure a minimum offset (e.g., 1.0 to 1.5 nm) between the solute and the box edges in all dimensions. For more complex systems like a water-chloroform interface, a reasonable starting box might be 4 x 4 x 8 nm, which can then be equilibrated under pressure [30].
Q: I am simulating in vacuum. Why is my box collapsing? A: Placing a few molecules in a large vacuum box models a gas-phase system. The observed box shrinkage is the natural condensation of these gas-phase molecules into a droplet under the influence of the barostat. This is an expected physical outcome, not necessarily an error, but it may not represent the liquid-phase environment you intended to model [20].
Q: My simulation fails with a domain decomposition error in GROMACS. Is this related to box size? A: Yes. This error occurs when the simulation box is too small to be efficiently divided across the number of processor ranks (MPI processes) you are using. The solution is to run the simulation on fewer MPI ranks (often just one) and use more OpenMP threads instead, or to use a larger simulation box [30].
Experimental Protocol: Determining Minimum Box Size
The following protocol, adapted from published research, provides a methodology to empirically determine the minimum acceptable box size for a given system [4].
1. System Preparation:
- Oligomer Selection: Select stable and unstable oligomers of your protein of interest. For example, in a lysozyme crystallization solution, a stable dimer and an unstable hexamer can be used [4].
- Model Creation: Extract structural models from a known crystal structure (e.g., using PyMOL).
- Protonation: Assign protonation states to amino acid residues at the desired pH using a server like PROPKA [4].
- Box Generation: Create multiple cubic simulation boxes with the solute at the center. Vary the box size so the minimum distance between the solute atoms and the box face (the "offset") is 1.0, 1.5, 2.0, and 2.5 nm [4].
- Solvation and Ions: Fill the box with an appropriate water model (e.g., TIP4P-Ew). Add ions to match the experimental precipitant concentration and neutralize the system's total charge [4].
2. Simulation Parameters:
- Force Field: Use a modern force field (e.g., Amber ff99SB-ILDN within GROMACS) [4].
- Energy Minimization: Use the steepest descent method until the maximum force is below 1000 kJ/(mol·nm) [4].
- Equilibration:
- NVT Ensemble: Equilibrate for 100 ps using a thermostat (e.g., V-rescale).
- NPT Ensemble: Equilibrate for 100 ps using a thermostat and a barostat (e.g., Parrinello-Rahman).
- Production MD: Run multiple relatively long simulations (e.g., 700 ns to 1 μs) for each box size in the NPT ensemble. Use a 2-fs time step, the PME algorithm for electrostatics, and constrain bond lengths with LINCS [4].
3. Data Analysis:
- Stability as Benchmark: The correct box size is the smallest one in which the known-stable oligomer (e.g., dimer) remains stable, while the known-unstable oligomer (e.g., hexamer) dissociates, matching experimental evidence [4].
- Metrics: Calculate and compare the Root-Mean-Square Fluctuation (RMSF), Root-Mean-Square Deviation (RMSD), and Radius of Gyration (Rg) for the oligomers across different box sizes [4].
Diagram 1: Workflow for determining the minimum acceptable simulation box size.
Quick Reference Tables
Table 1: Common "Box Too Small" Errors and Immediate Fixes
| Error Message / Symptom | Software | Primary Cause | Immediate Solution |
|---|---|---|---|
| Continuous box shrinkage | GROMACS (NPT) | Incorrect initial density or force field [20] | Check topology; ensure initial configuration is realistic |
| "Simulation box is too small! Particles would be interacting with themselves." | HOOMD-blue | 2 * (r_cut + r_buff) > Shortest box dimension [47] |
Increase box size; reduce r_cut or r_buff |
| "No domain decomposition ... minimum cell size" | GROMACS | Box too small for the number of MPI ranks [30] | Run on a single MPI rank; increase -rdd; use larger box |
Table 2: Recommended Minimum Offsets for Different Systems
| System Type | Recommended Minimum Offset | Key Consideration |
|---|---|---|
| Protein Crystallization Solutions [4] | 1.0 nm | Established as minimum for correct oligomer stability |
| Typical Solvated Protein / Polymer | 1.5 nm | Provides a good safety margin for most biomolecular systems |
| Systems with Long Cut-offs | >1.5 nm | Necessary if r_cut is large to satisfy 2*r_cut < L_min |
Table 3: Research Reagent Solutions
| Item | Function in Simulation Box Optimization |
|---|---|
| GAFF/GAFF2 Force Field | Provides parameters for small organic molecules or drug-like compounds; must be applied carefully to large molecules [20]. |
| Amber ff99SB-ILDN | A high-quality force field for proteins, often used in benchmark studies for box size determination [4]. |
| TIP4P-Ew Water Model | A four-site water model known for its accuracy, used to solvate the system and reproduce correct liquid densities [4]. |
| Parrinello-Rahman Barostat | A popular barostat for pressure coupling in NPT simulations, allowing the box size and shape to fluctuate correctly [4]. |
| PME (Particle Mesh Ewald) | The standard algorithm for handling long-range electrostatic interactions in periodic systems, essential for energy calculations [4]. |
Diagram 2: Logical troubleshooting flowchart for simulation box errors.
Frequently Asked Questions (FAQs)
Q1: My trajectory analysis shows a warning about atoms being outside the simulation box. Does this mean my simulation is invalid and I need to rerun it with a larger box?
Not necessarily. In molecular dynamics (MD) with Periodic Boundary Conditions (PBCs), the "box" is replicated infinitely in all directions. An atom "outside" one box is simultaneously inside an adjacent image. The gmx check tool may report this, but it often notes that "These may occur often and are normally not a problem" [34]. The key is to correct the trajectory for meaningful analysis, not to rerun the simulation, unless a specific artifact is identified.
Q2: I have already used gmx trjconv for PBC correction, but some analyses still seem affected. What else can I do?
Different analyses require different PBC-handling strategies. A single trjconv command might not suffice for all purposes. For instance, calculating the Root-Mean-Square Deviation (RMSD) requires a continuous trajectory where the protein does not jump across box edges. A combination of -pbc nojump (to remove jumps) followed by -fit rot+trans (to align structures) is often necessary for a clean analysis [48]. The choice of molecule for centering (e.g., the protein backbone) is also critical.
Q3: How do I know if my simulation box is truly too small and needs to be enlarged for future simulations?
A box that is too small can cause artifacts because a molecule may interact with its own periodic image. You can investigate this using tools like gmx mindist with the -pi flag to monitor the minimum distance between your protein and its periodic images [34]. A recent study on lysozyme oligomers established that a minimum distance of 1 nm between the protein atoms and the box face was the smallest permissible offset, as smaller boxes led to unrealistic destabilization of specific oligomers [4]. If this distance becomes too small during simulation, your box should be enlarged.
Troubleshooting Guide: Common Problems and Solutions
| Problem Description | Primary Cause | Recommended Solution | Key GROMACS Command(s) |
|---|---|---|---|
Atoms reported "outside the box" by gmx check. |
Normal PBC artifact; the trajectory is stored with molecules broken across box boundaries. | Apply PBC correction to output a continuous trajectory for analysis. | gmx trjconv -pbc mol -center [48] |
| Jumps in the trajectory causing spikes in RMSD analysis. | Molecules crossing the box boundary and appearing on the opposite side. | Use the nojump option to remove these discontinuous jumps. |
gmx trjconv -pbc nojump [34] [48] |
| Protein complex is split across the box, distorting interactions. | The entire complex was not properly recentered as a whole. | Center the trajectory on the protein (or complex) of interest. | gmx trjconv -center -pbc mol -ur compact [48] |
| Suspected artificial interaction with periodic images. | Simulation box is too small. | Check minimal distance to own image; if small, rerun with a larger box (e.g., >1 nm offset) [4]. | gmx mindist -pi [34] |
Trajectory Correction Workflow
The following diagram outlines a standard workflow for processing an MD trajectory to correct for PBC effects and prepare it for analysis.
Quantitative Data on Box Size Optimization
Recent research provides quantitative guidelines for selecting an appropriate simulation box size. The following table summarizes findings from a study investigating the minimum acceptable box size for lysozyme oligomers, ensuring results are not compromised by finite-size effects [4].
| System | Minimum Offset (nm) | Box Edge (nm) | Key Finding on Stability |
|---|---|---|---|
| Lysozyme Dimer | 1.0 | 8.6 | Dimer remained stable even in the smallest box (1 nm offset). |
| Lysozyme Hexamer | 1.0 | 10.9 | Hexamer was unstable (dissociated) in all box sizes, as expected from experiment. |
| Recommendation | 1.0 | System-dependent | A minimum distance of 1 nm between protein atoms and the box face is the established lower limit [4]. |
Experimental Protocol: Trajectory Correction for RMSD Analysis
This protocol details the steps to generate a continuous, aligned trajectory suitable for RMSD and other structural analyses [34] [48].
Remove Jumps: The first step is to ensure molecules do not jump across the box. This creates a trajectory where the center of mass of each molecule moves continuously.
When prompted, select the group (e.g., "Protein") for handling PBC.
Align to Reference Structure: To remove global rotation and translation, fit the trajectory to a reference structure (usually the first frame).
When prompted, select the group for least-squares fitting (e.g., "Protein Backbone") and the group to output (e.g., "System" or "Protein").
Center in Box (Optional but Recommended): For visualization or specific analyses, ensure the protein is centered in the middle of the box.
When prompted, select the group to center in the box (e.g., "Protein") and the group for output (e.g., "System").
The Scientist's Toolkit: Essential Software and Commands
| Item | Function in Trajectory Correction |
|---|---|
GROMACS (gmx trjconv) |
The primary tool for manipulating trajectories, handling PBC, centering, and fitting [34] [48]. |
-pbc nojump flag |
Corrects the trajectory so that molecules do not discontinuously "jump" across periodic boundaries [48]. |
-fit rot+trans flag |
Fits the trajectory to a reference structure to remove overall rotation and translation, essential for RMSD [48]. |
-center flag |
Moves a selected group of atoms (e.g., your protein) to the center of the simulation box [48]. |
GROMACS (gmx check) |
A utility that checks the integrity of topology and trajectory files, often reporting atoms "outside the box" [34]. |
GROMACS (gmx mindist) |
Calculates the minimum distance between two groups, which can be used with the -pi flag to check the distance to periodic images [34]. |
Optimizing Cutoff Radius and Buffer Settings for Efficiency
Conceptual Foundations: Understanding Cutoff Radius and Buffer Settings
What are the cutoff radius and buffer settings, and why are they critical for my simulation?
The cutoff radius is a predetermined distance beyond which non-bonded particle-particle interactions are neglected or approximated. Using a cutoff transforms an O(N²) computational problem into a more manageable O(N) one, drastically reducing the number of pairwise calculations required at each simulation step [49]. The buffer (or "skin") region is an additional zone surrounding the cutoff within which particles are considered potential neighbors between periodic neighbor-list updates. Proper settings are foundational to simulation box size optimization, as they directly determine the minimum required spacing between a protein and its periodic images to prevent artifacts [4].
How do these parameters impact the trade-off between computational speed and physical accuracy?
The primary trade-off is straightforward: a shorter cutoff speeds up computation but risks truncating significant long-range interactions, potentially leading to physical inaccuracies. A larger buffer reduces the frequency of neighbor-list updates (saving computation time) but increases the size of the neighbor list, making each update slower and consuming more memory.
- Computational Cost: Using a cutoff makes computational work scale linearly with the number of particles, which is essential for simulating large systems [49].
- Accuracy Risk: Cutting off interactions too abruptly can cause energy drift and artifacts. A switching function that smoothly reduces the interaction force to zero at the cutoff distance is often employed to mitigate this [49].
- Buffer Zone Impact: A larger buffer means the neighbor list remains valid for more steps, but it also includes more particles that are not within the cutoff at the time of the list build, leading to some unnecessary calculations.
Optimization Strategies and Protocols
What is the recommended methodology for determining the optimal cutoff radius?
The optimal cutoff is a balance between computational efficiency and scientific validity for your specific system.
- Consult Your Force Field: Always start with the recommended cutoff value specified by the force field you are using. This is non-negotiable for maintaining the parametrization's integrity.
- Benchmark Performance: Conduct short test runs with different cutoff values (e.g., 0.9 nm, 1.0 nm, 1.1 nm, 1.2 nm) and compare the nanoseconds-per-day simulation throughput.
- Validate Physically: For your final candidate cutoffs, run longer simulations and check key physical properties (e.g., energy conservation, radial distribution functions, diffusion coefficients) against a reference simulation with a very long cutoff or experimental data.
The following table summarizes a systematic protocol for this optimization, adaptable for tools like GROMACS:
Table: Experimental Protocol for Cutoff and Buffer Optimization
| Step | Action | Measurement | Goal |
|---|---|---|---|
| 1. System Setup | Create a simulation box with a proven sufficient protein-to-wall offset (e.g., ≥ 1.0 nm) [4]. | Box dimensions, total atom count. | Establish a valid baseline system. |
| 2. Performance Screening | Run multiple short simulations (e.g., 1-5 ns), varying rcoulomb and rvdw. |
Simulation throughput (ns/day). | Identify settings that offer the best speedup. |
| 3. Physical Validation | Run longer simulations (e.g., 50-100 ns) with the fastest parameter sets from Step 2. | Energy drift, RMSD, Rg, protein oligomer stability [4]. | Ensure physical accuracy is not compromised. |
| 4. Buffer Calibration | Using the chosen cutoff, test different nstlist values (e.g., 10, 20, 40) with a fixed buffer. |
Simulation throughput; check for "lost particle" errors. | Find the update frequency that maximizes performance. |
How do I optimize the neighbor-list buffer size and update frequency?
The buffer size (rlist) and update frequency (nstlist in GROMACS) are interdependent. The buffer must be large enough that no particle can move from outside to inside the cutoff within nstlist steps. A general guideline is:
Buffer Size ≈ Maximum particle displacement per step × nstlist × Safety Factor
A typical safety factor is 2-3. In practice, a buffer of 0.1 to 0.2 nm is common. You can optimize this by:
- Monitoring Performance: Profile your simulation with different
nstlistvalues. There is a sweet spot where the cost of more frequent list updates is balanced by the cost of maintaining an excessively large list. - Verifying Stability: If the buffer is too small, the simulation will crash with errors about particles being "lost" from the neighbor list.
What is the minimum safe distance between my protein and the box edge?
Research on lysozyme oligomers in crystallization solutions provides a concrete guideline. The stability of biologically relevant dimers versus non-relevant hexamers was used as an accuracy metric across different box sizes [4].
Table: Minimum Box Size Validation from Lysozyme Oligomer Stability Studies [4]
| Minimum Protein-Wall Offset | Observation | Recommendation |
|---|---|---|
| 1.0 nm | Dimers remained stable; hexamers correctly identified as unstable. | This is the minimum permissible offset for studying protein crystallization solutions. |
| < 1.0 nm | High risk of serious artifacts due to unrealistic interactions between periodic protein images. | Not recommended. |
This study establishes that a 1.0 nm offset is the lower limit, but using a larger margin (e.g., 1.5 nm) is often prudent for systems where protein flexibility or large-scale motion is expected. The workflow below summarizes the relationship between core parameters and the optimization process.
Diagram: Parameter Optimization Workflow
Troubleshooting Common Problems
My simulation is slower than expected. How can cutoff and buffer settings be the cause?
- Problem: An excessively short cutoff forces the neighbor list to be updated very frequently (
nstlistmust be small), making the update overhead dominate the computational cost. - Solution: Increase the cutoff slightly. This might seem counterintuitive, but a longer cutoff can allow for a larger buffer and a higher
nstlistvalue, reducing the frequency of expensive neighbor-list rebuilds. Find the balance for your specific system.
I am encountering "particles lost in the domain" or similar neighbor-list errors.
- Problem: The neighbor-list buffer is too small for the chosen update frequency. Particles move further than anticipated between list updates, breaking the simulation's logic.
- Solution: Increase the buffer size (
rlist) or reduce the neighbor list update frequency (nstlist). Increasing the buffer is the safer and more common approach.
After changing to a shorter cutoff, my protein structure becomes unstable or my energy conservation is poor.
- Problem: The cutoff is too short, truncating significant long-range electrostatic or van der Waals interactions. This is a problem of physical accuracy, not just performance.
- Solution: Revert to the force field's recommended cutoff. For electrostatics, always use a long-range method like Particle Mesh Ewald (PME) [50] [4] instead of a plain cutoff. Do not use a cutoff shorter than 1.0 nm for Coulomb interactions without PME.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table: Key Components for Simulation Box Optimization Studies
| Item / Software | Function / Role in Research |
|---|---|
| GROMACS | A high-performance MD simulation package used extensively for benchmarking and production runs, with robust support for PME and neighbor-list optimizations [50] [4]. |
| AMBER ff99SB-ILDN | An example of a well-parameterized force field; provides the physical model and recommended cutoff values for interatomic interactions [4]. |
| TIP4P-Ew Water Model | A specially parameterized water model used to solvate the system, contributing to an accurate representation of the solvent environment [4]. |
| Particle Mesh Ewald (PME) | The standard algorithm for handling long-range electrostatic interactions, allowing for an accurate treatment of forces beyond the short-range cutoff [50] [4]. |
| Lysozyme Oligomers (Dimers/Hexamers) | A model system used experimentally to validate the minimum box size by comparing the stability of relevant vs. non-relevant oligomeric states [4]. |
Managing Periodic Image Interactions and Artifacts
Core Concepts: Periodic Boundary Conditions and Artifacts
What are Periodic Boundary Conditions (PBC) and why are they used? Molecular dynamics simulations are computationally expensive, and simulating a small, isolated system does not reflect the reality inside cells. Periodic Boundary Conditions (PBC) imitate an infinite space by treating the simulation box as a unit cell that is replicated infinitely in all directions. These copies are called "periodic images." This approach ensures that when a molecule exits one side of the box, it re-enters from the opposite side, conserving the number of atoms and allowing particles to experience forces as if they were in a bulk solution [51] [52].
What are the common artifacts associated with PBC? The primary artifacts arise from the fundamental nature of PBC and its implementation:
- Visual Discontinuities: Atoms or molecules may appear to drift out of the central simulation box, making it look like the system is disintegrating or "exploding." In reality, these atoms have simply moved into a neighboring periodic image and continue to interact with the system [51].
- Interaction Artifacts: Without proper handling, an atom could interact with multiple images of the same atom or with itself across the periodic boundaries [52].
- Edge Discontinuities in Analysis: When performing operations like a Fourier transform on simulation data, the sharp discontinuities at the edges of the box can create cross-patterned artifacts in the resulting frequency spectrum [53] [54].
- Incorrect Visualization of Complex Systems: In simulations containing multiple components, like a protein and a membrane, choosing an incorrect center for visualization can make it appear as though the protein is incorrectly crossing the membrane or that there is "vacuum" in the box [51] [55].
Frequently Asked Questions (FAQs) and Troubleshooting
FAQ 1: Why do my molecules leave the simulation box during the run? Is my simulation exploding? Answer: No, this is normal and does not indicate an "explosion" or error in your simulation [51]. MD simulations often store atomic coordinates without resetting them when they cross the box boundary. An atom that has left the box is simply residing in a neighboring periodic image and continues to interact with the system. This behavior is a direct consequence of using PBC and is preferred for maintaining numerical precision and tracking the total drift of atoms [51].
FAQ 2: How can I fix the visualization so my entire molecule is in one box? Answer: You can achieve this through a process called "wrapping." Wrapping places all atoms back into a single central box for visualization by translating them by integer multiples of the box vectors [51]. The key is to select an appropriate wrapping center.
- For a single protein in solvent, center the box on the protein's average coordinates.
- For two proteins, typically center on one of them, especially if unbinding events occur.
- For a protein-membrane system, you may need to center on a point between the protein and the membrane to keep both intact in the visual output [51].
FAQ 3: My membrane appears broken after centering and applying PBC corrections. What went wrong?
Answer: This is a common issue when the PBC correction workflow is not optimal for multi-component systems. A forum post on GROMACS troubleshooting highlights this exact problem [55]. The artifact occurs when the commands used to make the protein/membrane complex whole again are not applied in the correct order or with the correct grouping. The solution is to carefully use a combination of the -center, -pbc mol, and -pbc whole options in tools like gmx trjconv, ensuring that the entire complex (e.g., a group like "SOLU_MEMB") is selected for output to maintain molecular integrity [55].
FAQ 4: How do I avoid self-interaction or double-counting of interactions in PBC? Answer: This is prevented by implementing two key concepts [52]:
- Minimum Image Convention (MIC): This rule ensures that an atom interacts only with the closest periodic image of every other atom in the system.
- Cut-off Radius: A distance cutoff (( r{\text{cut}} )) is defined for non-bonded interactions. To fulfill the MIC, this cutoff must not exceed half the length of the smallest box side for a cubic box: ( r{\text{cut}} \leq L/2 ).
FAQ 5: How can I reduce edge artifacts in analysis, like Fourier transforms? Answer: The Periodic Plus Smooth (P+S) Decomposition method is an effective solution [53] [54]. This technique decomposes an image (u) into a periodic component (p) and a smooth component (s), such that u = p + s. The smooth component is calculated by solving Poisson's equation with boundary conditions set by the intensity gaps at the image edges. Subtracting this smooth component removes the edge discontinuities, thereby reducing the cross-pattern artifacts in the Fourier transform without degrading the signal from the entire field of view [54].
Experimental Protocols
Protocol 1: Wrapping a Trajectory for Visualization
This protocol uses the Python moleculekit library to wrap a simulation trajectory around a molecule of interest [51].
- Objective: Center the simulation box on a specific molecule to achieve a visually coherent trajectory.
- Materials: A molecular dynamics trajectory file (e.g., .xtc, .dcd) and its corresponding topology file (e.g., .prmtop, .psf).
- Troubleshooting: If the initial wrapping selection (e.g., "protein") does not give the desired result, visualize the system first (
mol.view()) and experiment with more specific selection commands, for examplemol.wrap("resid 15 and chain A")to wrap around a specific residue [51].
Protocol 2: Applying Periodic Plus Smooth Decomposition
This protocol is used to reduce edge artifacts in 2D Fourier transforms of images, which can be adapted for analysis of 2D density maps from simulations [54].
- Objective: Remove the cross-pattern artifact from a Fourier transform caused by periodic boundary discontinuities.
- Materials: A 2D image or array (e.g., an electron density map).
The Scientist's Toolkit
Research Reagent Solutions
| Tool / Reagent | Function in Managing PBC |
|---|---|
| Visualization Software (VMD) | Displays trajectories and allows for the visualization of periodic images. It can also center molecules and handle PBC through its trajectory processing menus [56]. |
Trajectory Processing Tools (gmx trjconv) |
A GROMACS utility for post-processing trajectories. It is essential for operations like centering, PBC correction (-pbc mol, -pbc whole), and removing atom jumps across box boundaries (-pbc nojump) [55]. |
Python (moleculekit library) |
Provides a programmable interface for reading, manipulating, and wrapping molecular dynamics trajectories and topologies [51]. |
Python (numpy, scipy) |
Offer the foundational mathematical and Fourier transform capabilities required to implement advanced artifact reduction algorithms like the Periodic Plus Smooth Decomposition [54]. |
| Cut-off Radius (( r_{\text{cut}} )) | A simulation parameter critical for correctly handling non-bonded interactions under PBC. It must be set to ≤ ( L/2 ) (for cubic boxes) to comply with the minimum image convention and prevent interaction artifacts [52]. |
Workflow and Relationship Diagrams
PBC Correction and Visualization Workflow
The diagram below outlines the logical workflow for correcting and visualizing a molecular dynamics trajectory, incorporating key decision points for handling different system types.
PBC Artifact Reduction Methods
This diagram categorizes the main types of PBC-related artifacts and connects them to the appropriate correction methodologies.
Box Size Optimization for Enhanced Sampling and Free Energy Calculations
Table of Contents
- Core Concepts: Why Box Size Matters
- Troubleshooting FAQs
- Quantitative Data Summary
- Experimental Protocols
- Workflow and System Setup Diagrams
- The Scientist's Toolkit
Core Concepts: Why Box Size Matters
In molecular dynamics (MD) simulations, the simulation box defines the periodic boundary conditions (PBCs) within which your system exists. Optimizing its size is critical for balancing computational cost against the accuracy of calculated thermodynamic and kinetic properties, especially for free energy calculations [3].
A box that is too small can introduce finite-size artifacts, where the periodic images of your solute interact in an unphysical way, distorting the simulated properties. Conversely, an excessively large box needlessly increases the number of solvent atoms, making the simulation computationally prohibitive [3]. The key is to ensure a sufficient distance between the solute and its periodic images, allowing for a fully formed solvation shell and preventing direct interaction between periodic copies. For most biomolecular systems in explicit solvent, a minimum distance of 1.0 nm to 1.5 nm between the solute and the box edge is considered a best practice [3]. Proper box size ensures that long-range electrostatic interactions, typically handled by methods like the Particle-Mesh Ewald (PME) algorithm, are correctly screened by the solvent and not artificially enhanced by the proximity of periodic images [3].
Troubleshooting FAQs
Q1: My solvation free energy calculation shows a strong dependence on the simulation box size. What is the most likely cause?
A: This is typically a symptom of inadequate sampling rather than a genuine physical effect. When statistical samples are insufficient, random fluctuations can create illusory trends. One study demonstrated that a box-size dependency in the hydration free energy of anthracene disappeared when the results were averaged over 20 independent repeats, whereas examining individual trajectories could misleadingly suggest upward, downward, or non-monotonic trends [3]. Solution: Prioritize extensive sampling and multiple independent repeats (at least 5-10) to assign reliable uncertainties to your results before investigating other potential causes.
Q2: Under what specific conditions does a small box size genuinely introduce artifacts?
A: A genuine box-size artifact occurs when the distance between the solute and its periodic image is too small, leading to direct interaction. This is especially problematic:
- In Vacuum Simulations: Without a solvent to provide electrostatic screening, even moderately small boxes can cause strong, unphysical interactions between periodic images when using PME for electrostatics [3].
- In Solvated Systems with Reduced Screening: If the solvent's screening capability is compromised (e.g., using a low-dielectric solvent or scaled charges on water), box-size dependence can emerge [3].
- When the Buffer is Too Small: If the distance from the solute to the box edge is less than ~1.0 nm, the solvation shells of periodic images can interact, and the Verlet pair list buffer may be insufficient, leading to energy drifts and inaccurate forces [46] [3].
Q3: How does simulation box size relate to the Verlet cutoff scheme and energy conservation?
A: The Verlist cutoff scheme uses a pair list, updated periodically, of atom pairs within a cutoff distance (rlist). To account for atom diffusion between updates, a buffer (r_b) is used, making the list cutoff larger than the interaction cutoff (r_c). The box size must accommodate this buffer. If the box is too small or the buffer is insufficient, atoms outside the list cutoff can move within the interaction cutoff between list updates, causing missed interactions and a steady energy drift [46]. GROMACS can automatically determine the buffer size to keep the energy drift below a tolerable level (default: 0.005 kJ/mol/ps per particle) [46].
Q4: Are box size effects more pronounced for large molecules like proteins compared to small ligands?
A: Research indicates that for sufficiently large boxes (maintaining a >1.0 nm buffer), the solvation free energy of both small molecules and proteins (e.g., the GB protein) shows no significant box-size dependence when proper sampling and electrostatic treatments are applied [3]. The primary risk for larger solutes is the increased computational temptation to use a minimally small box, which can inadvertently push the system into the artifact-prone regime.
The following tables summarize key quantitative findings from research on box-size effects.
Table 1: Box Size Independence in Solvation Free Energy Calculations
| System Type | Box Size Range (Number of Water Molecules) | Key Finding | Statistical Notes |
|---|---|---|---|
| Small Molecule (Anthracene) [3] | 473 to 5334 | No statistically significant trend in computed hydration free energy observed. | Conclusion required 20 independent repeats per box size; single realizations showed random, misleading trends. |
| Protein (GB) [3] | Varying box dimensions | No box size dependence for electrostatic component of solvation free energy in water. | Required a >1.0 nm distance between protein surface and box edge; used Hamiltonian replica exchange for enhanced sampling. |
Table 2: Impact of System Conditions on Box Size Dependence
| System Condition | Observed Box Size Dependence? | Underlying Physical Reason |
|---|---|---|
| Solvated Protein (Standard Water Model) [3] | No | Water molecules provide sufficient electrostatic screening between periodic images. |
| Protein in Vacuum (with PME) [3] | Yes | Absence of screening leads to direct, unphysical electrostatic interactions between images. |
| Solvated Protein with Low-Screening Solvent [3] | Yes | Reduced solvent screening strength makes the system behave more like a vacuum. |
| Extremely Small Solvated Box (< 0.5 nm buffer) [3] | Yes | Solvation shells of periodic images directly interact, and water cannot properly screen. |
Experimental Protocols
Protocol 1: Verifying Box Size Independence for Solvation Free Energy
This protocol outlines a robust methodology to empirically verify that your calculated solvation free energy is independent of simulation box size.
System Preparation:
- Generate multiple systems of your solute (e.g., a small molecule or protein) solvated in boxes of different sizes. A recommended range is from a minimal box (just exceeding 2x the non-bonded cut-off in all dimensions) to a box with a buffer of 1.5-2.0 nm from the solute.
- Use a consistent force field and water model for all systems.
- Ensure proper protonation states and employ tools like
CHAPERONgorStreaMDto automate and standardize the setup process across all boxes [57] [58].
Equilibration:
- Carry out a thorough energy minimization and equilibration protocol for each system (NVT and NPT ensembles) to ensure identical temperature and pressure conditions.
- For complex solutes like proteins, consider advanced sampling techniques (e.g., Hamiltonian Replica Exchange) during equilibration to ensure water molecules properly sample deep pockets and the solute surface [3].
Free Energy Calculation:
- Perform the solvation free energy calculation using an alchemical method (e.g., Thermodynamic Integration or Free Energy Perturbation) for each box size.
- Ensure each calculation uses extensive sampling. For protein solvation, investing over 1 μs of aggregate sampling per box size may be necessary for convergence [3].
- Crucially, conduct multiple (e.g., 10-20) independent repeats for each box size, starting from different initial velocities.
Data Analysis:
- Plot the computed free energy for each box size with error bars representing the standard error or confidence intervals across the repeats.
- A statistically sound result will show overlapping confidence intervals across all box sizes, indicating no dependence.
Protocol 2: Setting Up a Minimally Sufficient Simulation Box
This protocol describes the standard procedure for setting up a simulation box that is large enough to avoid artifacts but small enough to be computationally efficient.
Initial Solvation:
- Start with your solute (protein, ligand, etc.) centered in a simulation box.
- Use your MD preprocessing software (e.g.,
gmx solvatein GROMACS) to add solvent. Initially, choose a box type (e.g., cubic, dodecahedron) and a van der Waals buffer distance.
Box Size Check:
- Measure the minimum distance between any atom of the solute and the box edge. The recommended minimum is 1.0 nm for most biomolecular systems [3].
- Ensure this distance is maintained in all three dimensions. A dodecahedral box can often achieve this with fewer water molecules than a cubic box.
Ion Addition and Neutralization:
- Add ions to neutralize the system's net charge and then to achieve the desired physiological or experimental salt concentration.
Verlet Buffer Validation:
- In your MD parameter file (e.g.,
.mdpfile in GROMACS), set the Verlet list buffer to allow for automatic optimization (verlet-buffer-tolerance). The software will then determine the optimal pair-list update frequency and buffer size to control energy drift [46]. - After a short test simulation, check the log file for warnings about excessive energy drift, which would indicate a need for a larger box or a more conservative buffer setting.
- In your MD parameter file (e.g.,
Workflow and System Setup Diagrams
Box Size Optimization Workflow
This diagram outlines the iterative process for empirically determining the optimal simulation box size for a given system and property.
Minimal Box Setup Protocol
This diagram shows the standard protocol for setting up a sufficiently large simulation box to avoid finite-size artifacts.
The Scientist's Toolkit
Table 3: Essential Software and Methodological "Reagents"
| Item Name | Function / Role in Research | Key Application Notes |
|---|---|---|
| GROMACS [46] [3] [57] | A high-performance, open-source MD simulation package. The primary engine for running simulations. | Known for its speed and efficiency. Its Verlet cutoff scheme and automatic buffer tuning are critical for stable simulations in boxes of various sizes [46]. |
| CHAPERONg [57] | An automation tool for GROMACS-based simulation pipelines and trajectory analysis. | Simplifies and standardizes the setup of multiple systems (e.g., with different box sizes), reducing human error and ensuring consistency in repetitive tasks [57]. |
| StreaMD [58] | A Python-based tool for high-throughput MD simulations across multiple servers. | Enables the automated setup and execution of large-scale parameter studies, such as testing many box sizes or running numerous independent repeats for statistics [58]. |
| Particle-Mesh Ewald (PME) [3] [59] | The standard algorithm for handling long-range electrostatic interactions in periodic systems. | Correct implementation is essential. Artifacts can arise in vacuum or with poor screening if boxes are too small, as PME interactions between images become significant [3]. |
| Alchemical Free Energy Methods [3] | A class of computational techniques for calculating free energy differences by perturbing the system's Hamiltonian. | Used for calculating solvation free energies. Requires extensive sampling and statistical checks to avoid spurious box-size effects due to poor convergence [3]. |
| Hamiltonian Replica Exchange [3] | An enhanced sampling method that improves conformational sampling by exchanging states between replicas with different Hamiltonians. | Particularly useful for equilibrating solvent around complex solutes like proteins and for improving convergence in free energy calculations, thereby revealing true box-size effects [3]. |
Validation Frameworks and Comparative Analysis of Box Size Effects
Statistical Significance Testing for Box Size Independence
Frequently Asked Questions (FAQs)
What is the fundamental principle behind testing for box size independence?
The core principle is that for a properly configured molecular dynamics (MD) simulation, thermodynamic and kinetic properties should not vary systematically with the simulation box size, provided the box is sufficiently large. Observing such a dependence is often a statistical artifact resulting from insufficient sampling rather than a true physical effect. The goal of statistical testing is to demonstrate that any observed variations fall within the range expected from random statistical fluctuations [3].
My initial results show a clear trend with box size. Does this confirm a box size effect?
Not necessarily. Apparent trends can be misleading when based on limited data. As demonstrated in solvation free energy studies, selecting individual trajectory replicates can produce arbitrary upward, downward, or non-monotonic trends. These apparent effects typically disappear when a sufficient number of independent repeats are analyzed (e.g., N=20), and confidence intervals are applied. Always perform multiple replicates before concluding that a box size effect is genuine [3].
What are the minimum recommended distances between the solute and the box boundary?
Best practices dictate a minimum distance between the solute surface and the simulation box edge of at least 1.0 nm to ensure thermodynamic properties remain independent of box size. A distance of approximately 0.5 nm is considered extremely small and introduces artifacts, as the solvation shells of periodic images begin to interact, and water screening becomes insufficient [3].
How does the treatment of electrostatic interactions influence box size dependence?
The use of Particle-Mesh Ewald (PME) for long-range electrostatics is standard. In a fully solvated system, water molecules provide sufficient screening to prevent box size dependence. However, in vacuum simulations (e.g., for alchemical free energy steps), a strong box size dependence emerges because the periodic environment is not screened. Therefore, caution is required when interpreting results from non-solvated systems with PME [3].
Troubleshooting Guides
Problem: Observed thermodynamic property drifts with decreasing box size.
| Potential Cause | Diagnostic Steps | Recommended Solution |
|---|---|---|
| Insufficient sampling | 1. Run multiple independent replicates (e.g., 20 per box size).2. Calculate confidence intervals/error bars for the property at each box size.3. Check if the trend persists across all replicates. | Increase the sampling. If the trend disappears with more replicates, it was a statistical fluctuation [3]. |
| Box size is too small | Measure the shortest distance between any solute atom and the box edge. | Ensure the solute-box edge distance is ≥ 1.0 nm. If it is near or below 0.5 nm, increase the box size [3]. |
| Inadequate equilibration | Check the time series of properties like potential energy and density for drift at the start of the production run. | Extend the equilibration procedure. For complex solutes like proteins, consider using enhanced sampling methods (e.g., Hamiltonian replica exchange) to improve solvent equilibration around the solute [3]. |
Problem: High variance in results across different box sizes.
| Potential Cause | Diagnostic Steps | Recommended Solution |
|---|---|---|
| Poor phase space overlap (in alchemical calculations) | Analyze the energy distributions between neighboring alchemical windows. | Increase the amount of sampling per window or the number of windows. Employ Hamiltonian replica exchange to improve sampling efficiency [3]. |
| Single, short trajectory per box size | The property of interest (e.g., solvation free energy) shows a wide spread from a single measurement. | Do not rely on single realizations. Perform multiple independent simulation repeats for each box size condition to robustly estimate the mean and variance [3]. |
The following table synthesizes key quantitative findings from a foundational study that investigated box size effects across various systems [3].
Table 1: Summary of Key Experimental Findings on Box Size Effects
| System / Property Investigated | Box Size Range (Number of Waters) | Key Statistical Finding | Conclusion on Box Size Effect |
|---|---|---|---|
| Hydration Free Energy (Anthracene) | 473 to 5334 | With N=20 repeats per box size, no statistically significant trend was observed. Single replicates showed arbitrary, misleading trends. | No effect when sufficient sampling is used [3]. |
| Electrostatic Solvation Free Energy (GB Protein in Water) | Various, with >1.0 nm protein-box edge distance | ΔGQ was independent of box size for sufficiently large, solvated boxes. | No effect in solvated systems with proper screening [3]. |
| Electrostatic Solvation Free Energy (GB Protein in Vacuum) | Various box sizes | A strong box size dependence was observed due to the lack of screening in a periodic vacuum. | Strong artificial effect present; requires correction [3]. |
| GB Protein in Reduced-Screening Solvent | Various box sizes | With water charges scaled to 10%, a box size dependence emerged, mimicking the vacuum-like behavior. | Effect appears only when electrostatic screening is insufficient [3]. |
Experimental Protocol: Testing for Box Size Effects in Solvation Free Energy
This protocol outlines a robust methodology for statistically testing the influence of box size on the calculation of solvation free energies for a small molecule, following the approach used in the referenced research [3].
Objective: To determine whether the computed solvation free energy of a small molecule is independent of simulation box size through rigorous statistical testing.
System Setup:
- Molecule Selection: Select a small molecule ligand (e.g., anthracene).
- Box Size Selection: Create multiple simulation systems where the molecule is solvated in boxes of different sizes. The range should include a minimally acceptable box (e.g., just exceeding 2x the non-bonded cut-off) to substantially larger boxes.
- Solvation and Ions: Solvate the molecule in water (e.g., TIP3P model) and add ions to neutralize the system.
- Force Field: Choose an appropriate force field (e.g., CHARMM36m) [60].
Simulation Parameters:
- Software: Use an MD package like GROMACS [3] [60].
- Electrostatics: Employ Particle-Mesh Ewald (PME) for long-range interactions with a short-range cutoff of 1.2 nm [3] [60].
- Temperature & Pressure: Use a velocity-rescaling thermostat and a Parrinello-Rahman barostat to maintain constant temperature (e.g., 300 K) and pressure (1 bar) [60].
- Constraints: Apply the LINCS algorithm to constrain bond lengths [60].
Sampling and Replicates:
- Replicates: For each box size, perform a minimum of 20 independent repeats [3].
- Initialization: Each repeat must start from different initial random velocities (e.g., from a Maxwell-Boltzmann distribution).
- Enhanced Sampling: For the free energy calculation itself, use an alchemical method with Hamiltonian replica exchange (HREX) between neighboring lambda windows to ensure adequate sampling and phase space overlap [3].
Workflow Diagram: The following diagram illustrates the key stages of this experimental protocol.
Analysis and Statistical Testing:
- Data Collection: For each box size, you will have N estimates of the solvation free energy (ΔG).
- Descriptive Statistics: Calculate the mean and standard deviation for the ΔG values at each box size.
- Visualization: Create a scatter plot of ΔG versus box size, overlaying the mean value and error bars (e.g., 95% confidence intervals) for each box.
- Statistical Testing: Perform a one-way Analysis of Variance (ANOVA) to test if the means of the ΔG values across different box sizes are statistically significantly different. A non-significant p-value (e.g., > 0.05) supports the null hypothesis that box size has no effect.
- Conclusion: The box size is considered to have no significant effect if no systematic trend is visible in the plot and the ANOVA test is not significant.
The Scientist's Toolkit
Table 2: Essential Research Reagents and Computational Tools
| Item / Resource | Function / Purpose in Research |
|---|---|
| GROMACS | A versatile software package for performing molecular dynamics simulations; it includes tools for system setup, simulation execution, and trajectory analysis [60]. |
| CHARMM36m Force Field | A widely used set of parameters for biomolecular simulations, providing the energy functions needed to compute interatomic forces [60]. |
| Alchemical Free Energy Methods | A class of computational techniques used to calculate free energy differences (e.g., solvation free energy) by gradually turning interactions on or off along a non-physical "alchemical" pathway [3]. |
| Hamiltonian Replica Exchange (HREX) | An enhanced sampling method that improves conformational exploration by allowing exchanges between simulations at different stages of the alchemical pathway, crucial for converging free energy estimates [3]. |
| Particle-Mesh Ewald (PME) | The standard algorithm for accurately treating long-range electrostatic interactions in periodic systems, preventing artifacts from self-interactions of periodic images [3] [60]. |
| Statistical Analysis Scripts (Python/R) | Custom scripts for calculating descriptive statistics, performing ANOVA tests, and generating plots with confidence intervals to rigorously assess the significance of results [3]. |
Frequently Asked Questions
What is the minimum acceptable distance between my protein and the edge of the simulation box? Research on lysozyme oligomers suggests that a minimum distance (offset) of 1 nm is acceptable for studying protein crystallization solutions without significant loss of accuracy. Studies found that lysozyme dimers remained stable while hexamers dissociated as expected experimentally, even at this smallest box size [4] [61].
Will increasing my simulation box size always improve the accuracy of my results? Not necessarily. For protein oligomer stability studies, increasing box size beyond a certain point may not refine results and could even slightly worsen their quality. The key is ensuring the box is not so small that it causes artifacts, but excessively large boxes consume more computational resources without providing additional benefits for assessing oligomer stability [61].
How can an improperly sized simulation box affect my protein stability results? An overly small box can artificially influence protein behavior through periodic boundary conditions, where virtual copies of your protein in neighboring boxes exert unrealistic influence on each other. This may lead to exaggerated structural fluctuations and potentially incorrect conclusions about oligomer stability [4] [62].
What evidence supports the use of smaller box sizes for oligomer stability studies? Smaller boxes may better replicate real crystallization conditions where proteins are surrounded by other molecules rather than pure solvent. The "virtual copies" in neighboring boxes through periodic boundary conditions can mimic the crowded environment proteins experience in actual solution, potentially providing more biologically relevant stability data [61].
Troubleshooting Guide
Problem: Unstable oligomers in small simulation boxes
Issue: Your protein oligomers show unexpected instability or dissociation in smaller simulation boxes.
Diagnosis Steps:
- Calculate RMSD, RMSF, and radius of gyration (Rg) values and compare across different box sizes
- Check if the instability is consistent with experimental evidence (e.g., should the oligomer be stable?)
- Verify if the minimum distance between protein atoms and box edge is at least 1 nm
Solutions:
- For lysozyme-like systems, ensure at least 1 nm offset between protein and box edge [4]
- Run multiple replicates to distinguish genuine instability from random variation [62]
- For amyloid-forming peptides like GNNQQNY, consider that some oligomer instability may be biologically real rather than artifact [63]
Problem: Inconsistent results between different box sizes
Issue: Your oligomers show different stability patterns when simulated in different box sizes.
Diagnosis Steps:
- Compare RMSF values - significant differences may indicate box size artifacts
- Check for systematic changes in Rg values across box sizes
- Verify your periodic boundary condition handling is correct across all simulations
Solutions:
- Use the
gmx trjconvcommand with appropriate flags (-pbc nojump,-pbc mol -center) to properly remove periodicity before analysis [4] [62] - Ensure noncovalent interaction cutoffs are appropriately set (typically 1 nm radius) [4]
- Consider that some variation (e.g., RMSD differences of 0.2-0.3 nm) may be normal between replicates [62]
Table 1: Lysozyme Oligomer Stability Metrics Across Different Box Sizes [4]
| Minimum Protein-Box Offset (nm) | Dimer Maximum RMSF (nm) | Hexamer Maximum RMSF (nm) | Hexamer RMSD Range (nm) | Oligomer Behavior Consistent with Experimental Data? |
|---|---|---|---|---|
| 1.0 | ≤0.7 | ≥1.4 | Up to 1.2 | Yes |
| 1.5 | ≤0.7 | ≥1.4 | Up to 1.2 | Yes |
| 2.0 | ≤0.7 | ≥1.4 | Up to 1.2 | Yes |
| 2.5 | ≤0.7 | ≥1.4 | Up to 1.2 | Yes |
| 3.0 | ≤0.7 | ≥1.4 | Up to 1.2 | Yes |
Table 2: Lysozyme Radius of Gyration Variations Across Box Sizes [61]
| Minimum Protein-Box Offset (nm) | Dimer Rg Variation | Hexamer Rg Variation | Notes |
|---|---|---|---|
| 1.0 | <0.5% of absolute Rg | Significant deviation | Most noticeable structural changes |
| 1.5 | <0.5% of absolute Rg | Significant deviation | Noticeable structural changes |
| 2.0 | <0.5% of absolute Rg | Minimal deviation | Closest to original crystal structure |
| 2.5 | <0.5% of absolute Rg | Minimal deviation | Closest to original crystal structure |
| 3.0 | <0.5% of absolute Rg | Minimal deviation | Closest to original crystal structure |
Detailed Experimental Protocols
System Setup:
- Oligomer Sources: Dimers and hexamers isolated from tetragonal crystal structure of lysozyme (PDB ID: 6QWY) using PyMOL
- Simulation Box: Cubic boxes with periodic boundary conditions
- Box Offsets Tested: 1.0, 1.5, 2.0, 2.5, and 3.0 nm minimum distance between protein atoms and box faces
- Solvation: TIP4P-Ew water model
- Ions: 0.4 M NaCl concentration to match crystallization conditions, with additional ions to neutralize system charge
Simulation Parameters:
- Force Fields: Amber ff99SB-ILDN or Amber ff19SB
- Software: GROMACS 2021 or AMBER 2019
- Temperature: 10°C (283.15 K) for crystallization conditions
- Electrostatics: PME algorithm with 1 nm cutoff for noncovalent interactions
- Integration: Leap-frog algorithm with 2 fs time step
- Simulation Duration: 1 μs for dimers, 700 ns for hexamers
Analysis Methods:
- Root-mean-square fluctuation (RMSF) for Cα atoms
- Root-mean-square deviation (RMSD) from initial structure
- Radius of gyration (Rg) calculations
- Visual inspection of dissociation events in trajectories
System Setup:
- Peptide Sequence: GNNQQNY (from yeast prion protein Sup35)
- Simulation Box: Rectangular or cubic boxes with minimum 10 Å (1 nm) distance to edges
- Solvation: SPC water model
- Initial Structures: Random orientations and conformations from monomer MD trajectories
Simulation Parameters:
- Force Field: GROMACS96
- Software: GROMACS 3.3
- Temperature: 300 K (except monomer at 450 K for conformational sampling)
- Electrostatics: Particle mesh Ewald with 10 Å cutoff
- Simulation Duration: 50-80 ns depending on oligomer size
Analysis Methods:
- Secondary structure analysis using DSSP
- Backbone-backbone hydrogen bond counting
- Parallel vs. antiparallel β-sheet alignment assessment
- π-π stacking interactions of aromatic residues
Workflow Visualization
Box Size Optimization Workflow
Research Reagent Solutions
Table 3: Essential Materials and Computational Tools for Oligomer Stability Studies
| Resource | Function/Purpose | Example Specifications |
|---|---|---|
| MD Simulation Software | Engine for running molecular dynamics simulations | GROMACS 2021, AMBER 2019 [4] [61] |
| Force Fields | Mathematical representation of interatomic forces | Amber ff99SB-ILDN, Amber ff19SB, CHARMM36 [4] [61] [62] |
| Water Models | Solvation environment for proteins | TIP4P-Ew, SPC, TIP3P [4] [63] [64] |
| Structure Visualization | Analysis and rendering of protein structures and trajectories | PyMOL, VMD [4] [62] |
| Analysis Tools | Calculation of stability metrics (RMSD, RMSF, Rg) | GROMACS built-in tools, cpptraj (Amber) [4] [61] |
| Protein Structures | Starting coordinates for oligomers | PDB IDs: 6QWY (lysozyme), 1YJP (GNNQQNY) [4] [63] |
Troubleshooting Guides
Issue 1: Unexpectedly High RMSD Values After Simulation
| Symptom | Potential Cause | Diagnostic Steps | Solution |
|---|---|---|---|
| Steady, high RMSD throughout trajectory. [4] | System not properly equilibrated. | Check if RMSD plot reaches a stable plateau. Compare with RMSD from a different, longer equilibration protocol. [6] | Extend equilibration time. Try a more robust equilibration method (e.g., the proposed ultrafast method shown to be 200% more efficient than conventional annealing). [6] |
| High RMSD in a small simulation box. [4] | Artifacts from periodic boundary conditions; the box is too small. | Measure the minimum distance (offset) between the protein and the box face. | Increase the simulation box size. A minimum offset of 1 nm is recommended to prevent artifacts from protein copies in virtual boxes. [4] |
| High RMSD in one specific region. | Conformational change in a flexible loop or domain. | Calculate per-residue RMSF to identify the flexible regions causing the high global RMSD. [65] | Analyze if the change is biologically relevant. Consider using position restraints on stable regions if the focus is elsewhere. |
Issue 2: Interpreting RMSF and B-Factor Relationships
| Symptom | Potential Cause | Diagnostic Steps | Solution |
|---|---|---|---|
| Discrepancy between simulation RMSF and experimental B-factors. [65] | B-factors contain contributions from static disorder, crystal contacts, and refinement errors, not just dynamics. [65] | Ensure your simulation ensemble is representative. Compare the ensemble-average pairwise RMSD with the value derived from B-factors. | For a single molecular species, the root mean-square ensemble-average pairwise RMSD, |
| RMSF values are uniformly low/high. | Incorrect reference structure for alignment when calculating RMSF. [65] | Recalculate RMSF using every structure in the ensemble for alignment, not just one reference. [65] | Use a robust alignment method. The choice of reference can significantly affect RMSF in a heterogeneous ensemble. [65] |
Issue 3: Radius of Gyration (Rg) Values Indicating Overly Compact or Expanded Structures
| Symptom | Potential Cause | Diagnostic Steps | Solution |
|---|---|---|---|
| Rg is much smaller than expected, suggesting an unnaturally compact protein. [66] | Potential force field inaccuracies or lack of sufficient simulation time for proper solvation. | Check the Rg trend over time; ensure it has stabilized. Compare with experimental data (e.g., from SAXS) if available. | Verify your system setup, including water model and ion concentration. Consider if the force field is appropriate for your specific molecule. [67] |
| Rg is unstable and fluctuates wildly in a multi-chain system (e.g., a hexamer). [4] | The oligomer may be inherently unstable under the simulation conditions, or the box size is too small. | Compare the stability of different oligomers (e.g., dimer vs. hexamer). A hexamer showed an Rg variation of 0.8 nm in a small box, while a dimer was stable (~0.2 nm variation). [4] | This may be a correct result if the oligomer is not stable in solution. Validate with known experimental oligomerization states. Ensure the box size has a sufficient offset (e.g., ≥1 nm). [4] |
Frequently Asked Questions (FAQs)
Q1: What is the core mathematical relationship between RMSD, RMSF, and Rg?
These metrics, while distinct, are fundamentally connected through their definitions of spatial deviations.
- RMSD (Root Mean Square Deviation): Measures the global structural difference between two structures after optimal alignment. The all-against-all pairwise RMSD distribution characterizes the structural heterogeneity of an ensemble. [65]
- RMSF (Root Mean Square Fluctuation): Measures the local deviation of a particular atom (e.g., Cα) around its average position over time. [65] It is directly related to experimental X-ray B-factors by the formula: RMSFᵢ² = 3Bᵢ / 8π². [65]
- Rg (Radius of Gyration): Measures the overall compactness of a structure, defined as the mass-weighted average distance of atoms from the system's center of mass. [66]
The key relationship is analogous to the two definitions of the radius of gyration for a polymer. The root mean-square ensemble-average pairwise RMSD (
Q2: How does simulation box size directly impact these validation metrics?
The simulation box size is a critical factor for obtaining accurate and artifact-free results, especially when studying oligomers or large conformational changes. [4]
The table below summarizes findings from a study on lysozyme dimer and hexamer stability in different box sizes, defined by the "offset" (minimum distance between protein atoms and the box face). [4]
| Box Offset (nm) | Dimer Stability | Hexamer Stability | Key Observations on Metrics |
|---|---|---|---|
| 1.0 | Stable | Unstable | A 1 nm offset was found to be the minimum permissible size; the dimer was stable (Rg variation ~0.2 nm) while the hexamer was not, matching experimental data. [4] |
| 1.5 | Stable | Unstable | RMSF values for the hexamer were significantly higher (≥1.2 nm spread) than for the dimer (≤0.6 nm spread), indicating greater instability. [4] |
| 2.0 | Stable | Unstable | The hexamer showed the smallest spread of RMSF values (1.2 nm), but this was still twice that of the least stable dimer. [4] |
| 2.5 | Stable | Unstable | The dimer's RMSD stabilized at ~0.7 nm after 570 ns, while the hexamer's RMSD continued to grow, failing to stabilize within 700 ns. [4] |
Using a box that is too small can lead to artificial interactions between the protein and its periodic images, which can destabilize native oligomers that should be stable or mask the instability of non-native oligomers. [4]
Q3: What are the detailed protocols for calculating these metrics in GROMACS?
After running your simulation, use these GROMACS commands to calculate the key metrics. The following protocols assume you have a trajectory file (traj.xtc), a topology file (topol.tpr), and an index file (index.ndx).
When prompted by these commands, select the appropriate atom groups (e.g., "Backbone" for RMSD alignment, "C-alpha" for RMSF and Rg). The -tu ns flag in the RMSD command plots the X-axis in nanoseconds. [4]
Q4: My ligand's properties are inaccurate. How can I improve the force field parameters?
Standard automatic parameterization tools (like GAFF or CGenFF) use pre-tabulated parameters, which may be inadequate for specific molecules, particularly for van der Waals (LJ) interactions. [67] To improve accuracy:
- Use Advanced Parameterization Tools: Employ protocols like GAAMP (General Automated Atomic Model Parameterization) that refine bond, angle, dihedral, and atomic charge parameters using ab initio quantum mechanical (QM) data as a target, ensuring context-specific accuracy. [67]
- Optimize Lennard-Jones Parameters: For drug-like small molecules, consider using optimized LJ parameter sets that have been systematically refined to reproduce experimental neat liquid densities, enthalpies of vaporization, and hydration free energies for a broad set of compounds. [67] One such optimization for GAFF resulted in an average unsigned error in hydration free energies of 0.79 kcal/mol. [67]
- Validate with Key Experiments: Always validate your final parameters against available experimental data, such as hydration free energies or NMR data, to ensure reliability before running production simulations. [67]
Experimental Workflow and Relationships
The following diagram illustrates the logical workflow for calculating and interpreting the primary validation metrics in a molecular dynamics study, highlighting their interconnectedness.
Research Reagent Solutions
This table lists essential computational tools, force fields, and parameters critical for running and validating molecular dynamics simulations.
| Item Name | Function / Purpose | Key Features / Notes |
|---|---|---|
| GROMACS [4] [68] | A versatile software package for performing MD simulations and analysis. | High performance. Includes commands for rms, rmsf, and gyrate. Used with force fields like Amber and CHARMM. [4] |
| Amber ff99SB-ILDN [4] | A high-quality force field for proteins. | Provides parameters for atoms in biomolecules. Often used for simulating proteins and nucleic acids. [4] |
| GAFF & CGenFF [67] | General force fields for small molecules and drug-like compounds. | Provide parameters for a wide range of chemical functionalities. A starting point for ligand parameterization. [67] |
| GAAMP [67] | Tool for automated atomic model parameterization. | Refines GAFF/CGenFF parameters using QM data for bonds, angles, dihedrals, and charges, improving accuracy. [67] |
| Optimized LJ Parameters [67] | A refined set of van der Waals parameters for small molecules. | Optimized to reproduce experimental liquid densities and enthalpies of vaporization. Can improve hydration free energy calculations. [67] |
| TIP4P-Ew [4] | A four-site water model. | Used to solvate the simulation box. Provides an accurate model for water interactions in MD simulations. [4] |
Solvation Free Energy Calculations as Validation Tools
FAQs: Addressing Common Computational Challenges
FAQ 1: Does the size of my simulation box significantly affect the calculated solvation free energy? For most typical system setups, the simulation box size has a minimal impact on the calculated solvation free energy, provided the box is sufficiently large. A study investigating the hydration free energy of anthracene in boxes containing 473 to 5,334 water molecules found no statistically significant trend in the results across box sizes when sufficient sampling was performed. Apparent box-size dependencies often vanish with increased sampling and proper statistical analysis, highlighting the importance of reporting uncertainty estimates for calculated observables [3].
FAQ 2: What is the minimum acceptable distance to maintain between my solute and the box edge? A minimum distance (offset) of 1 nanometer between the solute atoms and the box face is generally acceptable for protein systems in crystallization solutions. Research on lysozyme oligomers demonstrated that a 1 nm offset was the smallest size that still produced correct stability results (i.e., stable dimers and unstable hexamers). Using a smaller offset, such as 0.5 nm, can lead to artifacts where periodic images interact unrealistically [4] [3]. For smaller molecules, ensuring the box size exceeds twice the non-bonded cut-off radius in all dimensions is a foundational rule [3].
FAQ 3: My solvation free energy calculation is not converging. What could be wrong? Poor convergence is often a sampling issue, not necessarily a box size problem. Alchemical free energy calculations, especially for large molecules like proteins, require extensive sampling to ensure phase space overlap between neighboring intermediate states (lambda windows). For example, achieving a converged electrostatic solvation free energy for a 56-residue protein (GB) required 1.38 microseconds of sampling per box size, enhanced by Hamiltonian replica exchange. Insufficient sampling can lead to non-convergent and unreliable results [3].
FAQ 4: How do I handle long-range electrostatic interactions correctly in a periodic box? The Particle-Mesh Ewald (PME) method is the standard for handling long-range electrostatics in systems with periodic boundaries. In a properly sized, solvated system, water molecules effectively screen the electrostatic interactions between a solute and its periodic images. However, if you were to perform a simulation in vacuum using PME, a strong box-size dependence would appear because there is no solvent screening. Therefore, calculations in explicit solvent are recommended to avoid such artifacts [3].
Troubleshooting Guides
Issue: Erroneous Box-Size Dependence in Results
Problem: Observed trends in solvation free energy seem to depend on the simulation box size.
- Solution A: Increase Sampling and Repeats
- Action: Do not rely on single simulation trajectories. Perform multiple independent repeats (e.g., N=20) for each box size condition.
- Rationale: Apparent trends often disappear with sufficient sampling and statistical analysis. Anecdotal evidence from single runs can be misleading [3].
- Solution B: Verify Box Size Setup
- Solution C: Check Solvent Screening
- Action: Confirm your system is properly solvated. Avoid simulations in vacuum or with poorly screening solvents when using periodic boundary conditions.
- Rationale: Electrostatic box-size artifacts are pronounced without proper dielectric screening [3].
Issue: Instability of Protein Oligomers in Simulation
Problem: Protein oligomers (e.g., dimers, hexamers) that are expected to be stable in experiment dissociate during simulation.
- Solution: Validate Box Size and Force Field
- Action 1: Check that the simulation box offset is not less than 1 nm [4].
- Action 2: Ensure the use of a modern, appropriate force field (e.g., Amber ff99SB-ILDN for proteins) and accurate water models (e.g., TIP4P-Ew) [4].
- Rationale: An undersized box can artificially destabilize complexes. Using validated force fields and simulation protocols is critical for obtaining physically meaningful results [4].
Issue: High Variance in Alchemical Free Energy Estimates
Problem: The calculated free energy differences have unacceptably large statistical errors.
- Solution A: Implement Soft-Core Potentials
- Action: Use a soft-core potential for Lennard-Jones interactions during alchemical transformations. This prevents energy singularities and numerical instabilities when atoms are partially decoupled and can overlap [69].
- Rationale: Soft-core potentials are designed to smooth the energy landscape along the alchemical path, which reduces the variance of the free energy estimate and improves convergence [69].
- Solution B: Enhance Sampling
- Action: Employ advanced sampling techniques like Hamiltonian Replica Exchange (HREM) between the lambda windows.
- Rationale: This technique improves phase space sampling and overlap between intermediate states, which is crucial for converging difficult transformations, such as solvating an entire protein [3].
Quantitative Data on System Sizes and Results
Table 1: Summary of Simulation Box Sizes and Key Findings from Cited Studies
| Study System | Box Sizes / Offsets Tested | Key Measured Property | Finding Related to Box Size | Citation |
|---|---|---|---|---|
| Anthracene Hydration | 473 to 5,334 water molecules | Hydration Free Energy (ΔG) | No statistically significant box-size effect observed with sufficient sampling (N=20 repeats). | [3] |
| Lysozyme Oligomers | 1.0, 1.5, 2.0, 2.5, 3.0 nm offset | Oligomer Stability (RMSF, RMSD, Rg) | A 1.0 nm offset was found to be the minimum acceptable size to correctly reproduce experimental stability. | [4] |
| Protein GB (Electrostatics) | ~1.0 nm offset and smaller | Solvation Free Energy Component (ΔGQ) | No box-size dependence in water with ≥1.0 nm offset. Strong dependence in vacuum or with poor screening. | [3] |
Table 2: Essential Research Reagents and Computational Tools
| Item Name | Function / Description | Example from Literature |
|---|---|---|
| GROMACS | A versatile software package for performing molecular dynamics simulations. | Used with the Amber ff99SB-ILDN force field for lysozyme oligomer stability studies [4]. |
| Amber ff99SB-ILDN | A well-validated force field for simulating proteins and nucleic acids. | Employed to model lysozyme dimers and hexamers in crystallization solutions [4]. |
| TIP4P-Ew | A four-site water model that provides a more accurate description of water properties. | Used to solvate lysozyme oligomers in their simulation boxes [4]. |
| Particle-Mesh Ewald (PME) | An algorithm for efficiently calculating long-range electrostatic interactions in periodic systems. | Standard method for handling electrostatics; crucial for avoiding artifacts [4] [3]. |
| Soft-Core Potential | A modified potential energy function that prevents singularities in alchemical free energy calculations. | Critical for achieving low-variance free energy estimates when atoms are decoupled [69]. |
| Alchemical Free Energy Methods | Non-physical pathways (e.g., thermodynamic integration) to compute free energy differences. | Used to compute solvation free energies with high precision [70] [69] [3]. |
| Machine Learned Potentials (MLPs) | A promising alternative to empirical forcefields, offering higher accuracy from first principles. | Can achieve sub-chemical accuracy for solvation free energies of organic molecules [69] [71]. |
Experimental Protocols & Workflows
Protocol: Determining Minimum Box Size for Protein Solutions
This protocol is adapted from research on lysozyme crystallization solutions [4].
- System Preparation:
- Obtain the initial protein structure(s) from a database (e.g., PDB ID: 6QWY for lysozyme octamer).
- Use a molecular visualization program (e.g., PyMOL) to isolate the oligomers of interest (e.g., dimer, hexamer).
- Set protonation states of amino acid residues to reflect the experimental pH using a tool like the PROPKA server.
- Box Construction and Solvation:
- Place the oligomer at the center of a cubic simulation box.
- Create multiple boxes where the minimum distance between the protein atoms and the box face (the offset) is systematically varied (e.g., 1.0, 1.5, 2.0, 2.5, 3.0 nm).
- Fill each box with water molecules (e.g., using the TIP4P-Ew model).
- Add ions to match the experimental precipitant concentration (e.g., 0.4 M NaCl) and neutralize the system's total charge.
- Simulation and Analysis:
- Run molecular dynamics simulations for all systems using the same parameters (e.g., GROMACS, Amber ff99SB-ILDN force field, 2 fs time step, PME for electrostatics).
- Simulate for a sufficiently long duration (e.g., 700 ns to 1 μs) to observe stability or dissociation.
- Analyze trajectories using metrics like Root-Mean-Square Fluctuation (RMSF), Root-Mean-Square Deviation (RMSD), and Radius of Gyration (Rg).
- Validation Criterion: The smallest box size that reproduces the experimentally known oligomer stability (e.g., stable dimers, unstable hexamers) is the minimum acceptable box size.
Protocol: Alchemical Hydration Free Energy Calculation
This protocol summarizes the well-established method for calculating solvation free energies [70] [3].
- Define End States:
- State A (Solvated): The solute molecule fully interacting with the solvent (e.g., water) in a periodic box.
- State B (Gas Phase): The solute molecule in vacuum, or effectively non-interacting with its surroundings.
- Create an Alchemical Pathway:
- Construct a series of non-physical intermediate states by mixing the Hamiltonians of State A and State B using a coupling parameter, λ (ranging from 0 to 1).
- A common approach is to use a two-step process: first turning off the van der Waals interactions (λv), and then turning off the electrostatic interactions (λe).
- Run Simulations with Soft-Core Potentials:
- Perform simulations at each discrete λ value.
- Use a soft-core potential for Lennard-Jones interactions to avoid singularities as atoms are decoupled.
- Compute Free Energy:
- Use an estimator like Thermodynamic Integration (TI) or Exponential Averaging (Free Energy Perturbation, FEP) to compute the free energy change along the λ pathway.
- The sum of the free energy changes for all steps gives the total solvation free energy.
Best Practices for Reporting and Uncertainty Estimation
Frequently Asked Questions (FAQs)
Q1: Why is simulation box size so important in Molecular Dynamics studies? The simulation box size directly creates a balance between computational efficiency and result accuracy. A box that is too small can lead to artifacts from periodic boundary conditions, where a molecule's periodic images interact unrealistically. Conversely, an excessively large box wastes computational resources. The goal is to find the smallest box size that does not compromise the physical validity of your results [4] [3].
Q2: How does insufficient sampling affect uncertainty related to box size? Insufficient sampling can create the illusion of box-size-dependent effects that disappear with more data. Conclusions drawn from single or a few simulation repeats are highly vulnerable to this statistical noise. To reliably detect a true box size effect, you must perform multiple independent simulations and report uncertainty estimates, such as confidence intervals [3].
Q3: What is the minimum recommended distance between the solute and the box edge? A minimum offset of 1 nanometer is often a safe starting point. One study on lysozyme oligomers found that a 1 nm minimum distance was the smallest permissible size that still correctly reproduced the known stability of protein dimers over hexamers in a crystallization solution. Smaller offsets, such as 0.5 nm, have been shown to introduce significant artifacts [4] [3].
Q4: What are the best practices for reporting uncertainty in MD simulations? Always report the results of multiple independent simulation repeats alongside their statistical uncertainty (e.g., standard deviation or confidence intervals). Visually present this uncertainty on graphs with error bars. Furthermore, transparently document the number of repeats, total simulation time, and the statistical methods used to calculate the uncertainty [3].
Troubleshooting Guides
Issue: Unexpected Box Size Dependence in Results
Symptoms: Your computed thermodynamic or kinetic properties (e.g., free energy, diffusion rates) show a clear trend when you change the simulation box size.
Diagnosis and Resolution Steps:
Check Your Sampling: This is the most common cause. A perceived box size effect often vanishes with increased sampling.
- Action: For each box size, run multiple independent simulations (e.g., 20 repeats) starting from different initial velocities.
- Verification: Calculate the mean and confidence intervals for your observable across all repeats. If the confidence intervals between different box sizes overlap significantly, the observed dependence is likely not statistically significant [3].
Verify the Offset is Sufficient: Ensure your solute is not too close to its periodic images.
- Action: Measure the minimum distance between any atom of your solute and the box face. Increase the box size if this distance is less than 1 nm.
- Verification: After increasing the offset, re-run your simulations. The artifact should be reduced, particularly for properties influenced by long-range interactions [4] [3].
Validate Your Electrostatics Treatment: Incorrect handling of long-range electrostatics in a small box can skew results.
- Action: Confirm you are using a method like Particle-Mesh Ewald (PME) for electrostatic calculations. Be aware that in non-solvated systems (vacuum), PME can itself cause box-size dependence, so an infinite non-periodic box correction may be needed for vacuum calculations [3].
Issue: Achieving Equilibration is Computationally Expensive
Symptoms: Reaching a stable, equilibrated system density or energy for a large simulation cell (e.g., containing many polymer chains) takes an impractically long time using conventional methods like annealing.
Diagnosis and Resolution Steps:
- Evaluate Your Equilibration Protocol: Traditional annealing cycles between temperature sets (e.g., 300 K to 1000 K) are robust but computationally expensive.
- Action: Investigate more efficient equilibration algorithms. Recent research on ion exchange polymers like Nafion has demonstrated a novel approach that was ~200% more efficient than conventional annealing and ~600% more efficient than a "lean" two-step (NVT/NPT) method [6].
- Verification: Monitor the convergence of key properties like density and potential energy. The new protocol should achieve a stable plateau in these values faster than your previous method.
Key Experimental Data
Table 1: Lysozyme Oligomer Stability in Different Box Sizes [4] This table summarizes a study that determined the minimum acceptable box size by comparing the stability of lysozyme dimers and hexamers. The criterion for correctness was that the dimer (known to be stable) must be more stable than the hexamer (known to be absent) in the simulation.
| Minimum Protein-Box Offset (nm) | Dimer Stability (RMSF in nm) | Hexamer Stability (RMSF in nm) | Box Size Correctness |
|---|---|---|---|
| 1.0 | ≤ 0.7 | ≥ 1.4 | Correct |
| 1.5 | ≤ 0.7 | ≥ 1.4 | Correct |
| 2.0 | ≤ 0.7 | ≥ 1.4 | Correct |
| 2.5 | ≤ 0.7 | ≥ 1.4 | Correct |
Table 2: Impact of Polymer Chain Count on Property Convergence [6] This data shows how the number of polymer chains in a simulation cell for a Nafion membrane affects the variability of calculated diffusion coefficients, highlighting a morphological threshold for reliable results.
| Number of Nafion Chains | Variation in Diffusivity | Error at High Hydration | Notes |
|---|---|---|---|
| 4 | High | Significant | Basic model, higher uncertainty |
| 8 | Moderate | Moderate | Improved convergence |
| 14 | Low | Significantly Reduced | Recommended threshold for low error |
| 16 | Low | Significantly Reduced | Recommended threshold for low error |
Experimental Protocol: Establishing Minimum Box Size
This protocol is adapted from a study investigating protein crystallization solutions to determine the smallest viable simulation box [4].
Objective: To establish the minimum permissible simulation box size for studying a given molecular system without loss of accuracy.
Methodology:
System Preparation:
- Construct initial coordinates for your molecule(s) of interest.
- Using simulation software like GROMACS, place the molecule in the center of a cubic simulation box with periodic boundary conditions.
- Generate a series of boxes where the minimum distance between the solute atoms and the box face (the "offset") varies (e.g., 1.0, 1.5, 2.0, 2.5 nm).
- Solvate the system with a water model (e.g., TIP4P-Ew) and add ions to neutralize the system and match experimental conditions.
Simulation Parameters:
- Software: GROMACS [4] [3].
- Force Field: Choose an appropriate force field (e.g., Amber ff99SB-ILDN for proteins) [4].
- Energy Minimization: Use the steepest descent method until the maximum force is below a threshold (e.g., 1000 kJ/(mol·nm)).
- Equilibration:
- NVT ensemble: 100 ps with a thermostat (e.g., V-rescale).
- NPT ensemble: 100 ps with a thermostat and a barostat (e.g., Parrinello-Rahman).
- Production MD: Run multiple long-timescale simulations (e.g., 700 ns to 1 μs) for each box size in the NPT ensemble.
Analysis:
- Calculate stability metrics such as Root-Mean-Square Fluctuation (RMSF) and Radius of Gyration (Rg).
- Compare the relative stability of different molecular configurations (e.g., dimers vs. hexamers) against known experimental data or theoretical expectations.
- The smallest box size that reproduces the correct, experimentally-verified behavior is the minimum acceptable box size for your system.
Workflow and Relationship Diagrams
Diagram 1: Box size optimization workflow. The process hinges on robust sampling and statistical analysis to distinguish real effects from noise [3].
Diagram 2: Uncertainty sources in MD. Uncertainty arises from both statistical shortcomings and methodological choices, both of which must be addressed [4] [3].
The Scientist's Toolkit
Table 3: Essential Research Reagents and Computational Tools
| Item Name/Software | Primary Function/Brief Explanation |
|---|---|
| GROMACS | A versatile software package for performing MD simulations; used for energy minimization, equilibration, and production runs [4] [72]. |
| AMBER Force Fields (e.g., ff99SB-ILDN) | A family of force fields providing parameters for calculating potential energy of biomolecules, crucial for simulating proteins and nucleic acids [4]. |
| Particle-Mesh Ewald (PME) | The standard algorithm for handling long-range electrostatic interactions in periodic systems, preventing artifacts [4] [3]. |
| Thermostat (e.g., V-rescale, Nose-Hoover) | A algorithm to regulate the temperature of the simulation system, maintaining a constant average temperature [4]. |
| Barostat (e.g., Parrinello-Rahman, Berendsen) | A algorithm to control the pressure of the simulation system by adjusting the box volume [73] [4]. |
| COMPASS, DREIDING Force Fields | Force fields commonly used for simulating materials and polymers, such as ion exchange membranes [73] [6]. |
| Kernel Density Estimate (KDE) & Information Entropy | A model-free method from information theory used to quantify uncertainty, diversity, and detect outliers in atomistic data [74]. |
Conclusion
Molecular dynamics simulation box size optimization represents a critical balance between computational efficiency and physical accuracy. Evidence indicates that with proper statistical validation, sufficiently large boxes (maintaining ≥1 nm protein-surface offset) yield reliable thermodynamic and kinetic properties for most biomolecular systems. Future directions should focus on developing adaptive box resizing algorithms, establishing system-specific size recommendations, and integrating machine learning approaches for automated optimization. For biomedical research, these advancements will enhance drug discovery pipelines by ensuring more accurate predictions of protein-ligand interactions, peptide folding behavior, and macromolecular dynamics while optimizing computational resource allocation. The field must continue to prioritize rigorous statistical validation to distinguish genuine box size effects from sampling artifacts.
References
- 1. Periodic Boundary Conditions - an overview [https://www.sciencedirect.com/topics/medicine-and-dentistry/periodic-boundary-conditions]
- 2. Periodic Boundary Conditions - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/periodic-boundary-conditions]
- 3. On the importance of statistics in molecular simulations for ... [https://elifesciences.org/articles/57589]
- 4. Minimum Acceptable Simulation Box Size Based on a ... [https://link.springer.com/article/10.1134/S2635167624602304]
- 5. Asymmetric Periodic Boundary Conditions for All-Atom ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10544013/]
- 6. Ultrafast molecular dynamics approach to quantify structural ... [https://pubs.rsc.org/en/content/articlehtml/2025/ra/d5ra05434f]
- 7. Periodic boundary conditions [https://manual.gromacs.org/2024.4/reference-manual/algorithms/periodic-boundary-conditions.html]
- 8. Simulation Box - an overview [https://www.sciencedirect.com/topics/engineering/simulation-box]
- 9. Lecture 15 : Periodic Box and Minimum Image Convention [https://archive.nptel.ac.in/content/storage2/courses/103103036/module3/lec15/1.html]
- 10. Who can explain the minimum image convention in ... [https://scicomp.stackexchange.com/questions/41171/who-can-explain-the-minimum-image-convention-in-molecular-dynamic-simulations]
- 11. dmc_lec2 - Materials Computation Center [https://www.mcc.uiuc.edu/summerschool/2001/David%20Ceperley/dmc_lec2.htm]
- 12. Periodic boundary conditions [https://en.wikipedia.org/wiki/Periodic_boundary_conditions]
- 13. CP2K Open Source Molecular Dynamics [https://www.cp2k.org/exercises:2021_uzh_acpc2:ex01]
- 14. Comment on 'Valid molecular dynamics simulations ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6586461/]
- 15. Getting good performance from mdrun [https://manual.gromacs.org/documentation/2025.2/user-guide/mdrun-performance.html]
- 16. Effect of periodic box size on aqueous molecular dynamics ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1184438/]
- 17. Which Optimizer Should You Use With NNPs? | Rowan [https://www.rowansci.com/blog/which-optimizer-should-you-use-with-nnps]
- 18. Protein rotating in triclinic box - GROMACS forums [https://gromacs.bioexcel.eu/t/protein-rotating-in-triclinic-box/96]
- 19. LAMMPS Warning - Triclinic Box Skew is Large [https://matsci.org/t/lammps-warning-triclinic-box-skew-is-large/50698]
- 20. Simulation box keeps shrinking in MD - GROMACS forums [https://gromacs.bioexcel.eu/t/simulation-box-keeps-shrinking-in-md/3511]
- 21. A beginner's guide to molecular dynamics simulations and ... [https://www.sciencedirect.com/science/article/abs/pii/S0076687920301488]
- 22. Optimal Molecular Dynamics System Size for Increased ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11075082/]
- 23. Identifying the minimal sets of distance restraints for FRET ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11118665/]
- 24. VMD - Visual Molecular Dynamics [https://www.ks.uiuc.edu/Research/vmd/]
- 25. LAMMPS Molecular Dynamics Simulator [https://www.lammps.org/]
- 26. Molecular simulation of protein-surface interactions [https://pmc.ncbi.nlm.nih.gov/articles/PMC2756768/]
- 27. Optimized protein-water interactions and torsional ... [https://www.nature.com/articles/s41467-025-65603-4]
- 28. Mapping the binding sites of challenging drug targets - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9790766/]
- 29. Comparison of software for molecular mechanics modeling [https://en.wikipedia.org/wiki/Comparison_of_software_for_molecular_mechanics_modeling]
- 30. How can I calculate the number of atoms and the size of the ... [https://gromacs.bioexcel.eu/t/how-can-i-calculate-the-number-of-atoms-and-the-size-of-the-simulation-box-for-the-md-simulation/9082]
- 31. Protocol for Molecular Dynamics Simulations of Proteins [https://en.bio-protocol.org/en/bpdetail?id=2051&type=0]
- 32. Possible sexual error: the box size is too small [https://gromacs.bioexcel.eu/t/possible-sexual-error-the-box-size-is-too-small/7159]
- 33. Valid molecular dynamics simulations of human hemoglobin require... [https://elifesciences.org/articles/35560]
- 34. re- Box After running the sizing ... - GROMACS forums md simulation [https://gromacs.bioexcel.eu/t/box-re-sizing-after-running-the-md-simulation/12167]
- 35. From adaptive resolution to molecular dynamics of open ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8547219/]
- 36. Common errors when using GROMACS [https://manual.gromacs.org/documentation/2025.4/user-guide/run-time-errors.html]
- 37. Accurate and efficient constrained molecular dynamics of ... [https://www.sciencedirect.com/science/article/pii/S0010465523000875]
- 38. Conformational Exploration of two Peptides and their Hybrid ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7512015/]
- 39. A guide to fixing broken AMBER MD trajectory files and ... [https://www.blopig.com/blog/2025/09/a-guide-to-fixing-broken-amber-md-trajectory-files-and-visualisations/]
- 40. How to adaptively change the box size or include only ... [https://matsci.org/t/how-to-adaptively-change-the-box-size-or-include-only-those-particles-which-agree-with-condition/57410]
- 41. Re: [AMBER] Setting the cut-off and size of box [http://archive.ambermd.org/201204/0049.html]
- 42. [AMBER] Change simulation box size from Anthony Cruz ... [http://archive.ambermd.org/201902/0058.html]
- 43. fix box/relax command [https://docs.lammps.org/fix_box_relax.html]
- 44. Common errors when using GROMACS [https://manual.gromacs.org/current/user-guide/run-time-errors.html]
- 45. 7.2. Errors and warnings details [https://docs.lammps.org/Errors_details.html]
- 46. Molecular Dynamics - GROMACS 2025.4 documentation [https://manual.gromacs.org/documentation/current/reference-manual/algorithms/molecular-dynamics.html]
- 47. Simulation box too small error [https://groups.google.com/g/hoomd-users/c/DgonBgnCG9c/m/aZXyOzl-BQAJ]
- 48. Automated MD Simulation Trajectory Analyses - Part I [https://abeebyekeen.com/post-sim-analysis-1/]
- 49. Cutoff Distance - an overview [https://www.sciencedirect.com/topics/computer-science/cutoff-distance]
- 50. Getting good performance from mdrun [https://manual.gromacs.org/current/user-guide/mdrun-performance.html]
- 51. Wrapping tutorial or “why is my simulation exploding?” [https://software.acellera.com/htmd/tutorials/WrappingTutorial.html]
- 52. Molecular Dynamics: Periodic Boundary Conditions (PBC) [https://www.compchems.com/molecular-dynamics-periodic-boundary-conditions-pbc/]
- 53. Periodic artifact reduction in Fourier transforms of full field ... [https://pubmed.ncbi.nlm.nih.gov/25597865/]
- 54. Reducing Edge Artifacts in 2D Fourier Tranforms with Python [https://www.roberthovden.com/tutorial/2015_fftartifacts.html]
- 55. Protein Crossing Membrane in Visualization Despite Centering [https://gromacs.bioexcel.eu/t/gromacs-pbc-artifact-protein-crossing-membrane-in-visualization-despite-centering/9872]
- 56. Frequently asked questions about VMD [http://www.ks.uiuc.edu/Research/vmd/allversions/vmd_faq.html]
- 57. CHAPERONg: A tool for automated GROMACS-based ... [https://www.sciencedirect.com/science/article/pii/S2001037023003367]
- 58. StreaMD: the toolkit for high-throughput molecular dynamics ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-024-00918-w]
- 59. Best Practices for Foundations in Molecular Simulations ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6884151/]
- 60. A Study of the Ara h 6 Peanut Protein [https://www.mdpi.com/2227-9717/13/2/581]
- 61. Influence of Simulation Box Sizes on the Stability ... [https://link.springer.com/article/10.1134/S2635167624600767]
- 62. Regarding box size - User discussions - GROMACS forums [https://gromacs.bioexcel.eu/t/regarding-box-size/1122]
- 63. Molecular Dynamics Simulations on the Oligomer ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1948036/]
- 64. Application of conventional molecular dynamics simulation ... [https://www.nature.com/articles/srep44651]
- 65. Determination of Ensemble-Average Pairwise Root Mean- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2830444/]
- 66. Lesson 10: Center of Mass and Radius of Gyration [https://charmm-gui.org/?doc=lecture&module=scientific&lesson=10]
- 67. Optimized Lennard-Jones Parameters for Drug-Like Small ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5997559/]
- 68. Molecular dynamics parameters (.mdp options) [https://manual.gromacs.org/2025.0-beta/user-guide/mdp-options.html]
- 69. Computing solvation free energies of small molecules with ... [https://arxiv.org/html/2405.18171v4]
- 70. Approaches for calculating solvation free energies and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5648357/]
- 71. Computing solvation free energies of small molecules with ... [https://arxiv.org/abs/2405.18171]
- 72. Molecular dynamics parameters (.mdp options) [https://manual.gromacs.org/2025-rc/user-guide/mdp-options.html]
- 73. Uncertainty analysis of the molecular dynamics method for ... [https://www.sciencedirect.com/science/article/abs/pii/S0360319924025163]
- 74. Model-free estimation of completeness, uncertainties, and ... [https://www.nature.com/articles/s41467-025-59232-0]