Research Articles
Assessing Phylogenetic Network Accuracy: A Comprehensive Guide for Introgression Characterization in Biomedical Research
This article provides a comprehensive examination of the accuracy and application of phylogenetic networks for characterizing introgression in evolutionary genomics.
Unraveling Genomic Introgression: A Hidden Markov Model Framework for Comparative Genomics
This article provides a comprehensive exploration of Hidden Markov Models (HMMs) and their transformative role in detecting and analyzing introgression in comparative genomic studies.
Navigating Gene Tree Estimation Error: A Practical Guide for Robust Introgression Detection in Genomic Studies
Accurate detection of introgression—the transfer of genetic material between species—is crucial for understanding evolutionary history, adaptation, and the genetic basis of traits with biomedical relevance.
Discrete Trait Analysis vs. Structured Birth-Death Models: A Comprehensive Guide for Biomedical Researchers
This article provides a comprehensive comparison of Discrete Trait Analysis (DTA) and Structured Birth-Death Models (SBDM), two foundational methods in phylogenetic inference for studying trait evolution and population dynamics.
Troubleshooting Molecular Clock Model Selection: A Practical Guide for Biomedical Researchers
Selecting an appropriate molecular clock model is a critical, yet often challenging, step in Bayesian phylogenetic analysis for studying pathogen evolution, drug resistance, and disease origins.
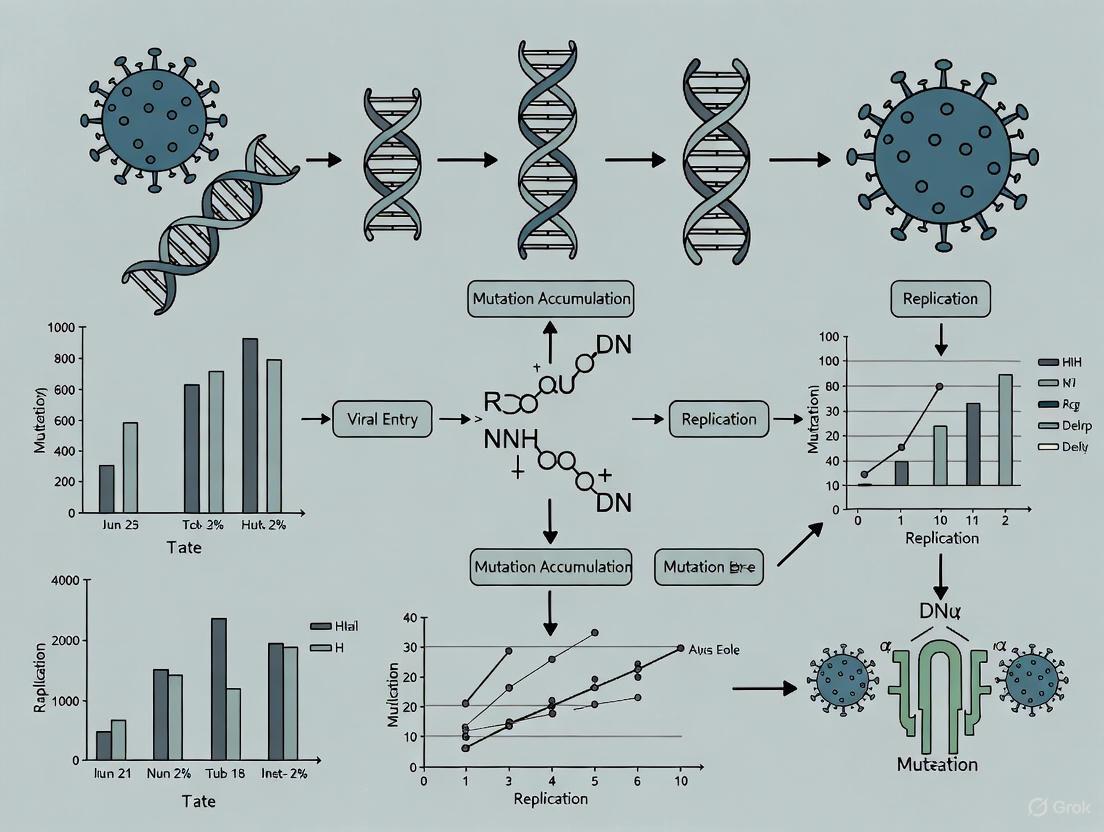
Viral Mutation Accumulation Studies: From Evolutionary Dynamics to Antiviral Drug Development
This article provides a comprehensive analysis of mutation accumulation studies in viruses, exploring the fundamental principles that govern viral evolution and their direct applications in biomedical research.
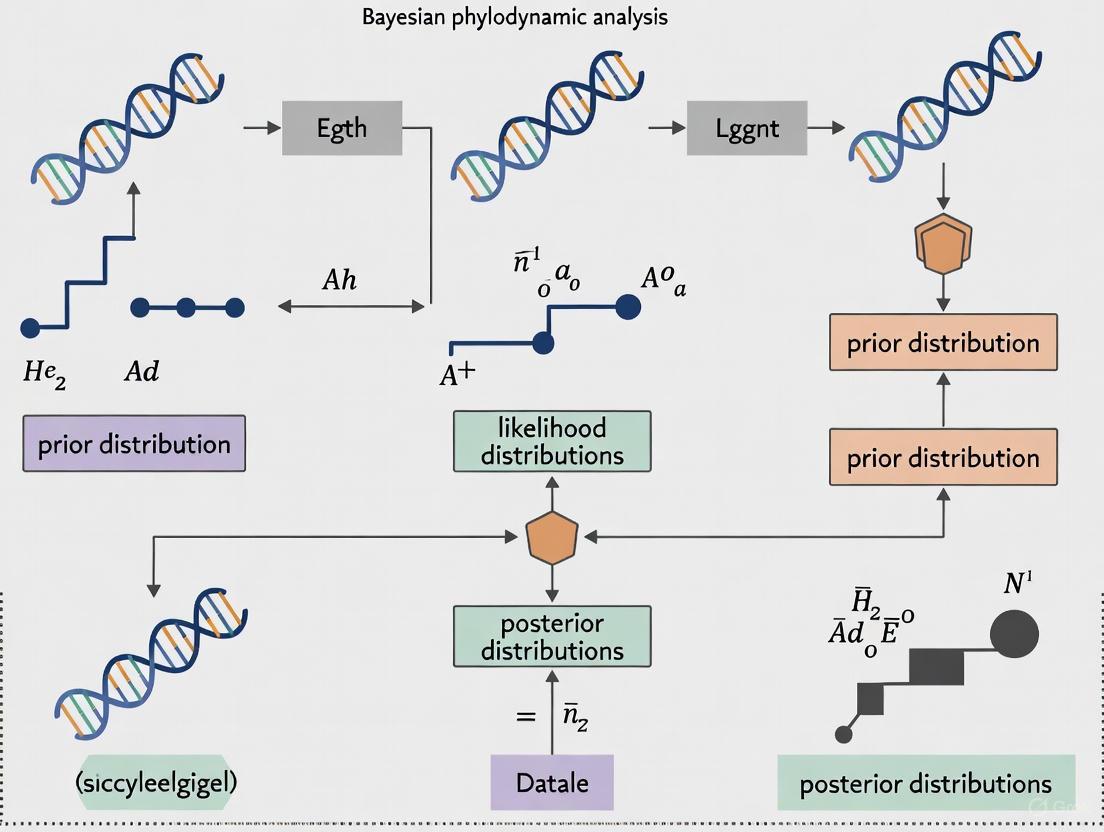
Bayesian Phylodynamics: Modeling Epidemic Spread from Genomic Data
This article provides a comprehensive overview of Bayesian phylodynamic methods for analyzing epidemic spread, tailored for researchers, scientists, and drug development professionals.
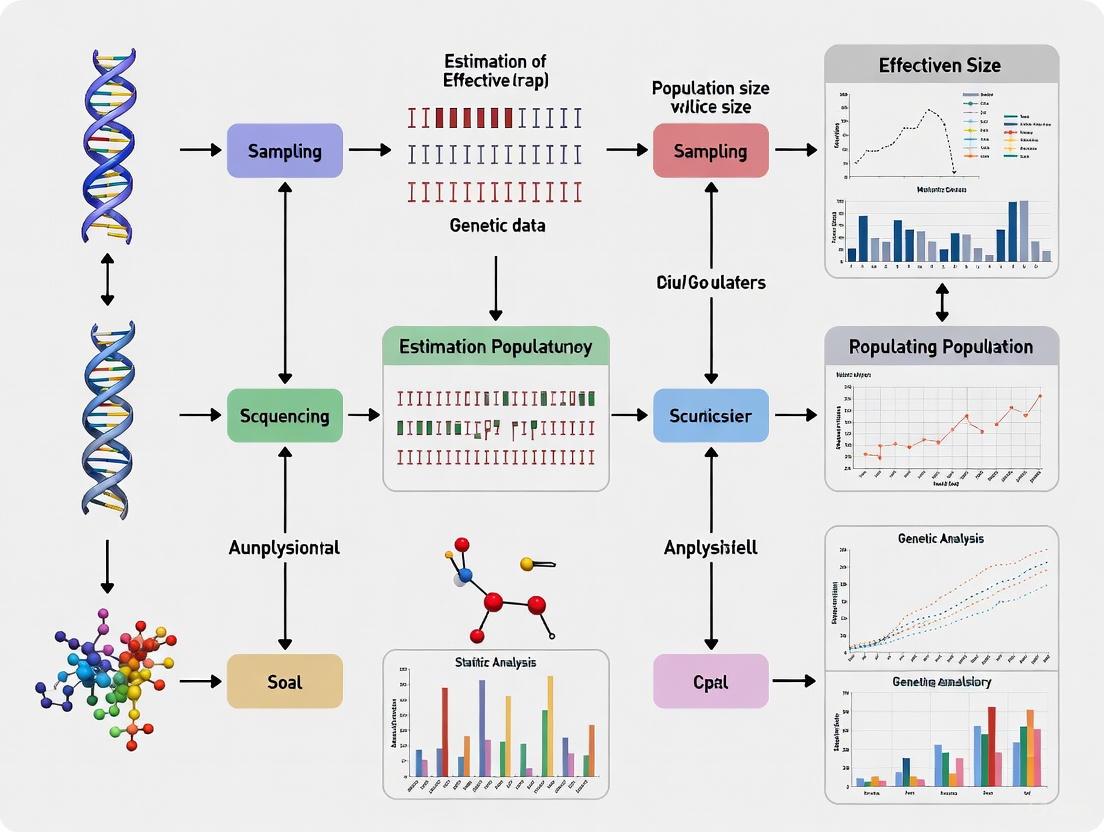
Estimating Effective Population Size from Genetic Data: A Comprehensive Guide for Biomedical Researchers
This article provides a comprehensive overview of methodologies for estimating effective population size (Ne) from genetic data, tailored for researchers, scientists, and drug development professionals.
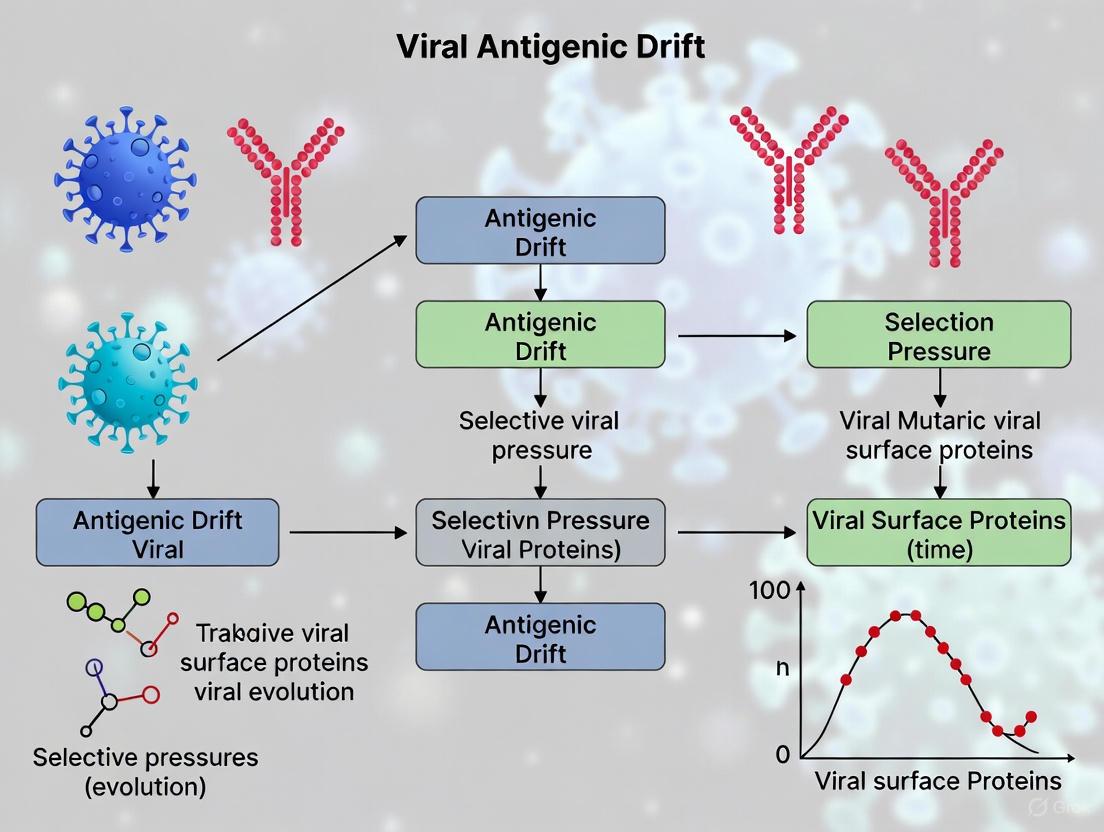
Selective Pressures Driving Viral Antigenic Drift: Mechanisms, Models, and Therapeutic Challenges
Antigenic drift, the gradual accumulation of mutations in viral surface proteins, is a primary mechanism for viral immune evasion, fundamentally challenging the durability of vaccines and therapeutics.
Codon Capture vs. Ambiguous Intermediate: Resolving the Mechanisms of Genetic Code Evolution
This article provides a comprehensive comparison of the Codon Capture and Ambiguous Intermediate theories, the two leading frameworks explaining genetic code evolution and reassignment.