Research Articles
A Practical Guide to Explicit Solvent Molecular Dynamics for Drug Discovery and Biomolecular Research
This comprehensive guide provides researchers and drug development professionals with foundational knowledge and advanced methodologies for performing molecular dynamics simulations with explicit solvent.
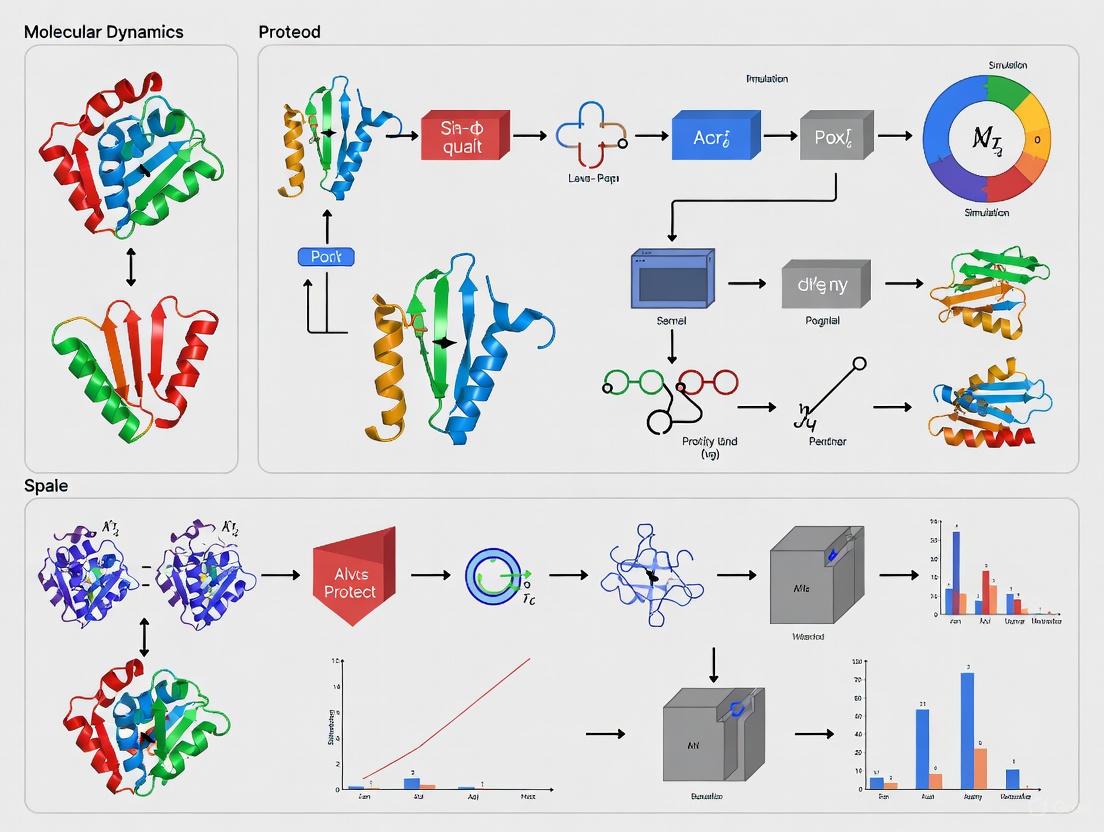
From Folding Pathways to Drug Discovery: Leveraging Molecular Dynamics for Protein Simulation
This article provides a comprehensive overview of how Molecular Dynamics (MD) simulations have revolutionized the study of protein folding, moving beyond static structures to capture dynamic conformational ensembles.
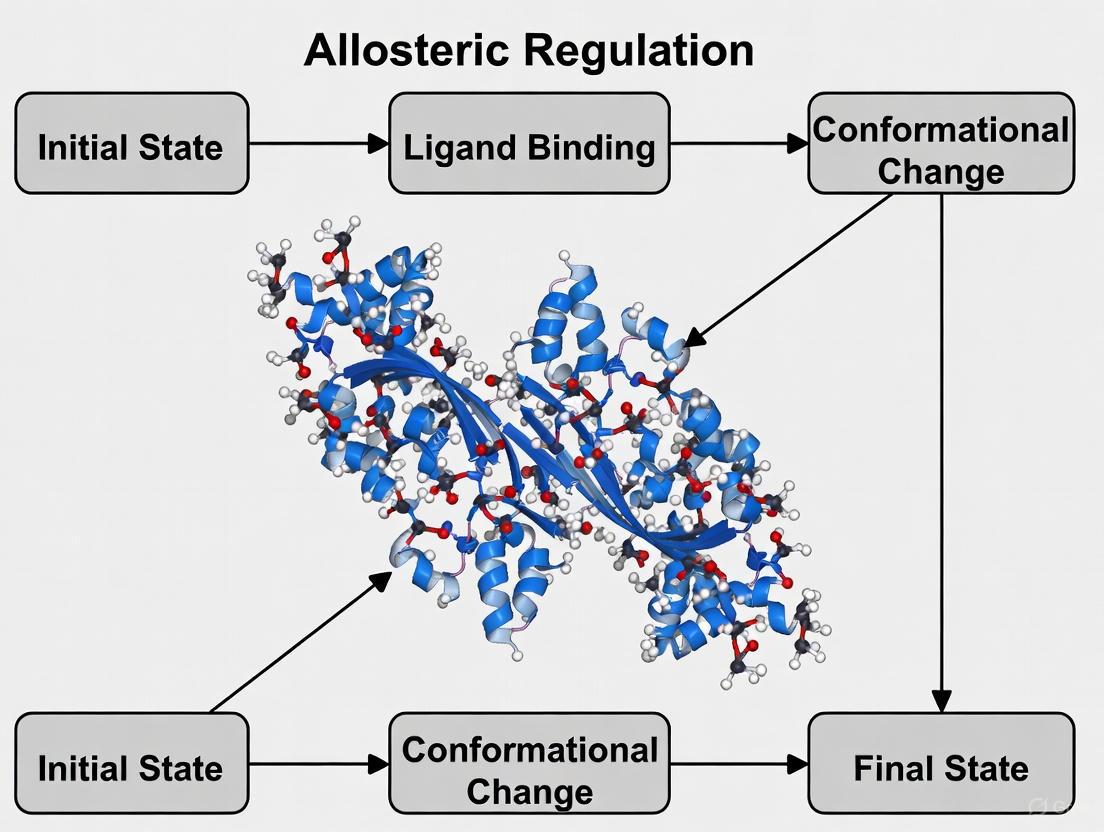
Decoding Allostery: How Molecular Dynamics Simulations Are Revolutionizing Drug Discovery
Allosteric regulation, the process of controlling protein function through binding at distal sites, offers a promising avenue for developing highly selective therapeutics.
Molecular Dynamics for Protein-Ligand Complex Analysis: A Comprehensive Guide from Fundamentals to Advanced Applications in Drug Discovery
This article provides a comprehensive overview of Molecular Dynamics (MD) simulations for analyzing protein-ligand complexes, a critical methodology in structural biology and rational drug design.
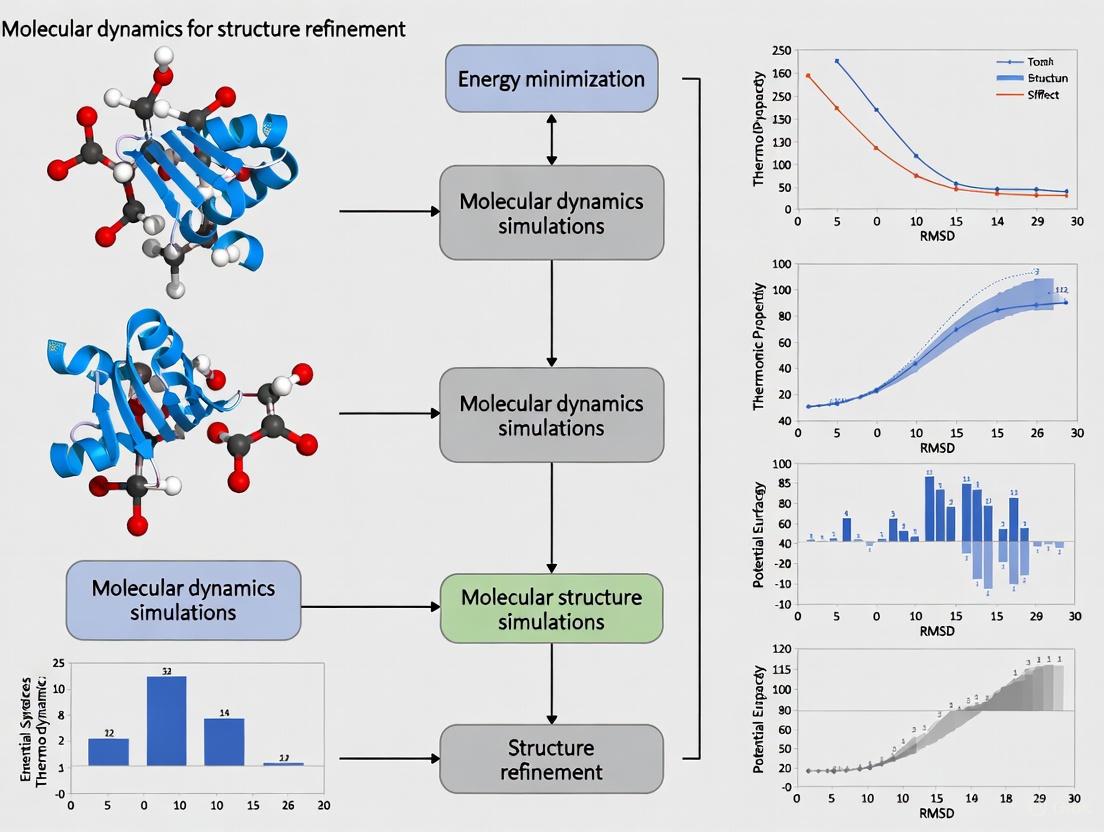
Molecular Dynamics for Protein Structure Refinement: A Practical Guide for Biomedical Researchers
This article provides a comprehensive overview of Molecular Dynamics (MD) simulation as a powerful tool for refining predicted and experimental protein structures.
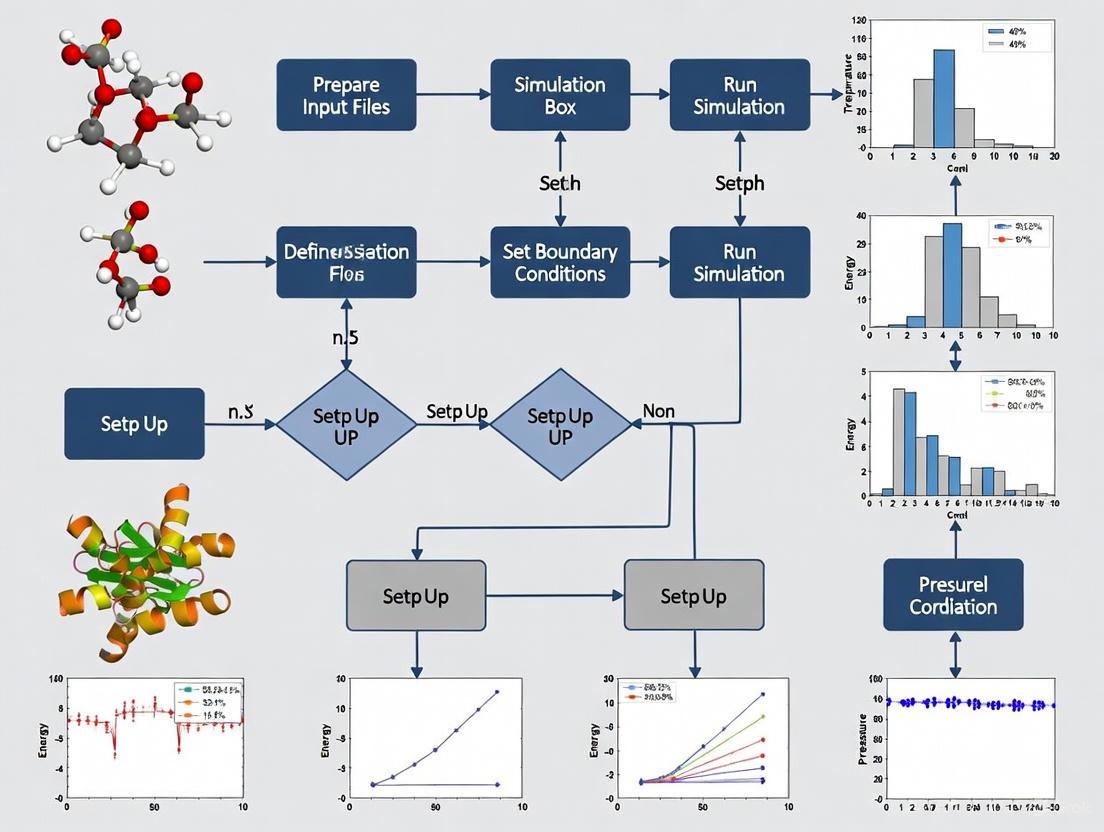
A Practical Guide to Setting Up Molecular Dynamics Simulations in GROMACS: From Fundamentals to Advanced Applications
This guide provides a comprehensive, structured approach for researchers, scientists, and drug development professionals to set up and execute molecular dynamics (MD) simulations using GROMACS.
Molecular Dynamics Ensembles: A Guide to Theory, Application, and Validation for Biomedical Research
This article provides a comprehensive guide to thermodynamic ensembles in Molecular Dynamics (MD) simulations, tailored for researchers and professionals in drug development.
A Comprehensive Guide to Molecular Dynamics Simulation Protocols for Protein Systems
This article provides a complete roadmap for researchers and drug development professionals to design, execute, and validate molecular dynamics (MD) simulations of protein systems.
Beyond Guesswork: A Systematic Framework for Molecular Dynamics Equilibration in Biomedical Research
This article provides a comprehensive guide to the molecular dynamics (MD) equilibration process, a critical yet often heuristic step for obtaining physically meaningful simulation results.
From Atoms to Action: How Newton's Equations Power Molecular Dynamics in Drug Discovery
This article provides a comprehensive overview of the central role of Newton's equations of motion in molecular dynamics (MD) simulations, tailored for researchers and professionals in drug development.