Research Articles
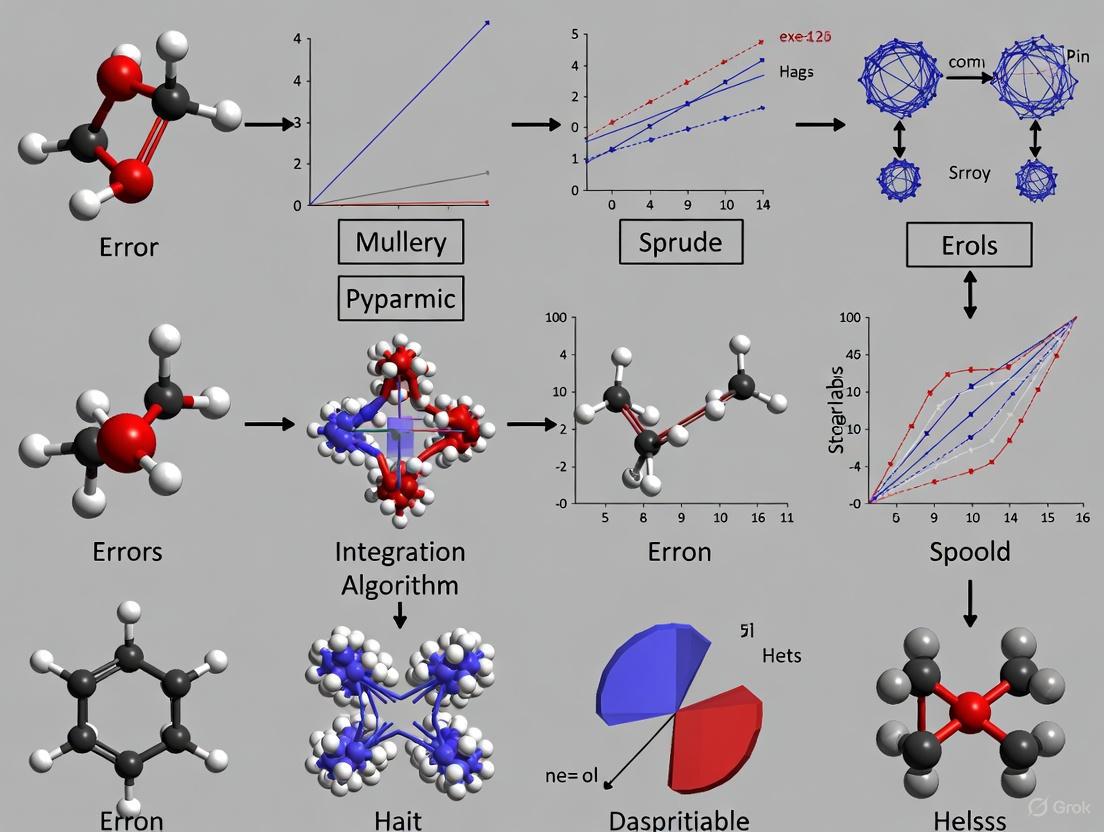
Navigating Molecular Dynamics Integration Errors: From Fundamentals to AI-Enhanced Solutions in Drug Discovery
This article provides a comprehensive analysis of molecular dynamics integration algorithm errors, a critical yet often overlooked factor determining the reliability of simulations in biomedical research.
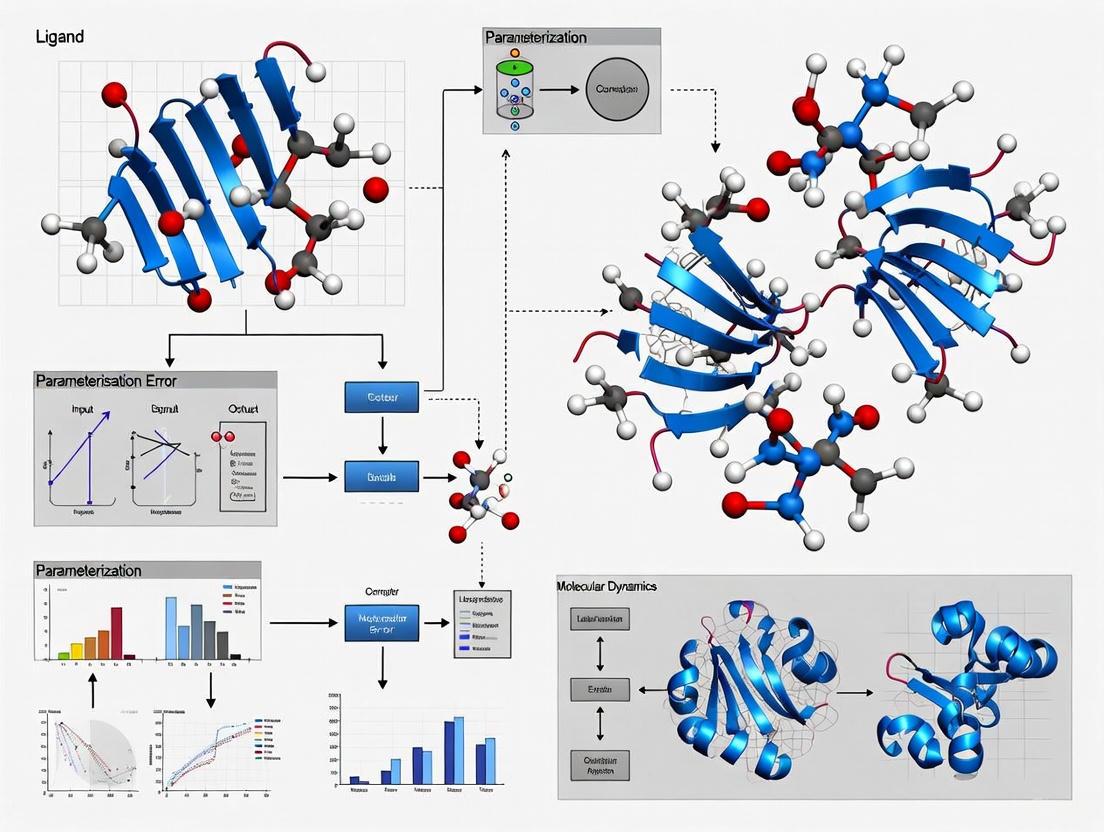
Navigating Ligand Parameterization Errors in Molecular Dynamics: From Force Field Pitfalls to Reliable Drug Discovery
Accurate ligand parameterization is a critical, yet often error-prone, foundation for molecular dynamics (MD) simulations in drug discovery.
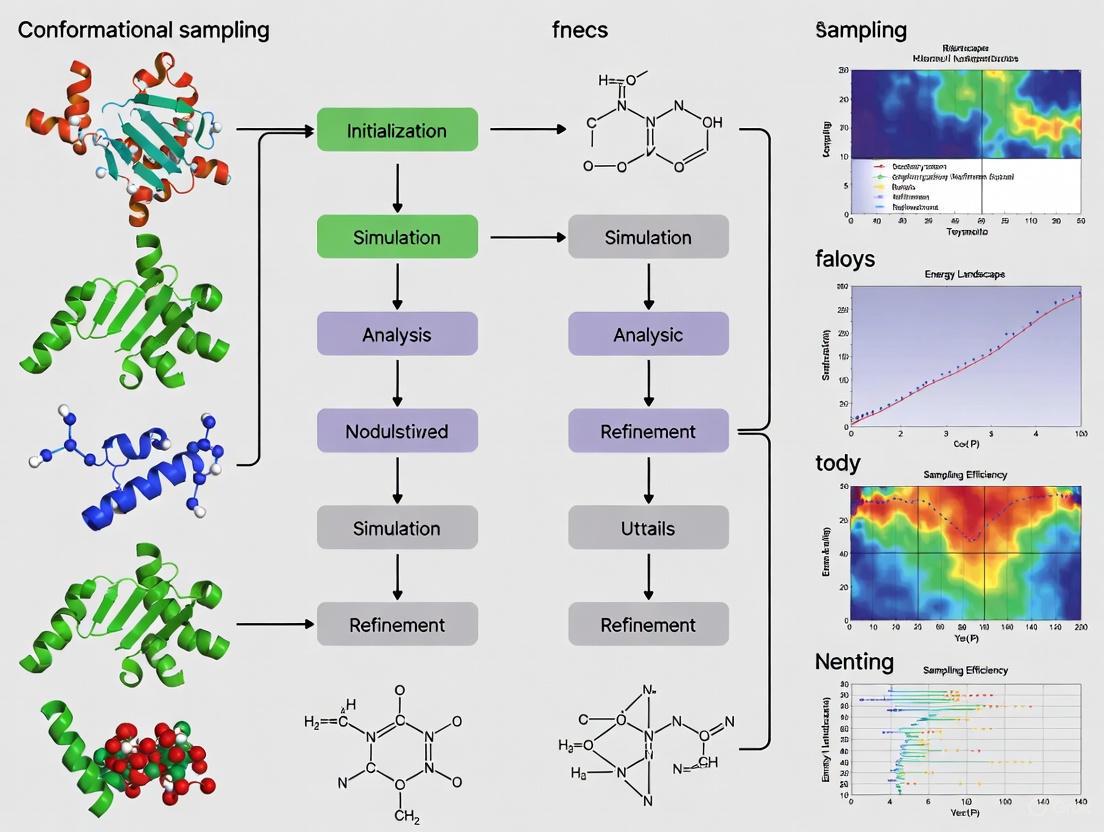
Beyond the Time Scale Barrier: Advanced Strategies for Enhanced Conformational Sampling in Molecular Dynamics
Adequately sampling the vast conformational landscape of proteins, especially highly dynamic or disordered systems, remains a central challenge in molecular dynamics (MD) simulations.
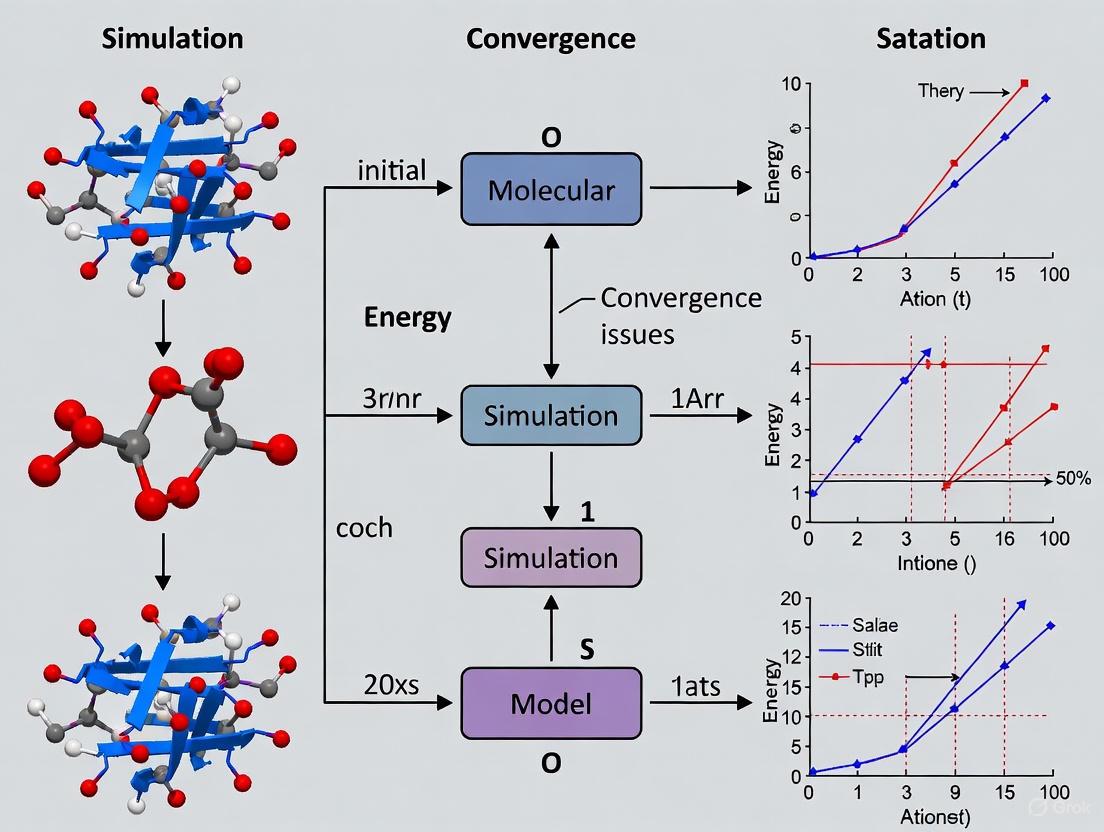
Navigating Molecular Dynamics Convergence: From Fundamental Challenges to Advanced Solutions in Drug Discovery
This article provides a comprehensive guide to molecular dynamics (MD) simulation convergence, a critical challenge in computational drug discovery.
Solving Molecular Dynamics Equilibration: A Practical Guide to Temperature and Pressure Control for Reliable Simulations
Achieving proper temperature and pressure equilibration in Molecular Dynamics (MD) simulations is a critical yet often challenging step for obtaining physically meaningful results in biomedical research, particularly in drug development.
Molecular Dynamics Simulation Crashes: A Comprehensive Guide from Prevention to Recovery for Biomedical Researchers
This article provides a systematic framework for researchers and drug development professionals to handle molecular dynamics simulation crashes.
Accelerate Your Research: A Practical Guide to Optimizing Slow Molecular Dynamics Simulations
Molecular dynamics (MD) simulations are a cornerstone of computational chemistry, biophysics, and drug discovery, yet their extreme computational cost often hinders research progress.
Solving Molecular Dynamics Energy Minimization Failures: A Troubleshooting Guide for Computational Researchers
This article provides a comprehensive framework for understanding, troubleshooting, and resolving common failures in molecular dynamics (MD) energy minimization.
Avoiding Common Pitfalls in Molecular Dynamics Simulations: A Guide to Error Correction, Force Field Selection, and Best Practices
This article provides a comprehensive guide for researchers and drug development professionals on identifying, troubleshooting, and preventing common errors in molecular dynamics (MD) simulations.
How to Fix Molecular Dynamics System Instability: A Troubleshooting Guide for Researchers
Molecular dynamics (MD) simulations are powerful but prone to instability that can invalidate results and waste computational resources.