Research Articles
Cross-Method Validation of ARG Concentration Techniques: A Guide for Robust Antimicrobial Resistance Surveillance
Accurate monitoring of Antibiotic Resistance Genes (ARGs) is critical for public health, but the diversity of concentration and detection methods challenges data comparability.
Comparative Genomic Analysis of Multidrug-Resistant Escherichia coli: Decoding the Resistome, Virulome, and Evolutionary Pathways
This article provides a comprehensive analysis of multidrug-resistant (MDR) Escherichia coli through a comparative genomics lens, tailored for researchers, scientists, and drug development professionals.
Mitigating Contamination in Metagenomic Antibiotic Resistance Gene Analysis: A Comprehensive Guide for Researchers
Accurate metagenomic analysis of Antibiotic Resistance Genes (ARGs) is paramount for public health surveillance and drug development, yet is critically threatened by contamination and analytical artifacts.
Standardizing ARG Concentration Methods for Wastewater Surveillance: A Comprehensive Guide for Researchers
The accurate monitoring of antibiotic resistance genes (ARGs) in wastewater is critical for public health surveillance and understanding environmental resistance dissemination.
Optimizing CARD Database Bit-Score Thresholds: A Guide for Enhanced ARG Detection in Biomedical Research
Accurate identification of Antibiotic Resistance Genes (ARGs) is critical for public health surveillance and drug development.
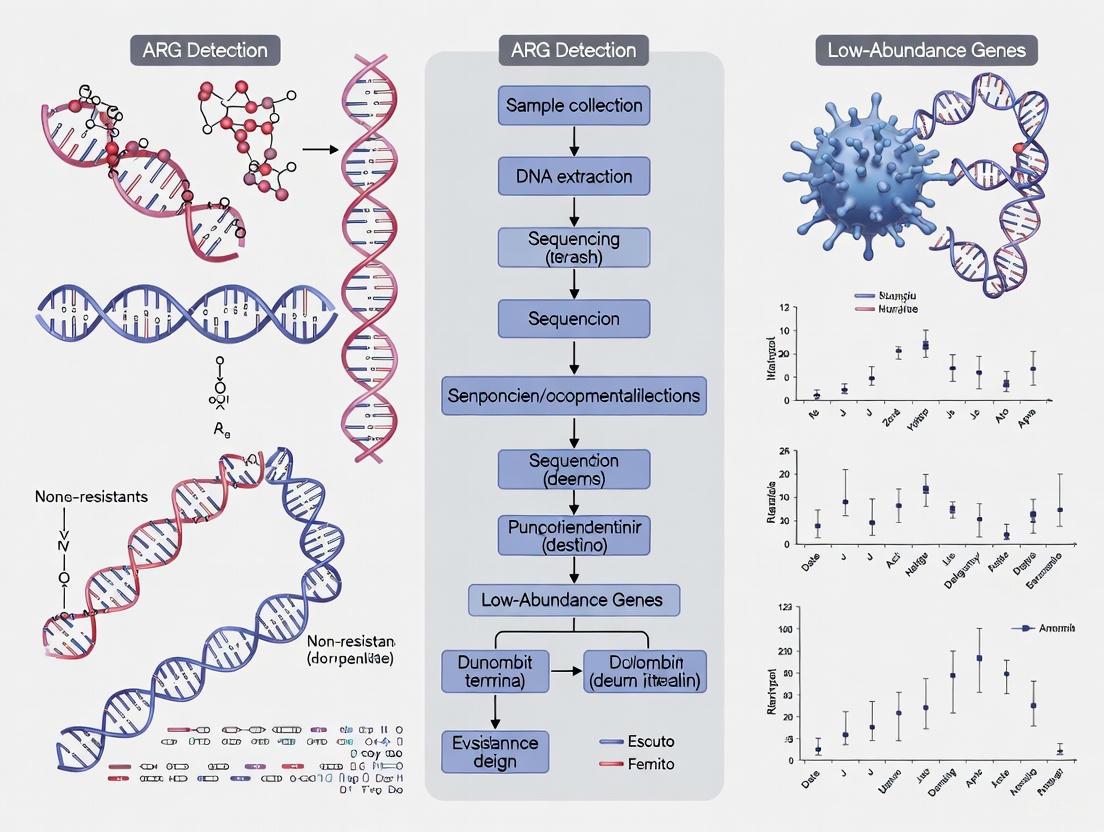
Breaking the Detection Limit: Advanced Strategies for Identifying Low-Abundance Antibiotic Resistance Genes in Complex Matrices
The accurate detection of low-abundance antibiotic resistance genes (ARGs) in complex environmental and clinical matrices is critical for global antimicrobial resistance (AMR) surveillance and risk assessment.
Advanced Metagenomic Sequencing Protocols for Comprehensive Resistome Analysis: From Workflow Design to Clinical Application
This article provides researchers, scientists, and drug development professionals with a comprehensive framework for implementing metagenomic sequencing in resistome analysis.
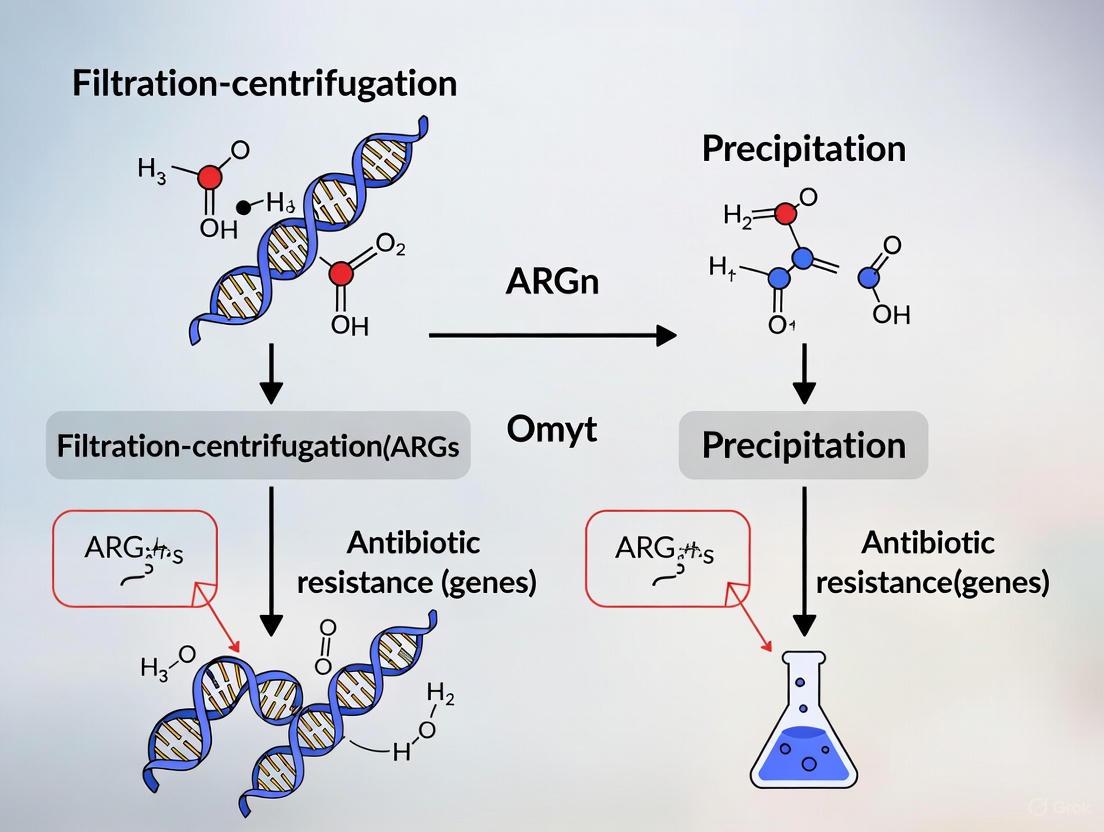
Filtration-Centrifugation vs. Precipitation: A Comparative Guide for Optimizing ARG Concentration in Environmental Samples
Antimicrobial resistance (AMR) poses a significant global health threat, and effective environmental surveillance of antibiotic resistance genes (ARGs) is critical under the One Health framework.
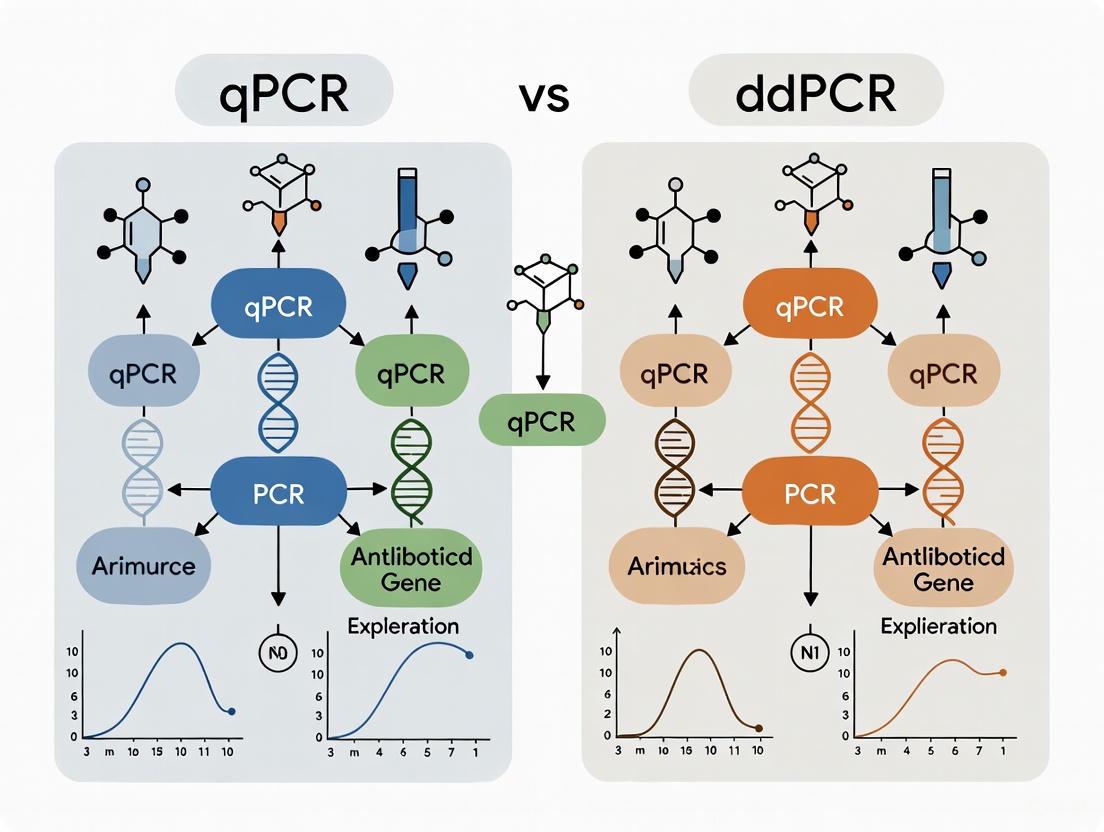
qPCR vs ddPCR: A Strategic Guide for Antibiotic Resistance Gene Quantification in Biomedical Research
This article provides a comprehensive comparison of Quantitative Real-Time PCR (qPCR) and Droplet Digital PCR (ddPCR) for the detection and quantification of antibiotic resistance genes (ARGs), a critical task in...
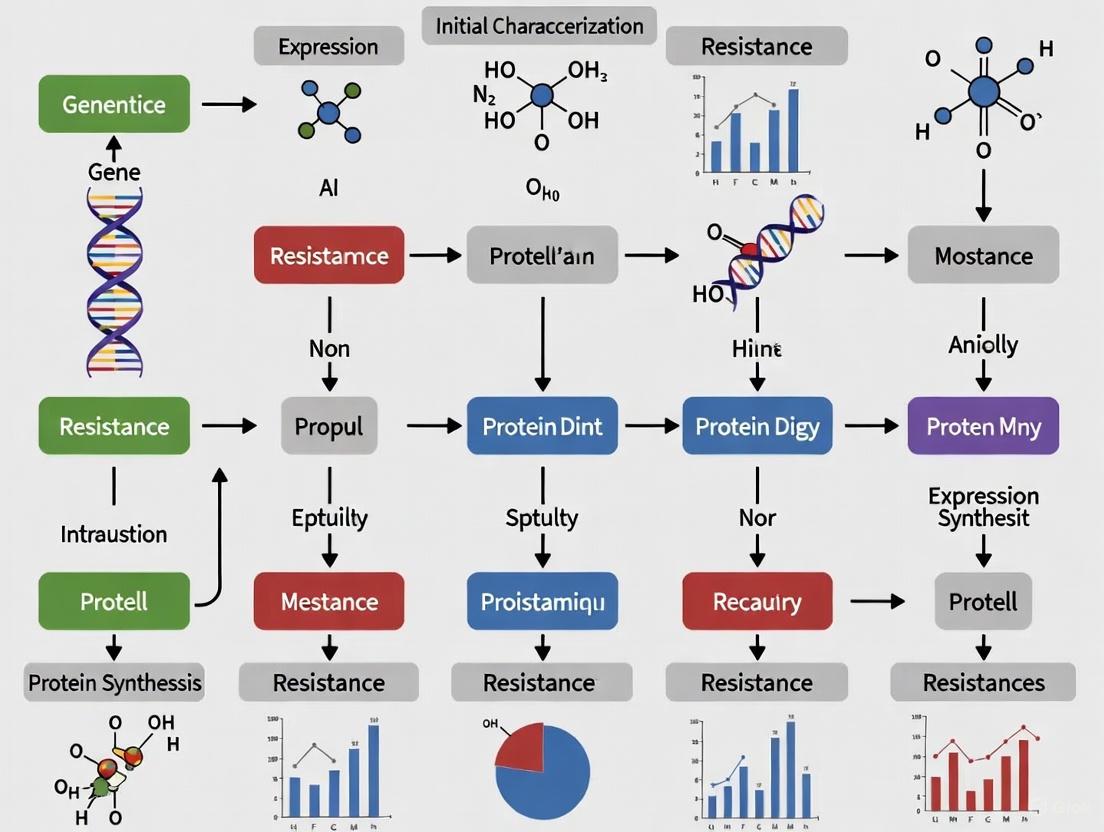
Decoding Multidrug Resistance: Genomic Characterization, Mechanisms, and Novel Therapeutic Strategies
This comprehensive review explores the initial characterization of multidrug resistance (MDR) genes, addressing a critical challenge in modern healthcare.