Research Articles
Validating Molecular Dynamics Thermodynamic Properties: From Foundational Principles to AI-Enhanced Workflows in Drug Discovery
This article provides a comprehensive guide for researchers and drug development professionals on the validation of thermodynamic properties derived from Molecular Dynamics (MD) simulations.
GROMACS vs AMBER vs NAMD: A 2025 Comparative Guide for Molecular Dynamics Simulations
This article provides a comprehensive, up-to-date comparison of the three leading molecular dynamics software packages—GROMACS, AMBER, and NAMD—tailored for researchers, scientists, and drug development professionals.
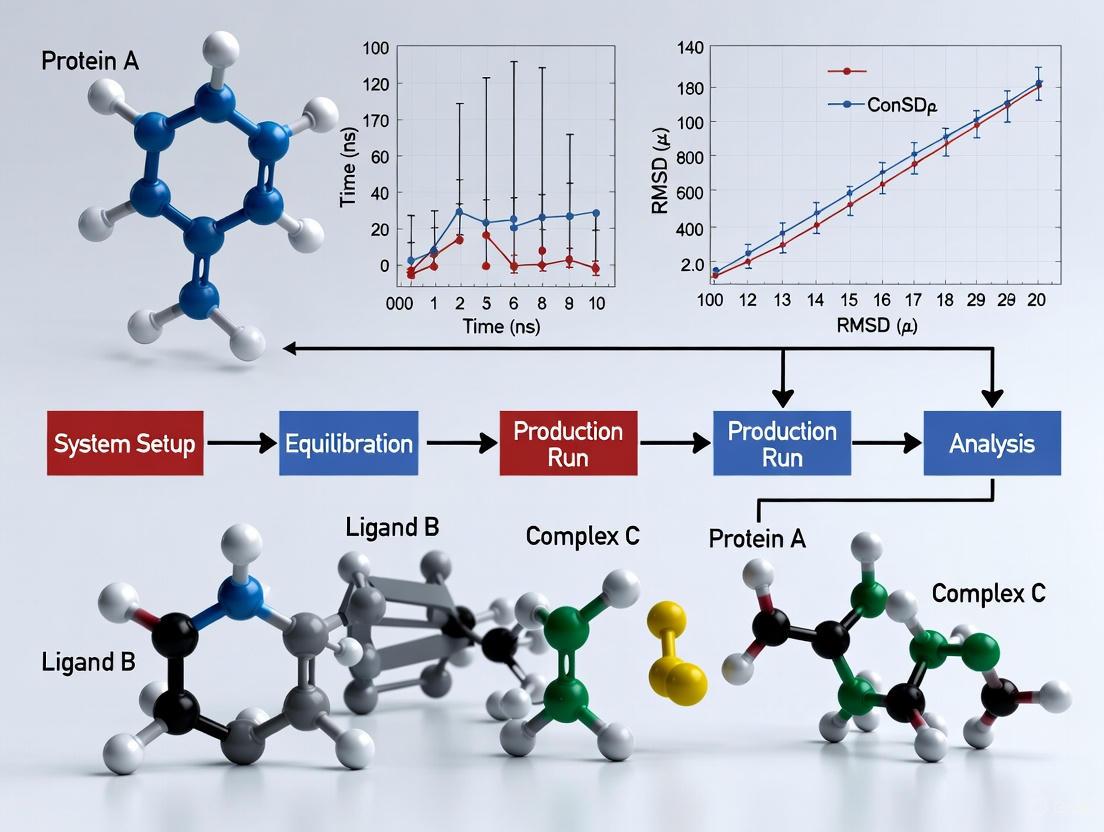
Validating Protein-Ligand Binding with Molecular Dynamics: A Comprehensive Guide from Theory to Practice
This article provides a comprehensive framework for researchers and drug development professionals to validate protein-ligand binding using Molecular Dynamics (MD) simulations.
Validating Molecular Dynamics Convergence: A Comprehensive Guide for Reliable Biomolecular Simulations
This article provides a comprehensive framework for validating convergence in molecular dynamics (MD) simulations, a critical step for ensuring the reliability and reproducibility of results in biomedical research and drug...
Bridging the Virtual and the Real: A Practical Guide to Validating Molecular Dynamics Simulations with Experimental Data
This article provides a comprehensive guide for researchers and drug development professionals on the critical process of validating molecular dynamics (MD) simulations against experimental data.
Overcoming Molecular Dynamics Trajectory Analysis Challenges: A Guide for Biomedical Researchers
Molecular dynamics (MD) simulations generate vast, complex trajectory data that poses significant analysis challenges for researchers in drug development and biomedical sciences.
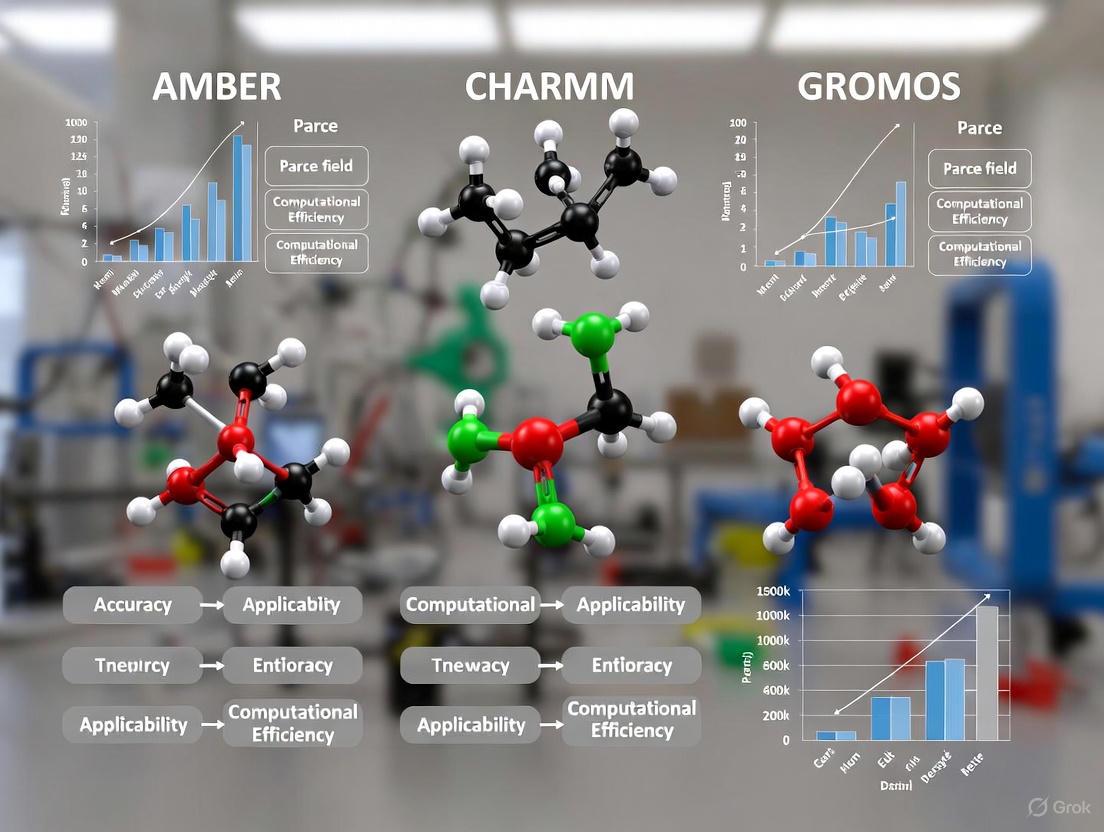
AMBER vs CHARMM vs GROMOS: A Comprehensive Force Field Comparison for Biomolecular Simulation
This article provides a systematic comparison of the AMBER, CHARMM, and GROMOS molecular dynamics force fields, essential tools for researchers in drug discovery and pharmaceutical development.
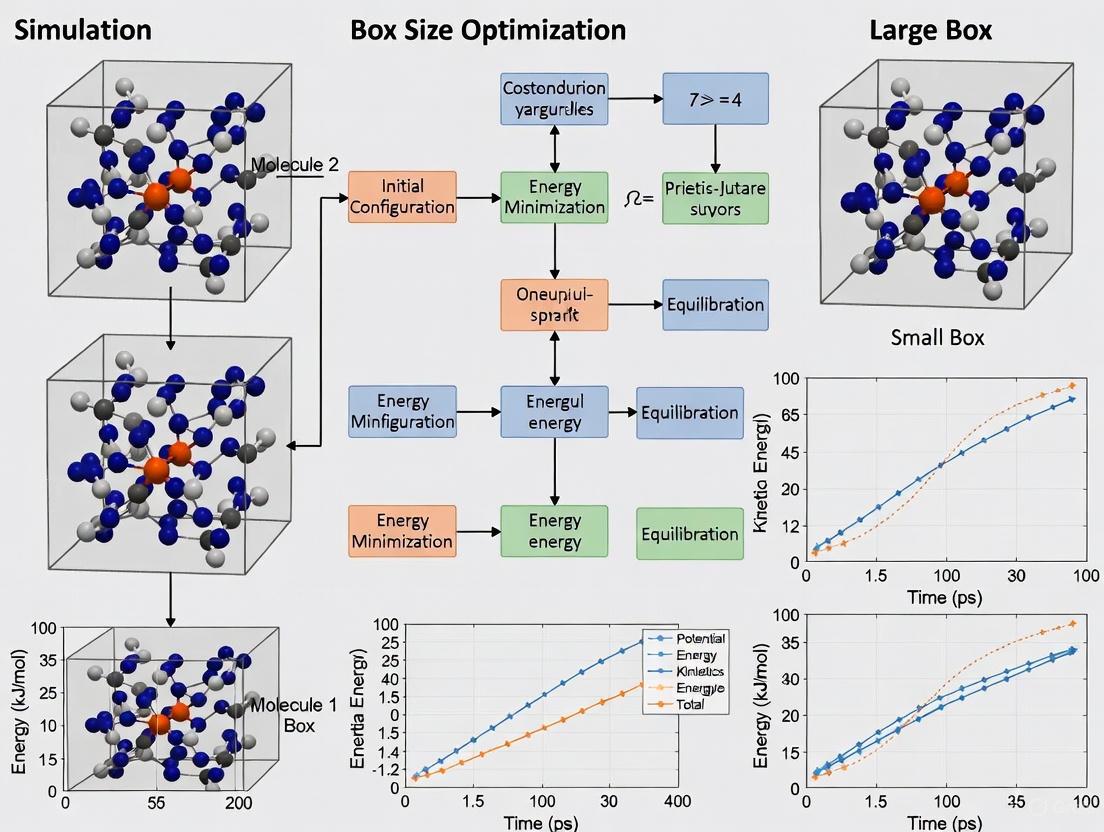
Molecular Dynamics Simulation Box Size Optimization: Balancing Accuracy and Computational Efficiency
This comprehensive review examines molecular dynamics simulation box size optimization strategies for biomedical and drug development applications.
Navigating Molecular Dynamics Integration Errors: From Fundamentals to AI-Enhanced Solutions in Drug Discovery
This article provides a comprehensive analysis of molecular dynamics integration algorithm errors, a critical yet often overlooked factor determining the reliability of simulations in biomedical research.
Navigating Ligand Parameterization Errors in Molecular Dynamics: From Force Field Pitfalls to Reliable Drug Discovery
Accurate ligand parameterization is a critical, yet often error-prone, foundation for molecular dynamics (MD) simulations in drug discovery.